

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC122724.4 - phase: 0
(357 letters)
Database: LJGI
28,460 sequences; 14,692,800 total letters
Searching..................................................done
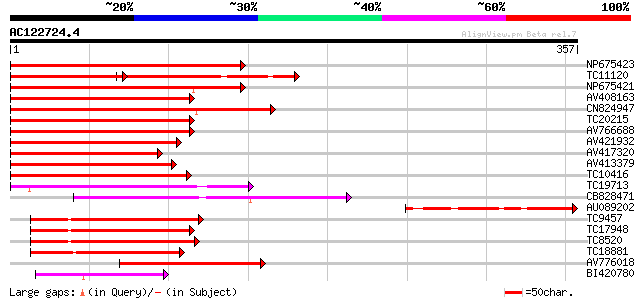
Score E
Sequences producing significant alignments: (bits) Value
NP675423 transcription factor MYB101 [Lotus japonicus] 232 8e-62
TC11120 UP|Q84PP4 (Q84PP4) Transcription factor MYB102, complete 142 1e-58
NP675421 transcription factor MYB103 [Lotus japonicus] 205 1e-53
AV408163 178 1e-45
CN824947 176 5e-45
TC20215 similar to AAS10104 (AAS10104) MYB transcription factor,... 174 2e-44
AV766688 166 4e-42
AV421932 163 5e-41
AV417320 161 2e-40
AV413379 160 2e-40
TC10416 weakly similar to UP|O04108 (O04108) MYB factor, partial... 160 3e-40
TC19713 homologue to UP|P93474 (P93474) Myb26, partial (55%) 141 2e-34
CB828471 129 8e-31
AU089202 125 8e-30
TC9457 homologue to UP|Q9FZ15 (Q9FZ15) Tuber-specific and sucros... 112 1e-25
TC17948 similar to UP|Q9FZ13 (Q9FZ13) Tuber-specific and sucrose... 110 4e-25
TC8520 homologue to UP|Q9FZ15 (Q9FZ15) Tuber-specific and sucros... 110 5e-25
TC18881 similar to UP|Q9FZ15 (Q9FZ15) Tuber-specific and sucrose... 104 3e-23
AV776018 86 7e-18
BI420780 71 2e-13
>NP675423 transcription factor MYB101 [Lotus japonicus]
Length = 1047
Score = 232 bits (591), Expect = 8e-62
Identities = 105/148 (70%), Positives = 123/148 (82%)
Frame = +1
Query: 1 MGRSPCCDKVGLKKGPWTPEEDQKLLAYIDEHGHGSWRALPAKAGLQRCGKSCRLRWTNY 60
MGR+PCCD++GLKKGPWTPEED L+ YI +HGHGSWRALP AGL RCGKSCRLRWTNY
Sbjct: 1 MGRTPCCDEIGLKKGPWTPEEDAILVDYIQKHGHGSWRALPKLAGLNRCGKSCRLRWTNY 180
Query: 61 LRPDIKRGKFSLQEEQTIIQLHALLGNRWSAIATHLAKRTDNEIKNYWNTHLKKRLTKMG 120
LRPDIKRGKF+ +EEQ II LHA+LGN+WSAIA HL RTDNEIKN+WNTHLKK+L +MG
Sbjct: 181 LRPDIKRGKFTEEEEQLIINLHAVLGNKWSAIAGHLPGRTDNEIKNFWNTHLKKKLLQMG 360
Query: 121 IDPVTHKPKNDALLSSSDNAHSKTASNL 148
+DPVTH+P+ D L S+ A+N+
Sbjct: 361 LDPVTHRPRLDHLNLLSNLQQFLAATNM 444
>TC11120 UP|Q84PP4 (Q84PP4) Transcription factor MYB102, complete
Length = 1281
Score = 142 bits (358), Expect(2) = 1e-58
Identities = 62/74 (83%), Positives = 67/74 (89%)
Frame = +1
Query: 1 MGRSPCCDKVGLKKGPWTPEEDQKLLAYIDEHGHGSWRALPAKAGLQRCGKSCRLRWTNY 60
MGRSPCCD+ GLKKGPWTPEEDQKL+ +I +HGHGSWRALP AGL RCGKSCRLRWTNY
Sbjct: 67 MGRSPCCDENGLKKGPWTPEEDQKLVEHIQKHGHGSWRALPKLAGLNRCGKSCRLRWTNY 246
Query: 61 LRPDIKRGKFSLQE 74
