

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
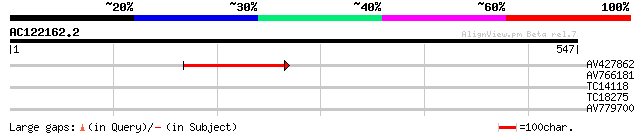
Query= AC122162.2 - phase: 0 /pseudo
(547 letters)
Database: LJGI
28,460 sequences; 14,692,800 total letters
Searching..................................................done
Score E
Sequences producing significant alignments: (bits) Value
AV427862 170 6e-43
AV766181 40 0.001
TC14118 homologue to UP|AAS18240 (AAS18240) Enolase , complete 32 0.26
TC18275 28 2.9
AV779700 28 5.0
>AV427862
Length = 314
Score = 170 bits (430), Expect = 6e-43
Identities = 83/103 (80%), Positives = 88/103 (84%)
Frame = +3
Query: 168 KQLQPVNFDELFASLYNDYTQKQEDKIWCIIHYFQRITSNMPKGVVSFERKVLPWEDDCI 227
K LQ +NFDELF SLY+ Y+QKQE KI CIIHYFQRITSNMP+GVVSFERKVLP EDD I
Sbjct: 6 KHLQSINFDELFGSLYDYYSQKQESKIQCIIHYFQRITSNMPQGVVSFERKVLPLEDDYI 185
Query: 228 HISYPNASFWSTSVKPLCRFEVKSSGLIEDHSSETVEVDFANE 270
HISYPNA WSTS+ PLCRFEV SSGLIED SE VEVDFANE
Sbjct: 186 HISYPNADVWSTSIIPLCRFEVHSSGLIEDQLSEAVEVDFANE 314
>AV766181
Length = 410
Score = 40.0 bits (92), Expect = 0.001
Identities = 18/24 (75%), Positives = 19/24 (79%)
Frame = -3
Query: 524 GLVLDFVAGMDSWRPMEHASGVLN 547
G LDFV MDSWRPMEHA+ VLN
Sbjct: 408 GCSLDFVTQMDSWRPMEHAN*VLN 337
>TC14118 homologue to UP|AAS18240 (AAS18240) Enolase , complete
Length = 1696
Score = 32.0 bits (71), Expect = 0.26
Identities = 22/76 (28%), Positives = 38/76 (49%), Gaps = 6/76 (7%)
Frame = -3
Query: 133 LDSQEAGIVFLTQELIAALLACSFLCLFPVHDRYEKQLQPVNFDELFASLYNDYTQKQED 192
L QE + L + L L +F+C P+ ++ + ++P++ LFA L D Q ++
Sbjct: 476 LSGQEHQLCRLQETLPIWHLLQAFVCTNPIR*QFHQAVEPLSCQLLFAEL--DPGQSKQG 303
Query: 193 KIWC------IIHYFQ 202
+ WC +IH FQ
Sbjct: 302 Q*WCSHC*QPLIHPFQ 255
>TC18275
Length = 800
Score = 28.5 bits (62), Expect = 2.9
Identities = 12/34 (35%), Positives = 18/34 (52%)
Frame = -1
Query: 151 LLACSFLCLFPVHDRYEKQLQPVNFDELFASLYN 184
L LC F DRY++ L P++F++ S N
Sbjct: 278 LFQMEILCSFNEEDRYQRDLLPLSFEDQTCSCLN 177
>AV779700
Length = 438
Score = 27.7 bits (60), Expect = 5.0
Identities = 16/52 (30%), Positives = 26/52 (49%)
Frame = +1
Query: 164 DRYEKQLQPVNFDELFASLYNDYTQKQEDKIWCIIHYFQRITSNMPKGVVSF 215
D+Y L VN E + + + KIWCI+H+ + T+ K ++SF
Sbjct: 1 DKYNISLHIVNSLERIFIPISMGEKNKHPKIWCIVHHEK*NTA*CAKCILSF 156
Database: LJGI
Posted date: Jul 30, 2004 11:16 AM
Number of letters in database: 14,692,800
Number of sequences in database: 28,460
Lambda K H
0.324 0.140 0.426
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 9,439,173
Number of Sequences: 28460
Number of extensions: 127012
Number of successful extensions: 700
Number of sequences better than 10.0: 10
Number of HSP's better than 10.0 without gapping: 699
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 700
length of query: 547
length of database: 4,897,600
effective HSP length: 95
effective length of query: 452
effective length of database: 2,193,900
effective search space: 991642800
effective search space used: 991642800
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 15 ( 7.0 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 40 (21.6 bits)
S2: 57 (26.6 bits)
Medicago: description of AC122162.2


