

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC121245.9 + phase: 2 /pseudo/partial
(223 letters)
Database: LJGI
28,460 sequences; 14,692,800 total letters
Searching..................................................done
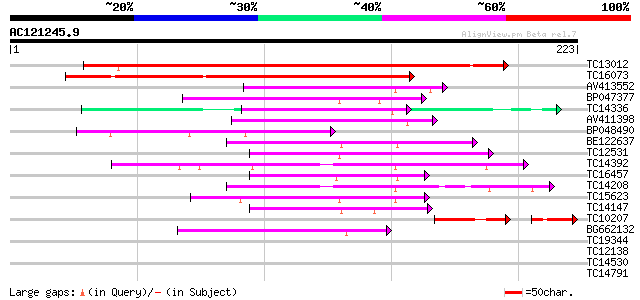
Score E
Sequences producing significant alignments: (bits) Value
TC13012 similar to PIR|T50004|T50004 RNA binding protein-like - ... 261 7e-71
TC16073 similar to UP|Q9FJN9 (Q9FJN9) Poly(A)-binding protein II... 199 4e-52
AV413552 47 3e-06
BP047377 47 3e-06
TC14336 similar to UP|Q9M549 (Q9M549) Poly(A)-binding protein, p... 45 1e-05
AV411398 44 2e-05
BP048490 44 3e-05
BE122637 43 4e-05
TC12531 similar to UP|Q9ASQ8 (Q9ASQ8) At1g22910/F19G10_13, parti... 43 5e-05
TC14392 similar to UP|Q9LEB4 (Q9LEB4) RNA binding protein 45, pa... 42 6e-05
TC16457 similar to UP|Q41124 (Q41124) Chloroplast RNA binding pr... 42 8e-05
TC14208 similar to UP|Q9LEB4 (Q9LEB4) RNA binding protein 45, pa... 42 8e-05
TC15623 similar to UP|P93486 (P93486) Glycine-rich RNA-binding p... 41 2e-04
TC14147 similar to UP|ROC1_NICSY (Q08935) 29 kDa ribonucleoprote... 40 2e-04
TC10207 homologue to UP|Q8LG48 (Q8LG48) Contains similarity to p... 33 3e-04
BG662132 40 4e-04
TC19344 weakly similar to GB|AAA34838.1|172092|YSCPABPG polyaden... 39 0.001
TC12138 similar to GB|AAP81806.1|32441262|BT009688 At5g46840 {Ar... 39 0.001
TC14530 similar to UP|P82277 (P82277) Plastid-specific 30S ribos... 38 0.001
TC14791 similar to UP|Q9LJ04 (Q9LJ04) ESTs AU082210(C53655), par... 38 0.002
>TC13012 similar to PIR|T50004|T50004 RNA binding protein-like - Arabidopsis
thaliana {Arabidopsis thaliana;}, partial (84%)
Length = 625
Score = 261 bits (667), Expect = 7e-71
Identities = 132/170 (77%), Positives = 149/170 (87%), Gaps = 3/170 (1%)
Frame = +3
Query: 30 DVDMIGADNNEA---DLDDMKKRLKEMEDEAAALKEMQAKVEKEMGSVQDPANASASQIN 86
DVDM AD++ A +LD+MK+RLKEME+EAAAL+EMQAKVEKE+G+VQDPA A+ASQ N
Sbjct: 117 DVDMSAADDDAAAVKELDEMKRRLKEMEEEAAALREMQAKVEKEIGTVQDPAAAAASQAN 296
Query: 87 REEVDARSIFVGNVDYACTPEDVQQHFQSCGTVNRVTIRTDKFGQPKGYAYVEFVEVEAV 146
+EE D RS+FVGNVDYACTPE+VQQHFQSCGTVNRVTI TDKFGQPKG+AYVEFVE EAV
Sbjct: 297 KEETDGRSVFVGNVDYACTPEEVQQHFQSCGTVNRVTILTDKFGQPKGFAYVEFVEAEAV 476