LRPDIKRGKFS +E
Sbjct: 247 LRPDIKRGKFSPEE 288
Score = 101 bits (251), Expect(2) = 1e-58
Identities = 50/115 (43%), Positives = 76/115 (65%)
Frame = +2
Query: 68 GKFSLQEEQTIIQLHALLGNRWSAIATHLAKRTDNEIKNYWNTHLKKRLTKMGIDPVTHK 127
G F ++EQTI+ LH++LGN+WS IATHL RTDNEIKN+WNTHLKK+L +MG DP+TH+
Sbjct: 269 GNFLQKKEQTILHLHSILGNKWSTIATHLPGRTDNEIKNFWNTHLKKKLIQMGFDPMTHQ 448
Query: 128 PKNDALLSSSDNAHSKTASNLSHMAQWESARLEAEARLVRESKIRSHNSLLHHNH 182
P+ D + S + +N++ + + +A V+ +K++ N LL ++
Sbjct: 449 PRTDLV---STLPYLLALANMTQLMDHNHQSFDEQA--VQLAKLQYLNYLLQSSN 598
>NP675421 transcription factor MYB103 [Lotus japonicus]
Length = 924
Score = 205 bits (521), Expect = 1e-53
Identities = 97/150 (64%), Positives = 117/150 (77%), Gaps = 2/150 (1%)
Frame = +1
Query: 1 MGRSPCCDKVGLKKGPWTPEEDQKLLAYIDEHGHGSWRALPAKAGLQRCGKSCRLRWTNY 60
M R+P D+ G+KKGPW+ EED+ L+ YI +HGHGSWRALP AGL RCGKSCRLRWTNY
Sbjct: 1 MMRTPSIDESGMKKGPWSQEEDKVLVDYIQKHGHGSWRALPQLAGLNRCGKSCRLRWTNY 180
Query: 61 LRPDIKRGKFSLQEEQTIIQLHALLGNRWSAIATHLAKRTDNEIKNYWNTHLKK--RLTK 118
LRPDIKRGKFS +EEQ II LHA LGN+W+ IA+HL RTDNEIKN WNTHLKK +L +
Sbjct: 181 LRPDIKRGKFSDEEEQLIINLHASLGNKWATIASHLPGRTDNEIKNLWNTHLKKKLKLLQ 360
Query: 119 MGIDPVTHKPKNDALLSSSDNAHSKTASNL 148
MG+DPVTH P++D + S+ A+N+
Sbjct: 361 MGLDPVTHCPRSDHMNILSNLQQILAAANI 450
>AV408163
Length = 425
Score = 178 bits (452), Expect = 1e-45
Identities = 76/116 (65%), Positives = 94/116 (80%)
Frame = +2
Query: 1 MGRSPCCDKVGLKKGPWTPEEDQKLLAYIDEHGHGSWRALPAKAGLQRCGKSCRLRWTNY 60
MGRSPCC+K + KG WT EED +L++YI HG G WR+LP AGL RCGKSCRLRW NY
Sbjct: 77 MGRSPCCEKAHINKGAWTKEEDDRLISYIRAHGEGCWRSLPKAAGLLRCGKSCRLRWINY 256
Query: 61 LRPDIKRGKFSLQEEQTIIQLHALLGNRWSAIATHLAKRTDNEIKNYWNTHLKKRL 116
LRPD+KRG F+ +E++ II+LH+LLGN+WS IA L RTDNEIKNYWNTH++++L
Sbjct: 257 LRPDLKRGNFTEEEDELIIKLHSLLGNKWSLIAGRLPGRTDNEIKNYWNTHIRRKL 424
>CN824947
Length = 738
Score = 176 bits (446), Expect = 5e-45
Identities = 89/174 (51%), Positives = 111/174 (63%), Gaps = 7/174 (4%)
Frame = +1
Query: 1 MGRSPCCDKVGLKKGPWTPEEDQKLLAYIDEHGHGSWRALPAKAGLQRCGKSCRLRWTNY 60
MGR+PCC+KVGLK+G WT EED+ L YI G GSWR+LP AGL RCGKSCRLRW NY
Sbjct: 151 MGRAPCCEKVGLKRGRWTAEEDELLTKYIQASGEGSWRSLPKNAGLLRCGKSCRLRWINY 330
Query: 61 LRPDIKRGKFSLQEEQTIIQLHALLGNRWSAIATHLAKRTDNEIKNYWNTHLKKRL---- 116
LR D+KRG S +EE I++LHA GNRWS IA+H+ RTDNEIKNYWN+HL ++L
Sbjct: 331 LRGDLKRGNISDEEESLIVKLHASFGNRWSLIASHMPGRTDNEIKNYWNSHLSRKLVYTF 510
