

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
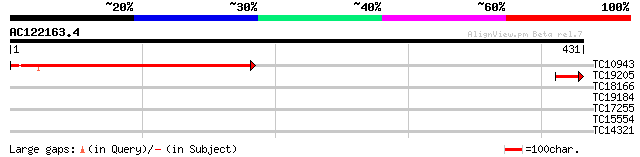
Query= AC122163.4 - phase: 0
(431 letters)
Database: LJGI
28,460 sequences; 14,692,800 total letters
Searching..................................................done
Score E
Sequences producing significant alignments: (bits) Value
TC10943 similar to GB|AAM98095.1|22655008|AY139777 AT3g20300/MQC... 279 6e-76
TC19205 45 2e-05
TC18166 33 0.091
TC19184 weakly similar to UP|Q9FXH4 (Q9FXH4) F6F9.19 protein, pa... 29 1.7
TC17255 28 3.8
TC15554 similar to GB|AAO24533.1|27808506|BT003101 At2g41420 {Ar... 27 6.5
TC14321 similar to UP|AAQ72789 (AAQ72789) 60S ribosomal protein ... 27 8.5
>TC10943 similar to GB|AAM98095.1|22655008|AY139777 AT3g20300/MQC12_5
{Arabidopsis thaliana;}, partial (5%)
Length = 710
Score = 279 bits (714), Expect = 6e-76
Identities = 135/187 (72%), Positives = 153/187 (81%), Gaps = 2/187 (1%)
Frame = +2
Query: 1 MEEEREVSFLAMNKENSGIY--HDGKELRRFRAYLRWVYVDQSNLCKAGISWSVFFTLAY 58
ME++R + L +NKE + Y +D ELR+ R+Y+RWVYVDQSN+CK + SVFFTLA+
Sbjct: 152 MEDDRGI-LLQVNKEENASYLLNDEAELRKIRSYIRWVYVDQSNVCKTVFTCSVFFTLAF 328
Query: 59 VVPILSHFLLDCSTTCDADHRRPYHVPVQISLSVFATLSFISLSKWDRKYGFSKFLFLDK 118
VVPILSHFLL C +TCD DH RPYH+PVQISL+VFATLSFISLS WDRKYG SKFLFLDK
Sbjct: 329 VVPILSHFLLYCPSTCDPDHSRPYHIPVQISLTVFATLSFISLSSWDRKYGLSKFLFLDK 508
Query: 119 VSDESLKIQRGYAIQMKRTMKLILLWGLPCFICQCVYKIWWYISGASQIPYYGEVYVSGI 178
V DESL+IQRGYA QMK TM LIL WGLPCFI +C YKIWWY SG SQIPYYGE+Y S I
Sbjct: 509 VGDESLRIQRGYAQQMKGTMNLILRWGLPCFIAECAYKIWWYASGTSQIPYYGEIYASSI 688
Query: 179 ILCTLEL 185
IL TLEL
Sbjct: 689 ILGTLEL 709
>TC19205
Length = 523
Score = 45.1 bits (105), Expect = 2e-05
Identities = 20/21 (95%), Positives = 20/21 (95%)
Frame = +1
Query: 411 HSIFGIQLALCLWLLNKTVGI 431
HSIF IQLALCLWLLNKTVGI
Sbjct: 1 HSIFAIQLALCLWLLNKTVGI 63
>TC18166
Length = 530
Score = 33.1 bits (74), Expect = 0.091
Identities = 18/46 (39%), Positives = 28/46 (60%)
Frame = -1
Query: 223 ETEVGTILLEHLRIRRNLRVISHRFRAFILSSLLLVTASQLIFLLM 268
ET + T+L++HLRI++ + H FR L LLL+ Q++ L M
Sbjct: 347 ETNLLTLLIKHLRIKKMKENM*HLFRTPSLHGLLLIWLKQMVGLDM 210
>TC19184 weakly similar to UP|Q9FXH4 (Q9FXH4) F6F9.19 protein, partial (19%)
Length = 627
Score = 28.9 bits (63), Expect = 1.7
Identities = 17/50 (34%), Positives = 27/50 (54%)
Frame = -1
Query: 313 TGLASKWHICATINSFDNIDGETPTALIASAQAMAADISWGSSEDESGDE 362
T + KW C T D+I G+T ++S+ A + IS S+ D +GD+