Query: 147 QEALLLNESELHGRQLKVTAKRTNVPGMKQFRPRRPSNPYMGFRGRTPYA 196
QEAL LNESELHGRQLKV KRTNVPGMKQ+RPRR +PYM + R PYA
Sbjct: 477 QEALRLNESELHGRQLKVLPKRTNVPGMKQYRPRR-YDPYMAYGFRRPYA 623
>TC16073 similar to UP|Q9FJN9 (Q9FJN9) Poly(A)-binding protein II-like,
partial (59%)
Length = 529
Score = 199 bits (505), Expect = 4e-52
Identities = 101/137 (73%), Positives = 117/137 (84%)
Frame = +1
Query: 23 D*DMENDDVDMIGADNNEADLDDMKKRLKEMEDEAAALKEMQAKVEKEMGSVQDPANASA 82
D D E D D N+ DL+DMKKRLKE+E+EA+AL+EMQAKVEK+MG+VQD A +SA
Sbjct: 124 DADAEQQDHDFASNHPNK-DLEDMKKRLKEIEEEASALREMQAKVEKDMGAVQD-AGSSA 297
Query: 83 SQINREEVDARSIFVGNVDYACTPEDVQQHFQSCGTVNRVTIRTDKFGQPKGYAYVEFVE 142
+Q +EEVD RSI+VGNVDYACTPE+VQQHFQSCGTVNRVTI TDKFGQPKG+AYVEFVE
Sbjct: 298 TQAEKEEVDGRSIYVGNVDYACTPEEVQQHFQSCGTVNRVTILTDKFGQPKGFAYVEFVE 477
Query: 143 VEAVQEALLLNESELHG 159
EAVQ A++LNESELHG
Sbjct: 478 AEAVQNAVMLNESELHG 528
>AV413552
Length = 285
Score = 46.6 bits (109), Expect = 3e-06
Identities = 30/82 (36%), Positives = 44/82 (53%), Gaps = 2/82 (2%)
Frame = +1
Query: 93 RSIFVGNVDYACTPEDVQQHFQSCGTVNRVTIRTDKFGQPKGYAYVEFVEVEAVQEAL-L 151
+ ++V N+ ++ + D++ F CGTV V I K G+ KGYA+V E Q A+
Sbjct: 19 KKLYVFNLPWSMSAADIKDLFGQCGTVTDVEIIRGKDGRGKGYAFVTMASGEEAQAAVDK 198
Query: 152 LNESELHGRQLKV-TAKRTNVP 172
+ EL GR L+V AKR P
Sbjct: 199 FDTLELSGRILRVELAKRFKKP 264
>BP047377
Length = 564
Score = 46.6 bits (109), Expect = 3e-06
Identities = 29/98 (29%), Positives = 50/98 (50%), Gaps = 2/98 (2%)
Frame = +3
Query: 69 KEMGSVQDPANASASQINREEVDARSIFVGNVDYACTPEDVQQHFQSCGTVNRVTIRTDK 128
+ S P++++ + + R +FVGN+ Y T E + + Q G V + D+
Sbjct: 57 RRWSSRASPSSSAVDFMASSQSQHRCVFVGNIPYDATEEQLIEICQEVGPVVSFRLVIDR 236
Query: 129 -FGQPKGYAYVEFVEVE-AVQEALLLNESELHGRQLKV 164
G+PKGY + E+ + E A+ L E++GRQL+V
Sbjct: 237 ETGKPKGYGFCEYKDEETALSARRNLQGYEINGRQLRV 350
>TC14336 similar to UP|Q9M549 (Q9M549) Poly(A)-binding protein, partial
(85%)
Length = 1917
Score = 44.7 bits (104), Expect = 1e-05
Identities = 24/73 (32%), Positives = 39/73 (52%), Gaps = 6/73 (8%)
Frame = +2
Query: 92 ARSIFVGNVDYACTPEDVQQHFQSCGTVNRVTIRTDKFGQPKGYAYVEFVEVEAVQEA-- 149
A +IF+ N+D A + + F + G + + TD GQ KGY +V+F EA Q A
Sbjct: 89 AGNIFIKNLDKAIDHKALHDTFSTFGNILSCKVATDSSGQSKGYGFVQFETEEAAQNAIE 268