Query: 117 ---TKMGIDPVTHKPKNDALLSSSDNAHSKTASNLSHMAQWESARLEAEARLVR 167
T I K + S +KT S +++ Q ++ LE E + R
Sbjct: 511 RGSTTKSIVDTPPKRRRCGRTSRWAMKKNKTYSLINNDLQNQNNELEKEDPVAR 672
>TC20215 similar to AAS10104 (AAS10104) MYB transcription factor, partial
(35%)
Length = 464
Score = 174 bits (442), Expect = 2e-44
Identities = 74/116 (63%), Positives = 95/116 (81%)
Frame = +2
Query: 1 MGRSPCCDKVGLKKGPWTPEEDQKLLAYIDEHGHGSWRALPAKAGLQRCGKSCRLRWTNY 60
MGR+PCC+KVGLK+G W+ EED+ L +I ++G GSW++LP AGL RCGKSCRLRW NY
Sbjct: 29 MGRAPCCEKVGLKRGRWSEEEDKILTDFIKQNGEGSWKSLPKNAGLLRCGKSCRLRWINY 208
Query: 61 LRPDIKRGKFSLQEEQTIIQLHALLGNRWSAIATHLAKRTDNEIKNYWNTHLKKRL 116
LR D+KRG + +EE+ I++LHA LGNRWS IA HL RTDNEIKNYWN+HL++++
Sbjct: 209 LREDVKRGNITSEEEEIIVKLHAALGNRWSVIAGHLPGRTDNEIKNYWNSHLRRKI 376
>AV766688
Length = 608
Score = 166 bits (421), Expect = 4e-42
Identities = 74/116 (63%), Positives = 91/116 (77%)
Frame = +2
Query: 1 MGRSPCCDKVGLKKGPWTPEEDQKLLAYIDEHGHGSWRALPAKAGLQRCGKSCRLRWTNY 60
M R+P CDK GL+KG WT EED+KL+AY+ +G +WR LP AGL RCGKSCRLRW NY
Sbjct: 77 MARTPSCDKSGLRKGTWTAEEDRKLIAYVTRYGCWNWRQLPKFAGLARCGKSCRLRWLNY 256
Query: 61 LRPDIKRGKFSLQEEQTIIQLHALLGNRWSAIATHLAKRTDNEIKNYWNTHLKKRL 116
LRPDIKRG F+ +EE+ II++H LGNRWS IA L RTDNE+KN+W+T LK+R+
Sbjct: 257 LRPDIKRGHFTQEEEEIIIRMHKNLGNRWSIIAAELPGRTDNEVKNHWHTSLKRRV 424
>AV421932
Length = 402
Score = 163 bits (412), Expect = 5e-41
Identities = 69/108 (63%), Positives = 87/108 (79%)
Frame = +3
Query: 1 MGRSPCCDKVGLKKGPWTPEEDQKLLAYIDEHGHGSWRALPAKAGLQRCGKSCRLRWTNY 60
MG CC++ +K+G W+PEED+KL+ YI HG+G W +P KAGLQRCGKSCRLRW NY
Sbjct: 78 MGHHSCCNQQKVKRGLWSPEEDEKLIRYITTHGYGCWSEVPEKAGLQRCGKSCRLRWINY 257
Query: 61 LRPDIKRGKFSLQEEQTIIQLHALLGNRWSAIATHLAKRTDNEIKNYW 108
LRPDI+RG+F+ +EE+ II LH ++GNRW+ IA+HL RTDNEIKNYW
Sbjct: 258 LRPDIRRGRFTPEEEKLIISLHGVVGNRWAHIASHLPGRTDNEIKNYW 401
>AV417320
Length = 369
Score = 161 bits (407), Expect = 2e-40
Identities = 70/96 (72%), Positives = 82/96 (84%)
Frame = +3
Query: 1 MGRSPCCDKVGLKKGPWTPEEDQKLLAYIDEHGHGSWRALPAKAGLQRCGKSCRLRWTNY 60
M R+PCC+K+GLKKGPWT EEDQ L++YI +HGHG+WRALP AGL RCGKSCRLRW NY
Sbjct: 78 MVRAPCCEKIGLKKGPWTSEEDQILISYIQKHGHGNWRALPKHAGLLRCGKSCRLRWINY 257
Query: 61 LRPDIKRGKFSLQEEQTIIQLHALLGNRWSAIATHL 96
LRPDIKRG F+ +EE+ II++H LLGNRWSAIA L
Sbjct: 258 LRPDIKRGNFTAEEEELIIKMHELLGNRWSAIAAKL 365
>AV413379
Length = 416
Score = 160 bits (406), Expect = 2e-40
Identities = 72/105 (68%), Positives = 83/105 (78%)
