

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC122162.15 - phase: 0
(342 letters)
Database: LJGI
28,460 sequences; 14,692,800 total letters
Searching..................................................done
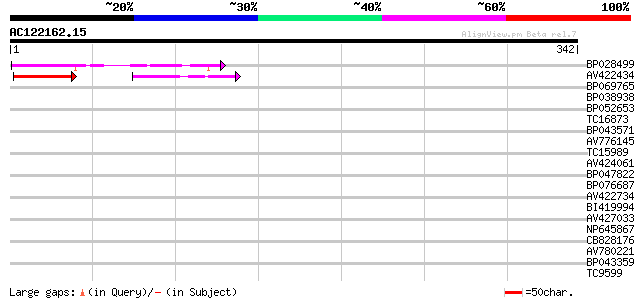
Score E
Sequences producing significant alignments: (bits) Value
BP028499 68 2e-12
AV422434 56 4e-11
BP069765 37 0.006
BP038938 33 0.090
BP052653 30 0.45
TC16873 30 0.45
BP043571 30 0.77
AV776145 30 0.77
TC15989 29 1.3
AV424061 29 1.3
BP047822 28 1.7
BP076687 28 2.2
AV422734 28 2.2
BI419994 28 2.9
AV427033 27 3.8
NP645867 proliferating floral organs protein [Lotus japonicus] 27 3.8
CB828176 27 5.0
AV780221 27 5.0
BP043359 27 5.0
TC9599 27 5.0
>BP028499
Length = 491
Score = 68.2 bits (165), Expect = 2e-12
Identities = 51/137 (37%), Positives = 71/137 (51%), Gaps = 8/137 (5%)
Frame = +2
Query: 2 ASVDWTELPRDILFLISQRLDVELDLIRFRSVCSTWH----SSPIPNDHNILPFQFPLLK 57
A DW++LP ++L LISQRL+ L L+RFRSVCSTW SS IP H PF FP L
Sbjct: 107 APPDWSQLPAELLHLISQRLNSPLYLLRFRSVCSTWRHSSASSFIPIHH--FPFNFPPLS 280
Query: 58 YVPAPDSINNNNEIIDNINNANTSFCYLSKRSFFLVKPPQEQQQQETLIRCRPWLIQITK 117
D+ ++ +SF L+KR+ L+ PP Q PWL++I +
Sbjct: 281 ---------------DHDHHDASSFS-LTKRTVVLITPPPPPHHQTQ----TPWLVKIGE 400
Query: 118 N----SSGKTQLLNPPF 130
+ +T+L +P F
Sbjct: 401 DLCDGDRTRTRLWHPLF 451
>AV422434
Length = 422
Score = 56.2 bits (134), Expect(2) = 4e-11
Identities = 25/38 (65%), Positives = 31/38 (80%)
Frame = +3
Query: 3 SVDWTELPRDILFLISQRLDVELDLIRFRSVCSTWHSS 40
+VDW++LP ++L LISQRLD L L+RFRSVCSTW S
Sbjct: 51 AVDWSQLPTELLHLISQRLDSPLYLLRFRSVCSTWRRS 164
Score = 27.3 bits (59), Expect(2) = 4e-11
Identities = 20/65 (30%), Positives = 34/65 (51%)
Frame = +1
Query: 75 INNANTSFCYLSKRSFFLVKPPQEQQQQETLIRCRPWLIQITKNSSGKTQLLNPPFVSLK 134
+++ S +LS+R+ FL+ PP Q + P L QI+++ +G T+ L P S +
Sbjct: 205 LSDDTASTFFLSQRTVFLIIPPPLQPTEP----LTPLLFQISQD-NG*TRRLYDPLSSHR 369
Query: 135 STAFS 139
FS
Sbjct: 370 DPPFS 384
>BP069765
Length = 417
Score = 36.6 bits (83), Expect = 0.006
Identities = 14/34 (41%), Positives = 23/34 (67%)
Frame = -2
Query: 303 KKWEKLKKLGDRILFVGSGCSFSASALDLCLPKG 336
+ W ++ + DR+LF+G C+FS SA DL + +G
Sbjct: 416 ESWVHVRNVRDRVLFIGDDCAFSVSASDLGVDEG 315
>BP038938
Length = 583
Score = 32.7 bits (73), Expect = 0.090
Identities = 14/34 (41%), Positives = 22/34 (64%)
Frame = +3
Query: 6 WTELPRDILFLISQRLDVELDLIRFRSVCSTWHS 39
W++LP ++L L+ RL ++ D +R VC WHS
Sbjct: 135 WSDLPAELLELVMTRLALD-DNVRASVVCKRWHS 233
>BP052653
Length = 552
Score = 30.4 bits (67), Expect = 0.45
Identities = 15/31 (48%), Positives = 20/31 (64%)
Frame = +1
Query: 9 LPRDILFLISQRLDVELDLIRFRSVCSTWHS 39
LP D++ I RL V L+RF+SVC +W S
Sbjct: 187 LPHDLIVQILLRLPVR-SLLRFKSVCKSWLS 276
>TC16873
Length = 1080
Score = 30.4 bits (67), Expect = 0.45