Query: 150 ----LLLNESELH 158
+LLN+ +++
Sbjct: 269 KLNGMLLNDKQVY 307
Score = 40.0 bits (92), Expect = 3e-04
Identities = 40/189 (21%), Positives = 74/189 (38%)
Frame = +2
Query: 29 DDVDMIGADNNEADLDDMKKRLKEMEDEAAALKEMQAKVEKEMGSVQDPANASASQINRE 88
DD N +DD + + + + ++ +E++ K E+ M D
Sbjct: 518 DDAARAVESLNGKKVDDKEWYVGKAQKKSEREQELKNKFEQSMKEAAD------------ 661
Query: 89 EVDARSIFVGNVDYACTPEDVQQHFQSCGTVNRVTIRTDKFGQPKGYAYVEFVEVEAVQE 148
+ +++V N+D + E +++ F S GT+ + D G +G +V F E
Sbjct: 662 KYQGANLYVKNLDDSIADEKLKELFSSYGTITSCKVMRDPNGISRGSGFVAFSTPEEASR 841
Query: 149 ALLLNESELHGRQLKVTAKRTNVPGMKQFRPRRPSNPYMGFRGRTPYAPPFAYAPAPYGY 208
ALL E++G+ + + K+ R R + R P A PAP
Sbjct: 842 ALL----EMNGKMVVSKPLYVTLAQRKEDRRARLQAQFAQMR-------PVAMPPAP--- 979
Query: 209 GKIPRFRMG 217
++P + G
Sbjct: 980 -RVPMYPPG 1003
Score = 30.4 bits (67), Expect = 0.26
Identities = 13/57 (22%), Positives = 28/57 (48%)
Frame = +2
Query: 94 SIFVGNVDYACTPEDVQQHFQSCGTVNRVTIRTDKFGQPKGYAYVEFVEVEAVQEAL 150
++FV N+ + T ++++ F GT+ + D G+ K + +V F + A+
Sbjct: 368 NVFVKNLSESTTDDELKNVFGEFGTITSAVVMRDGDGKSKCFGFVNFENADDAARAV 538
>AV411398
Length = 427
Score = 43.9 bits (102), Expect = 2e-05
Identities = 30/82 (36%), Positives = 39/82 (46%), Gaps = 1/82 (1%)
Frame = +3
Query: 88 EEVDARSIFVGNVDYACTPEDVQQHFQSCGTVNRVTIRTDKFGQPKGYAYVEFVEVEAVQ 147
EE + NV + TPEDV+ F+ GTV V + + +G A+VE E
Sbjct: 63 EEFSTTRLVAQNVPWTSTPEDVRSLFERYGTVLEVELSMYNKTRSRGLAFVEMSSPEEAL 242
Query: 148 EALLLNES-ELHGRQLKVTAKR 168
EAL ES E GR LK+ R
Sbjct: 243 EALNKLESYEFEGRVLKLNYAR 308
>BP048490
Length = 452
Score = 43.5 bits (101), Expect = 3e-05
Identities = 29/108 (26%), Positives = 55/108 (50%), Gaps = 6/108 (5%)
Frame = +3
Query: 27 ENDDVDMIGADN----NEADLDDMKKRLKEMEDEAAALKEMQAKVEK-EMGSVQDPANAS 81
++DD D +D+ ++ D+D+ E++ + K Q KV+ EM NA
Sbjct: 126 DSDDEDSSDSDSEVKKDKMDVDESDSSEDSDEEDEKSTKTPQKKVKDVEMVDASSGKNAP 305
Query: 82 ASQINREEVD-ARSIFVGNVDYACTPEDVQQHFQSCGTVNRVTIRTDK 128
+ + E + ++++FVGN+ ++ DV+ F+ CG V V + TD+
Sbjct: 306 KTPAAQTENEGSKTLFVGNLSFSVQRSDVENFFKDCGEVVDVRLATDE 449
>BE122637
Length = 332
Score = 43.1 bits (100), Expect = 4e-05
Identities = 28/101 (27%), Positives = 48/101 (46%), Gaps = 2/101 (1%)
Frame = +2