Frame = +3
Query: 1 MGRSPCCDKVGLKKGPWTPEEDQKLLAYIDEHGHGSWRALPAKAGLQRCGKSCRLRWTNY 60
MGRSPCC+K KG WT EED +L+AYI HG G WR+LP AGL RCGKSCRLRW NY
Sbjct: 102 MGRSPCCEKAHTNKGAWTKEEDDRLIAYIRAHGEGCWRSLPKSAGLLRCGKSCRLRWINY 281
Query: 61 LRPDIKRGKFSLQEEQTIIQLHALLGNRWSAIATHLAKRTDNEIK 105
LRPD+KRG F+ QE++ II+LH+LLGN+WS IA LA RTDNEIK
Sbjct: 282 LRPDLKRGNFTPQEDELIIKLHSLLGNKWSLIAGRLAGRTDNEIK 416
>TC10416 weakly similar to UP|O04108 (O04108) MYB factor, partial (41%)
Length = 1046
Score = 160 bits (405), Expect = 3e-40
Identities = 70/114 (61%), Positives = 89/114 (77%)
Frame = +2
Query: 1 MGRSPCCDKVGLKKGPWTPEEDQKLLAYIDEHGHGSWRALPAKAGLQRCGKSCRLRWTNY 60
M R+P CDK G++KG WT EED+KL+AY+ +G +WR LP AGL RCGKSCRLRW NY
Sbjct: 92 MARTPSCDKSGMRKGTWTAEEDKKLIAYVTRYGCWNWRQLPKFAGLARCGKSCRLRWLNY 271
Query: 61 LRPDIKRGKFSLQEEQTIIQLHALLGNRWSAIATHLAKRTDNEIKNYWNTHLKK 114
LRP+IKRG F+ +EE+ II++H LGNRWS IA L RTD+E+KN+W+T LK+
Sbjct: 272 LRPNIKRGNFTQEEEEIIIRMHKKLGNRWSTIAAELPGRTDSEVKNHWHTTLKR 433
>TC19713 homologue to UP|P93474 (P93474) Myb26, partial (55%)
Length = 532
Score = 141 bits (355), Expect = 2e-34
Identities = 70/155 (45%), Positives = 93/155 (59%), Gaps = 2/155 (1%)
Frame = +3
Query: 1 MGRSPCCDKVG--LKKGPWTPEEDQKLLAYIDEHGHGSWRALPAKAGLQRCGKSCRLRWT 58
M + PC ++KGPWT EED L+ YI HG G W L AGL+R GKSCRLRW
Sbjct: 66 MDKKPCNSSHDPEVRKGPWTIEEDLILINYIANHGEGVWNTLAKSAGLKRTGKSCRLRWL 245
Query: 59 NYLRPDIKRGKFSLQEEQTIIQLHALLGNRWSAIATHLAKRTDNEIKNYWNTHLKKRLTK 118
NYLRPD++RG + +E+ I++LHA GNRWS IA HL RTDNEIKN+W T ++K + +
Sbjct: 246 NYLRPDVRRGNITPEEQLLIMELHAKWGNRWSKIAKHLPGRTDNEIKNFWRTRIQKHIKQ 425
Query: 119 MGIDPVTHKPKNDALLSSSDNAHSKTASNLSHMAQ 153
+ S ++ H + S +S MA+
Sbjct: 426 ------AESLQQQQSSSEINDHHQTSTSQMSTMAE 512
>CB828471
Length = 525
Score = 129 bits (324), Expect = 8e-31
Identities = 78/179 (43%), Positives = 97/179 (53%), Gaps = 4/179 (2%)
Frame = +1
Query: 41 PAKAGLQRCGKSCRLRWTNYLRPDIKRGKFSLQEEQTIIQLHALLGNRWSAIATHLAKRT 100
P +AGL RCGKSCRLRW NYLRPDIKRG FS +EE II LH LLGNRWSAIA L RT
Sbjct: 1 PKQAGLLRCGKSCRLRWANYLRPDIKRGNFSREEEDEIINLHELLGNRWSAIAARLPGRT 180
Query: 101 DNEIKNYWNTHLKKRLTKMGIDPVTHKPKNDALLSSSDNAHSKTASNLSH----MAQWES 156
DNEIKN W+THLKKRL K N L +S N +K Q S
Sbjct: 181 DNEIKNVWHTHLKKRLQPQ----TKQKQTNLDLDASKSNKDAKKEHQEDEGPRSPQQCSS 348
Query: 157 ARLEAEARLVRESKIRSHNSLLHHNHLKAWNINLESPTSTLSFNENVAPIMNSGEIGGE 215
+ + + L S I + + N+N +S ++L+ +E+ + S + GE