Identities = 29/104 (27%), Positives = 46/104 (43%), Gaps = 10/104 (9%)
Frame = -3
Query: 43 PNDHNILPFQFPLLKYVPAPDSINNNNEIIDNINNANTSFCYLSKRSFF----LVKPPQE 98
P ++ ILP F +L Y P I N + +NN+NT F +L + ++ P+
Sbjct: 418 PLENEILPPPFRVLHYSGIPIPIFFRNITMCQLNNSNTRFRFLKHKPHRGNNPIIPAPKS 239
Query: 99 QQQQETLIRCRPWLIQIT------KNSSGKTQLLNPPFVSLKST 136
L R +QI+ KN+ T LL P ++L+ T
Sbjct: 238 PSLHNQL---RFLQLQISPCHRPIKNAEIPTNLLINPRIALRET 116
>BP043571
Length = 438
Score = 29.6 bits (65), Expect = 0.77
Identities = 13/42 (30%), Positives = 22/42 (51%)
Frame = +1
Query: 275 SEGELLLLDIHQTFFQFSMKIFKLDENRKKWEKLKKLGDRIL 316
S G +L +++ F +K FK+ + R W+ KLG + L
Sbjct: 85 SLGRILTCTLNKKIFTVEVKNFKMSKTRNSWKDHSKLGGKSL 210
>AV776145
Length = 275
Score = 29.6 bits (65), Expect = 0.77
Identities = 12/43 (27%), Positives = 25/43 (57%)
Frame = -3
Query: 53 FPLLKYVPAPDSINNNNEIIDNINNANTSFCYLSKRSFFLVKP 95
FP+L+Y PA ++N+ +I+ + + FC +F+++P
Sbjct: 258 FPILRYHPATFRESDNSHLINPLPHTTFFFCRYIYPLYFIIEP 130
>TC15989
Length = 526
Score = 28.9 bits (63), Expect = 1.3
Identities = 14/42 (33%), Positives = 24/42 (56%)
Frame = -3
Query: 65 INNNNEIIDNINNANTSFCYLSKRSFFLVKPPQEQQQQETLI 106
IN+ ++ +I N CY+ K S F + Q+QQQQ++ +
Sbjct: 494 INSVSQYTMSIKNNK*KSCYILKNS*FTIIQQQQQQQQKSYL 369
>AV424061
Length = 273
Score = 28.9 bits (63), Expect = 1.3
Identities = 10/37 (27%), Positives = 23/37 (62%)
Frame = +1
Query: 54 PLLKYVPAPDSINNNNEIIDNINNANTSFCYLSKRSF 90
P+LKY+ P + ++ + +I+N+N + ++S + F
Sbjct: 46 PILKYIQHPSTSSSRSITFSSISNSNPYYLFISPKCF 156
>BP047822
Length = 533
Score = 28.5 bits (62), Expect = 1.7
Identities = 23/67 (34%), Positives = 33/67 (48%)
Frame = -1
Query: 267 KNRKLLVESEGELLLLDIHQTFFQFSMKIFKLDENRKKWEKLKKLGDRILFVGSGCSFSA 326
+ KL + +E L + Q F + ++ K ENRK W+ L R + V CSF +
Sbjct: 392 RQEKLQLLNERTEELQNGAQDFASMATELAKRMENRKWWQ----L*QRCMIVVLLCSF*S 225
Query: 327 SALDLCL 333
SAL CL
Sbjct: 224 SALVNCL 204
>BP076687
Length = 389
Score = 28.1 bits (61), Expect = 2.2
Identities = 14/33 (42%), Positives = 21/33 (63%)
Frame = -3
Query: 7 TELPRDILFLISQRLDVELDLIRFRSVCSTWHS 39
+ LP +++ I RL V L+RF+SVC +W S
Sbjct: 387 SHLPEELIPQILLRLPVR-SLLRFKSVCKSWLS 292
>AV422734
Length = 448
Score = 28.1 bits (61), Expect = 2.2
Identities = 25/93 (26%), Positives = 46/93 (48%)
Frame = +1
Query: 15 FLISQRLDVELDLIRFRSVCSTWHSSPIPNDHNILPFQFPLLKYVPAPDSINNNNEIIDN 74
FL+ Q+L + + + S + SS D ++ P + L+ P P N NN I++
Sbjct: 178 FLLLQQLQKQQQQQQQAAAISRFPSSI---DAHLRPIRPINLQQNPNP---NPNNPILNL 339
Query: 75 INNANTSFCYLSKRSFFLVKPPQEQQQQETLIR 107
N N++ ++ + PQ+QQQQ+ ++R
Sbjct: 340 HQNPNSNHLQQQQQQ----QQPQQQQQQQKMVR 426
>BI419994
Length = 549
Score = 27.7 bits (60), Expect = 2.9
Identities = 12/32 (37%), Positives = 21/32 (65%)
Frame = +2
Query: 6 WTELPRDILFLISQRLDVELDLIRFRSVCSTW 37
++ LP +++ I RL V L+RF++VC +W