Query: 86 NREEVDARSIFVGNVDYACTPEDVQQHFQSCGTVNRVTIRTDKF-GQPKGYAYVEFVEVE 144
N V S+ V N+ C ++++ F+ G V V I D + G+P+G+A+V+FV+
Sbjct: 2 NSARVTKGSLLVRNIPLDCRADEIRVPFERFGPVRDVYIPKDYYSGEPRGFAFVQFVDPY 181
Query: 145 AVQEALL-LNESELHGRQLKVTAKRTNVPGMKQFRPRRPSN 184
EA +N GR++ V ++ R R S+
Sbjct: 182 DASEAQYHMNRQIFAGREITVVVASETRKRPEEMRHRTRSS 304
>TC12531 similar to UP|Q9ASQ8 (Q9ASQ8) At1g22910/F19G10_13, partial (39%)
Length = 688
Score = 42.7 bits (99), Expect = 5e-05
Identities = 26/97 (26%), Positives = 43/97 (43%), Gaps = 1/97 (1%)
Frame = -1
Query: 95 IFVGNVDYACTPEDVQQHFQSCGTVNRVTIRTDK-FGQPKGYAYVEFVEVEAVQEALLLN 153
+FVG + + E ++++F+ G + + TDK G+ KGY +V F E EA A +
Sbjct: 355 VFVGGLAWETQKETMKKYFEQFGDILEAVVITDKATGRSKGYGFVTFREPEAAMRACVDA 176
Query: 154 ESELHGRQLKVTAKRTNVPGMKQFRPRRPSNPYMGFR 190
+ GR+ V K P++ FR
Sbjct: 175 APVIDGRRANCNLASLGVQRSKPSTPKQHGGAGRNFR 65
>TC14392 similar to UP|Q9LEB4 (Q9LEB4) RNA binding protein 45, partial (77%)
Length = 1576
Score = 42.4 bits (98), Expect = 6e-05
Identities = 46/174 (26%), Positives = 79/174 (44%), Gaps = 10/174 (5%)
Frame = +3
Query: 41 ADLDDMKKRLKEMEDEAAALKEMQA--KVEKEMGS---VQDPANASASQINREEVDARS- 94
AD + + + EM+ + + M+ K +G+ Q A+ SQ + E D +
Sbjct: 765 ADESEQVRAMTEMQGVVCSTRPMRIGPAANKNLGTGTGTQPKASYQNSQGPQNENDPNNT 944
Query: 95 -IFVGNVDYACTPEDVQQHFQSCGTVNRVTIRTDKFGQPKGYAYVEFVEVEAVQEAL-LL 152
IFVGN+D T + ++Q F G + V K Q K +V+F + +EAL +L
Sbjct: 945 TIFVGNLDPNVTDDHLRQVFGLYGDLVHV-----KIPQGKRCGFVQFADRSCAEEALRVL 1109
Query: 153 NESELHGRQLKVTAKRTNVPGMKQFRPRRPSNPY--MGFRGRTPYAPPFAYAPA 204
N + L G+ ++++ R+ Q P + +N G+ G + YAPA
Sbjct: 1110NGTLLGGQNVRLSWGRSPSNKQTQSDPSQWNNSAGGAGYYGYAQGYENYGYAPA 1271
>TC16457 similar to UP|Q41124 (Q41124) Chloroplast RNA binding protein
precursor, partial (30%)
Length = 655
Score = 42.0 bits (97), Expect = 8e-05
Identities = 25/73 (34%), Positives = 42/73 (57%), Gaps = 2/73 (2%)
Frame = +2
Query: 95 IFVGNVDYACTPEDVQQHFQSCGTVNRVTIRTD-KFGQPKGYAYVEFVEVEAVQEALL-L 152
+FVGN+ + T E + Q F GTV + D + G+ +GY +V + + ++ AL+ L
Sbjct: 20 LFVGNLSWTVTSESLIQVFPEYGTVVGARVLYDGETGRSRGYGFVCYSKRSELETALISL 199
Query: 153 NESELHGRQLKVT 165
N EL GR ++V+
Sbjct: 200 NNVELEGRAIRVS 238
>TC14208 similar to UP|Q9LEB4 (Q9LEB4) RNA binding protein 45, partial (41%)