Sbjct: 349 SNSDNMSSLTNSSSIAPSTNHDDMSINDVHNVNFDSLENSLALDEDFWSEVLSSDNSGE 525
>AU089202
Length = 401
Score = 125 bits (315), Expect = 8e-30
Identities = 73/110 (66%), Positives = 81/110 (73%), Gaps = 2/110 (1%)
Frame = +3
Query: 250 VKEEGGDDQDQDQWKGY-ESSLTFTSNLHHELTMSMDQSVDDVIAEEGFTNLLLKTNSED 308
VKEEG +QWKGY ESS+TFTS H M+ V D + EEGFTNLLLKTNS+D
Sbjct: 9 VKEEG------EQWKGYHESSMTFTSGFHE----FMEGVVGDDVVEEGFTNLLLKTNSDD 158
Query: 309 LSLSESGGESNNGGDGGSGSGSEFYE-DNNNYWNNILNLVNSSPSDSPMF 357
SE GGESNNGG GSGS+FYE DNNNYW++ILNLVNSSPSDSPMF
Sbjct: 159 ---SEGGGESNNGG---RGSGSDFYEADNNNYWSSILNLVNSSPSDSPMF 290
>TC9457 homologue to UP|Q9FZ15 (Q9FZ15) Tuber-specific and
sucrose-responsive element binding factor, partial (43%)
Length = 860
Score = 112 bits (279), Expect = 1e-25
Identities = 51/109 (46%), Positives = 74/109 (67%)
Frame = +3
Query: 14 KGPWTPEEDQKLLAYIDEHGHGSWRALPAKAGLQRCGKSCRLRWTNYLRPDIKRGKFSLQ 73
KGPW+PEED+ L +++HG +W +L +K+ R GKSCRLRW N L P ++ F+ +
Sbjct: 78 KGPWSPEEDEALQRLVEKHGPRNW-SLISKSIPGRSGKSCRLRWCNQLSPQVEHRAFTAE 254
Query: 74 EEQTIIQLHALLGNRWSAIATHLAKRTDNEIKNYWNTHLKKRLTKMGID 122
E+ TII+ HA GN+W+ IA L+ RTDN IKN+WN+ LK++ +D
Sbjct: 255 EDDTIIRAHARFGNKWATIARLLSGRTDNAIKNHWNSTLKRKCASFMMD 401
>TC17948 similar to UP|Q9FZ13 (Q9FZ13) Tuber-specific and sucrose-responsive
element binding factor, partial (50%)
Length = 487
Score = 110 bits (275), Expect = 4e-25
Identities = 52/103 (50%), Positives = 71/103 (68%)
Frame = +2
Query: 14 KGPWTPEEDQKLLAYIDEHGHGSWRALPAKAGLQRCGKSCRLRWTNYLRPDIKRGKFSLQ 73
KGPW+ EED+ L ++ HG +W +L ++ R GKSCRLRW N L P ++ FS+Q
Sbjct: 170 KGPWSAEEDRILTRLVERHGARNW-SLISRYIKGRSGKSCRLRWCNQLSPTVEHRPFSVQ 346
Query: 74 EEQTIIQLHALLGNRWSAIATHLAKRTDNEIKNYWNTHLKKRL 116
E++TII HA GNRW+AIA L RTDN +KN+WN+ LK+R+
Sbjct: 347 EDETIIAAHARFGNRWAAIARLLPGRTDNAVKNHWNSTLKRRV 475
>TC8520 homologue to UP|Q9FZ15 (Q9FZ15) Tuber-specific and
sucrose-responsive element binding factor, partial (36%)
Length = 1778
Score = 110 bits (274), Expect = 5e-25
Identities = 50/106 (47%), Positives = 73/106 (68%)
Frame = +1
Query: 14 KGPWTPEEDQKLLAYIDEHGHGSWRALPAKAGLQRCGKSCRLRWTNYLRPDIKRGKFSLQ 73
KGPW+PEED+ L ++ HG +W +L +K+ R GKSCRLRW N L P ++ F+ +
Sbjct: 184 KGPWSPEEDEALQKLVERHGPRNW-SLISKSIPGRSGKSCRLRWCNQLSPQVEHRAFTPE 360
Query: 74 EEQTIIQLHALLGNRWSAIATHLAKRTDNEIKNYWNTHLKKRLTKM 119
E+ TII+ HA GN+W+ IA L+ RTDN IKN+WN+ LK++ + +
Sbjct: 361 EDDTIIRAHARFGNKWATIARLLSGRTDNAIKNHWNSTLKRKCSSI 498