Sbjct: 38 FSHLPDELILQILLRLPVR-SLLRFKTVCKSW 130
>AV427033
Length = 420
Score = 27.3 bits (59), Expect = 3.8
Identities = 14/31 (45%), Positives = 20/31 (64%)
Frame = +1
Query: 9 LPRDILFLISQRLDVELDLIRFRSVCSTWHS 39
LP D++ I RL V L++FR VC +W+S
Sbjct: 124 LPFDLVVEILCRLPVN-SLLQFRCVCKSWNS 213
>NP645867 proliferating floral organs protein [Lotus japonicus]
Length = 1350
Score = 27.3 bits (59), Expect = 3.8
Identities = 28/92 (30%), Positives = 38/92 (40%), Gaps = 9/92 (9%)
Frame = +1
Query: 1 MASVDWTELPRDILFLISQRLDVELDLIRFRSVCSTWHSSPIPNDHNILPFQ-------F 53
M S W++LP+ +L + L R RSVC W+S N L F
Sbjct: 160 MNSRIWSKLPQKLLDRVIAFLPTPA-FFRARSVCKRWYSLLFSNTFLELYLHLSPRRHWF 336
Query: 54 PLLKYVPAPDSI--NNNNEIIDNINNANTSFC 83
K+ + I NNNN II +A T+ C
Sbjct: 337 IFFKHKTRKNYIYKNNNNNII--TGSAGTASC 426
>CB828176
Length = 363
Score = 26.9 bits (58), Expect = 5.0
Identities = 13/35 (37%), Positives = 19/35 (54%)
Frame = +3
Query: 295 IFKLDENRKKWEKLKKLGDRILFVGSGCSFSASAL 329
IFKLD+ +WE+++ L LF S S + L
Sbjct: 15 IFKLDQTLMEWEEMRTLDGATLFASFLSSHSRTEL 119
>AV780221
Length = 461
Score = 26.9 bits (58), Expect = 5.0
Identities = 11/29 (37%), Positives = 17/29 (57%)
Frame = -1
Query: 34 CSTWHSSPIPNDHNILPFQFPLLKYVPAP 62
C+ + I N N++P PLLK+ P+P
Sbjct: 329 CTLGNGILILNQQNLIP*SIPLLKHPPSP 243
>BP043359
Length = 479
Score = 26.9 bits (58), Expect = 5.0
Identities = 17/59 (28%), Positives = 28/59 (46%)
Frame = +3
Query: 48 ILPFQFPLLKYVPAPDSINNNNEIIDNINNANTSFCYLSKRSFFLVKPPQEQQQQETLI 106
I FP+L Y + ++N N TS ++SKR+F PP+ +Q ++I
Sbjct: 222 IFKVMFPILSYTGSGYDLSNANPFSIK---PFTSSSWISKRNFGFNFPPKLSKQSRSII 389
>TC9599
Length = 579
Score = 26.9 bits (58), Expect = 5.0
Identities = 19/54 (35%), Positives = 26/54 (47%), Gaps = 1/54 (1%)
Frame = +2
Query: 148 IKFSAQHLRTDFIIDEDDFTFQNQHG-SYLYPQTVLAVTCHGKKPLVLGTLSYC 200
I+ A H + DF +Q QH Y+Y Q +L+V CH L+ G YC
Sbjct: 296 IQLMALH*EISSCLHPADFIYQLQHSRKYIYTQ-LLSVLCH--FILLSGNALYC 448
Database: LJGI
Posted date: Jul 30, 2004 11:16 AM
Number of letters in database: 14,692,800
Number of sequences in database: 28,460
Lambda K H
0.322 0.138 0.426
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 7,124,527
Number of Sequences: 28460
Number of extensions: 109759
Number of successful extensions: 599
Number of sequences better than 10.0: 46
Number of HSP's better than 10.0 without gapping: 595
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 599
length of query: 342
length of database: 4,897,600
effective HSP length: 91
effective length of query: 251
effective length of database: 2,307,740
effective search space: 579242740
effective search space used: 579242740
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 16 ( 7.4 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.9 bits)
S2: 55 (25.8 bits)
Medicago: description of AC122162.15


