

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
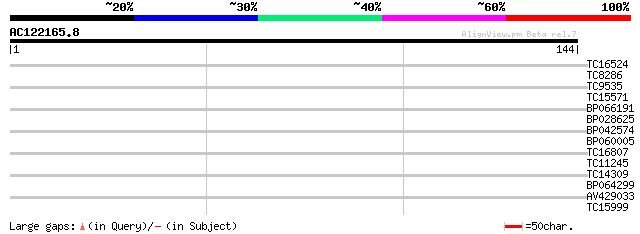
Query= AC122165.8 - phase: 1 /pseudo
(144 letters)
Database: LJGI
28,460 sequences; 14,692,800 total letters
Searching..................................................done
Score E
Sequences producing significant alignments: (bits) Value
TC16524 similar to GB|AAL69537.1|18491137|AY074839 AT5g45420/MFC... 30 0.16
TC8286 homologue to GB|AAD14482.1|4249385|T2K10 T2K10.11 {Arabid... 28 0.61
TC9535 28 0.80
TC15571 weakly similar to PIR|S19652|S19652 cellodextrinase C - ... 26 2.3
BP066191 25 4.0
BP028625 25 5.2
BP042574 25 5.2
BP060005 25 5.2
TC16807 similar to UP|Q9LPN1 (Q9LPN1) F2J10.2 protein, partial (... 25 5.2
TC11245 similar to UP|Q8KBE2 (Q8KBE2) Naphthoate synthase, parti... 25 6.8
TC14309 homologue to UP|H2B_GOSHI (O22582) Histone H2B, partial ... 24 8.8
BP064299 24 8.8
AV429033 24 8.8
TC15999 similar to UP|Q9LLT5 (Q9LLT5) Receptor-like protein kina... 24 8.8
>TC16524 similar to GB|AAL69537.1|18491137|AY074839 AT5g45420/MFC19_9
{Arabidopsis thaliana;}, partial (41%)
Length = 997
Score = 30.0 bits (66), Expect = 0.16
Identities = 18/51 (35%), Positives = 26/51 (50%)
Frame = +1
Query: 81 SNRKQRFKQKLKEADKRISGTGRENKVDNLRELVGGEKTSVGMAKGSSPKD 131
S+ ++F +K K ADKR G E KV+ GE ++ A + PKD
Sbjct: 538 SDSYEQFLKKRKAADKRGGGEEEEGKVEEETNWSNGEDIALLNALKAFPKD 690
>TC8286 homologue to GB|AAD14482.1|4249385|T2K10 T2K10.11 {Arabidopsis
thaliana;}, partial (6%)
Length = 671
Score = 28.1 bits (61), Expect = 0.61
Identities = 10/20 (50%), Positives = 14/20 (70%)
Frame = +2
Query: 65 PHGAPPSQPKQQEVIPSNRK 84
PH PP+QP Q V+PS ++
Sbjct: 146 PHLPPPTQPPQSHVLPSQQR 205
>TC9535
Length = 1083
Score = 27.7 bits (60), Expect = 0.80
Identities = 18/54 (33%), Positives = 24/54 (44%)
Frame = -3
Query: 29 PSLGIQDSDQSKNNAASVESPNSAAKTKKEEKIYLGPHGAPPSQPKQQEVIPSN 82
P+ GI + SK + SV +S TK Y +PPS P +PSN
Sbjct: 520 PTTGIPEDMASKTTSPSVS--DSEGMTKTSPLAYA*DSSSPPSIPVNTAGMPSN 365
>TC15571 weakly similar to PIR|S19652|S19652 cellodextrinase C - Pseudomonas
fluorescens {Pseudomonas fluorescens;} , partial (6%)
Length = 784
Score = 26.2 bits (56), Expect = 2.3
Identities = 20/72 (27%), Positives = 28/72 (38%), Gaps = 9/72 (12%)
Frame = -2
Query: 34 QDSDQSKNNAASV-----ESPNSAAKTKKEEKIYLGPH----GAPPSQPKQQEVIPSNRK 84
QD D + S E +S+ EE L H G P +QP +++ IP
Sbjct: 756 QDDDDDEEETDSFDQGDEEETDSSLSHSDEEIALLDTHASAEGVPVAQPAEEQPIPPPPT 577
Query: 85 QRFKQKLKEADK 96
Q L+E K
Sbjct: 576 TSMSQLLQELQK 541
>BP066191
Length = 518
Score = 25.4 bits (54), Expect = 4.0
Identities = 12/32 (37%), Positives = 16/32 (49%)
Frame = -3
Query: 65 PHGAPPSQPKQQEVIPSNRKQRFKQKLKEADK 96
P PP Q Q +I N +Q+ + LKE K
Sbjct: 411 PFSKPPEQEFSQSIISKNNQQKPCKSLKEECK 316
>BP028625
Length = 566
Score = 25.0 bits (53), Expect = 5.2
Identities = 8/11 (72%), Positives = 9/11 (81%)
Frame = -2
Query: 132 WLDPHCHEAEF 142
WLDPHC+ EF
Sbjct: 142 WLDPHCNFLEF 110
>BP042574
Length = 505
Score = 25.0 bits (53), Expect = 5.2
Identities = 12/34 (35%), Positives = 18/34 (52%)
Frame = +1
Query: 49 PNSAAKTKKEEKIYLGPHGAPPSQPKQQEVIPSN 82
PN A ++ E ++ P P +P+ E IPSN
Sbjct: 19 PNQRACFRRIESLHSLPPSHFPCKPRDPEPIPSN 120
>BP060005
Length = 339
Score = 25.0 bits (53), Expect = 5.2
Identities = 19/68 (27%), Positives = 27/68 (38%), Gaps = 8/68 (11%)
Frame = +1