>TC18881 similar to UP|Q9FZ15 (Q9FZ15) Tuber-specific and sucrose-responsive
element binding factor, partial (28%)
Length = 346
Score = 104 bits (259), Expect = 3e-23
Identities = 47/97 (48%), Positives = 66/97 (67%)
Frame = +1
Query: 14 KGPWTPEEDQKLLAYIDEHGHGSWRALPAKAGLQRCGKSCRLRWTNYLRPDIKRGKFSLQ 73
KGPW+PEED+ L + HG +W A+ +K+ R GKSCRLRW N L P+++ F+ +
Sbjct: 55 KGPWSPEEDEALTRLVQVHGPKNWTAI-SKSIPGRSGKSCRLRWCNQLSPEVEHRPFTPE 231
Query: 74 EEQTIIQLHALLGNRWSAIATHLAKRTDNEIKNYWNT 110
E+ TII+ HA GN+W+ IA L RTDN +KN+WN+
Sbjct: 232 EDSTIIRAHARCGNKWATIARILNGRTDNAVKNHWNS 342
>AV776018
Length = 476
Score = 86.3 bits (212), Expect = 7e-18
Identities = 45/93 (48%), Positives = 60/93 (64%), Gaps = 1/93 (1%)
Frame = +1
Query: 70 FSLQEEQTIIQLHALLGNRWSAIATHLAKRTDNEIKNYWNTHLKKRLTKMGIDPVTHKPK 129
F +E+ II+LHA+LGNRWS AT RTDNEIKN WN+ LKK+L + GIDPVTHKP
Sbjct: 13 FHKEEDNLIIELHAVLGNRWSQDATQWPGRTDNEIKNLWNSCLKKKLRQRGIDPVTHKPL 192
Query: 130 NDALLSSSD-NAHSKTASNLSHMAQWESARLEA 161
++ D N + SN ++ + ES + +A
Sbjct: 193 SEVENGKEDKNKSQEKVSNELNLLKSESFKSDA 291
>BI420780
Length = 475
Score = 71.2 bits (173), Expect = 2e-13
Identities = 34/86 (39%), Positives = 49/86 (56%), Gaps = 2/86 (2%)
Frame = +1
Query: 17 WTPEEDQKLLAYIDEHGHGSWRALPAKAG--LQRCGKSCRLRWTNYLRPDIKRGKFSLQE 74
W+ EED L AY+ ++G W + + L R KSC RW NYL+P IK+G + +E
Sbjct: 211 WSSEEDALLHAYVQQYGPREWNLVSQRMNTPLNRDTKSCLERWKNYLKPGIKKGSLTKEE 390
Query: 75 EQTIIQLHALLGNRWSAIATHLAKRT 100
++ +I L A GN+W IA + RT
Sbjct: 391 QRLVILLQANYGNKWKKIAAEVPGRT 468
Database: LJGI
Posted date: Jul 30, 2004 11:16 AM
Number of letters in database: 14,692,800
Number of sequences in database: 28,460
Lambda K H
0.309 0.128 0.377
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 6,849,412
Number of Sequences: 28460
Number of extensions: 96409
Number of successful extensions: 756
Number of sequences better than 10.0: 86
Number of HSP's better than 10.0 without gapping: 684
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 708
length of query: 357
length of database: 4,897,600
effective HSP length: 91
effective length of query: 266
effective length of database: 2,307,740
effective search space: 613858840
effective search space used: 613858840
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 16 ( 7.1 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 42 (21.7 bits)
S2: 56 (26.2 bits)
Medicago: description of AC122724.4