Sbjct: 369 TNI*HKWQRCQTRKRCDSIKGKT*LLQVSSSSAAPSSIS--SAIDLAGDQ 226
>TC17255
Length = 485
Score = 27.7 bits (60), Expect = 3.8
Identities = 16/68 (23%), Positives = 32/68 (46%), Gaps = 3/68 (4%)
Frame = +2
Query: 138 MKLILLWGLPC---FICQCVYKIWWYISGASQIPYYGEVYVSGIILCTLELCSWFYRTSI 194
+K+ ++W L F C + + Y+S + ++G V + + L L F+ I
Sbjct: 254 LKVKIVW*LSH*VPFCCNVRWHVITYLSWFVLVFFFGIV*IFSCVFSFLNLSIAFFGIDI 433
Query: 195 FFLVCVLF 202
F +C++F
Sbjct: 434 IFHICIVF 457
>TC15554 similar to GB|AAO24533.1|27808506|BT003101 At2g41420 {Arabidopsis
thaliana;}, partial (77%)
Length = 679
Score = 26.9 bits (58), Expect = 6.5
Identities = 20/85 (23%), Positives = 34/85 (39%), Gaps = 4/85 (4%)
Frame = -2
Query: 141 ILLWGLPCFICQCVYKIWWYISGASQIPYYGEVYVSGIILCTLELCSWFYRTSIFFLVCV 200
I LW +PC+ WW +S IP + +W+ + +V
Sbjct: 216 IPLWRVPCWRIPLCRVPWWRVSVLRWIPLW------------RRNTNWWL---LLMIVVA 82
Query: 201 LFRLIGYLQ----IQRLDEFAPVFQ 221
L+GYL+ +Q+L F +F+
Sbjct: 81 HVNLLGYLKCCAFVQKLQRFRKLFE 7
>TC14321 similar to UP|AAQ72789 (AAQ72789) 60S ribosomal protein L5, partial
(98%)
Length = 1185
Score = 26.6 bits (57), Expect = 8.5
Identities = 28/98 (28%), Positives = 47/98 (47%), Gaps = 5/98 (5%)
Frame = +1
Query: 208 LQIQRLDEFAPVFQRETEVGTILLEHLRIRRNLRVISHRFRAFILSSLLLVTASQLIFLL 267
LQ+ LD + PV L+HL+ RN+R + I +LL ++ L+
Sbjct: 325 LQLIALDSYWPVES---------LKHLKWMRNMRAMLRLLERTIQWNLLRPGGHSVLSLM 477
Query: 268 MVI-KPH-ADVDILKGGELGLV---SITLVSGLYILLR 300
+V+ +P A V + EL +V +T+ GL +L+R
Sbjct: 478 LVLSRPQLATVSSVPLREL*MVVWIFLTVTRGLLVLIR 591
Database: LJGI
Posted date: Jul 30, 2004 11:16 AM
Number of letters in database: 14,692,800
Number of sequences in database: 28,460
Lambda K H
0.327 0.140 0.434
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 8,488,219
Number of Sequences: 28460
Number of extensions: 133872
Number of successful extensions: 1091
Number of sequences better than 10.0: 14
Number of HSP's better than 10.0 without gapping: 1082
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 1091
length of query: 431
length of database: 4,897,600
effective HSP length: 93
effective length of query: 338
effective length of database: 2,250,820
effective search space: 760777160
effective search space used: 760777160
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 15 ( 7.1 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 40 (21.7 bits)
S2: 57 (26.6 bits)
Medicago: description of AC122163.4