Query: 56 KKEEKIYLGPHGAPPSQPKQQ--------EVIPSNRKQRFKQKLKEADKRISGTGRENKV 107
+ E + + HGAPP+ P + + S RK + KQ G N +
Sbjct: 76 QSSELVIVSAHGAPPTHPNRTLYFGDNILHISSSGRKFKHKQ----------GGSNNN*L 225
Query: 108 DNLRELVG 115
LR LVG
Sbjct: 226 SILRGLVG 249
>TC16807 similar to UP|Q9LPN1 (Q9LPN1) F2J10.2 protein, partial (13%)
Length = 486
Score = 25.0 bits (53), Expect = 5.2
Identities = 14/62 (22%), Positives = 25/62 (39%)
Frame = -3
Query: 29 PSLGIQDSDQSKNNAASVESPNSAAKTKKEEKIYLGPHGAPPSQPKQQEVIPSNRKQRFK 88
P LG +N +AS + + + + PP P ++ SNR+Q +
Sbjct: 259 PGLGGASPGSDRNTSASSHHF*TLSSISRRRSLRAKLFPPPPPPPSVAAMLVSNRRQLYM 80
Query: 89 QK 90
+K
Sbjct: 79 EK 74
>TC11245 similar to UP|Q8KBE2 (Q8KBE2) Naphthoate synthase, partial (23%)
Length = 488
Score = 24.6 bits (52), Expect = 6.8
Identities = 24/100 (24%), Positives = 40/100 (40%)
Frame = -2
Query: 8 FVLKCMFFGLLLVVDTDGFVIPSLGIQDSDQSKNNAASVESPNSAAKTKKEEKIYLGPHG 67
F+++ FG +V T + SL ++ D ++ + + SA K +GP G
Sbjct: 484 FIIRISIFGPQGLVTTTTKCLGSLPGENDDPNRGIVSCIIE--SADKLLNR----MGPKG 323
Query: 68 APPSQPKQQEVIPSNRKQRFKQKLKEADKRISGTGRENKV 107
P P + S Q + E R+ G+G N V
Sbjct: 322 IPSLWPINSNLGYSFPNSLLVQNVSEVLARVIGSGTPNDV 203
>TC14309 homologue to UP|H2B_GOSHI (O22582) Histone H2B, partial (93%)
Length = 682
Score = 24.3 bits (51), Expect = 8.8
Identities = 16/56 (28%), Positives = 24/56 (42%), Gaps = 2/56 (3%)
Frame = +1
Query: 44 ASVESPNSAAKTKKEEKIYLGPHGAPPSQ--PKQQEVIPSNRKQRFKQKLKEADKR 97
A+ E +A K EEK APP++ PK + +P K K+ K+
Sbjct: 70 AAAEKKPAAEKKPAEEKKSTAAEKAPPAEKKPKAGKKLPKEGASAAGDKKKKRSKK 237
>BP064299
Length = 531
Score = 24.3 bits (51), Expect = 8.8
Identities = 14/62 (22%), Positives = 24/62 (38%)
Frame = -3
Query: 29 PSLGIQDSDQSKNNAASVESPNSAAKTKKEEKIYLGPHGAPPSQPKQQEVIPSNRKQRFK 88
P LG +N +AS + + + PP P ++ SNR+Q +
Sbjct: 304 PGLGGASPGSDRNTSASSHHF*TLRSISRRRSLRAKLFPPPPPPPSVAAMLVSNRRQLYM 125
Query: 89 QK 90
+K
Sbjct: 124 EK 119
>AV429033
Length = 557
Score = 24.3 bits (51), Expect = 8.8
Identities = 13/46 (28%), Positives = 21/46 (45%)
Frame = +1
Query: 29 PSLGIQDSDQSKNNAASVESPNSAAKTKKEEKIYLGPHGAPPSQPK 74
PSLG + +S+ + SP +++ PHG PP P+
Sbjct: 151 PSLGRHNRCRSRRRQRLLPSP---------PRLHRRPHGPPPPHPQ 261
>TC15999 similar to UP|Q9LLT5 (Q9LLT5) Receptor-like protein kinase
(Fragment), complete
Length = 999
Score = 24.3 bits (51), Expect = 8.8
Identities = 8/16 (50%), Positives = 9/16 (56%)
Frame = +3
Query: 122 GMAKGSSPKDWLDPHC 137
G G P+DWL HC
Sbjct: 663 G*TNGRGPQDWLSVHC 710
Database: LJGI
Posted date: Jul 30, 2004 11:16 AM
Number of letters in database: 14,692,800
Number of sequences in database: 28,460
Lambda K H
0.320 0.137 0.406
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 2,405,100
Number of Sequences: 28460
Number of extensions: 24819
Number of successful extensions: 176
Number of sequences better than 10.0: 28
Number of HSP's better than 10.0 without gapping: 175
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 176
length of query: 144
length of database: 4,897,600
effective HSP length: 82
effective length of query: 62
effective length of database: 2,563,880
effective search space: 158960560
effective search space used: 158960560
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 16 ( 7.4 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.8 bits)
S2: 51 (24.3 bits)
Medicago: description of AC122165.8