Length = 837
Score = 42.0 bits (97), Expect = 8e-05
Identities = 39/136 (28%), Positives = 63/136 (45%), Gaps = 7/136 (5%)
Frame = +2
Query: 86 NREEVDARSIFVGNVDYACTPEDVQQHFQSCGTVNRVTIRTDKFGQPKGYAYVEFVEVEA 145
N + +IFVGN+D + ++Q F G + V K K +V+F +
Sbjct: 170 NENDPSNTTIFVGNLDPNVNDDHLRQVFSPYGELVHV-----KIPAGKRCGFVQFADRSG 334
Query: 146 VQEAL-LLNESELHGRQLKVTAKRTNVPGMKQFRPRRPSNPYM---GFRGRTPYAPPFAY 201
+EAL +LN + L G+ ++++ R+ P KQ +P SN Y G+ G + Y
Sbjct: 335 AEEALRVLNGTLLGGQNVRLSWGRS--PSNKQAQP--DSNQYAAAGGYYGYGQGYENYGY 502
Query: 202 APA---PYGYGKIPRF 214
AP+ P YG P +
Sbjct: 503 APSGQDPNAYGSYPGY 550
>TC15623 similar to UP|P93486 (P93486) Glycine-rich RNA-binding protein
PsGRBP, partial (88%)
Length = 683
Score = 40.8 bits (94), Expect = 2e-04
Identities = 25/97 (25%), Positives = 49/97 (49%), Gaps = 3/97 (3%)
Frame = +1
Query: 72 GSVQDPANASASQINREE-VDARSIFVGNVDYACTPEDVQQHFQSCGTVNRVTIRTDK-F 129
G+ Q AS +N + + +F+G + Y + ++ F S GTV + D+
Sbjct: 118 GAAQRSQAPVASMLNYIRCMSSSKLFIGGLSYGVDDKSLEDAFSSFGTVAEARVIVDRDT 297
Query: 130 GQPKGYAYVEFVEVEAVQEAL-LLNESELHGRQLKVT 165
G+ +G+ +V F E+ AL ++ +L+GR ++V+
Sbjct: 298 GRSRGFGFVSFDSEESASSALSSMDGQDLNGRNIRVS 408
>TC14147 similar to UP|ROC1_NICSY (Q08935) 29 kDa ribonucleoprotein A,
chloroplast precursor (CP29A), partial (63%)
Length = 1602
Score = 40.4 bits (93), Expect = 2e-04
Identities = 25/74 (33%), Positives = 37/74 (49%), Gaps = 2/74 (2%)
Frame = +3
Query: 95 IFVGNVDYACTPEDVQQHFQSCGTVNRVTIRTDKF-GQPKGYAYVEFVE-VEAVQEALLL 152
+FVGN+ ++ + + FQ G V V + DK G +G+A+V EA A
Sbjct: 351 VFVGNLPFSVDSAQLAELFQDAGNVEVVEVIYDKMTGNSRGFAFVTMSSAAEAEAAAQQF 530
Query: 153 NESELHGRQLKVTA 166
N EL GR L+V +
Sbjct: 531 NNYELEGRALRVNS 572
>TC10207 homologue to UP|Q8LG48 (Q8LG48) Contains similarity to
poly(A)-binding protein II, partial (8%)
Length = 521
Score = 33.5 bits (75), Expect(2) = 3e-04
Identities = 16/30 (53%), Positives = 19/30 (63%)
Frame = +2
Query: 168 RTNVPGMKQFRPRRPSNPYMGFRGRTPYAP 197
RTNVPGM + RRP+ GFR R P+ P
Sbjct: 2 RTNVPGMNPYFGRRPA----GFRSRRPFMP 79
Score = 25.8 bits (55), Expect(2) = 3e-04
Identities = 11/18 (61%), Positives = 12/18 (66%)
Frame = +1
Query: 206 YGYGKIPRFRMGMRYSPY 223
Y YG PR+R MRY PY
Sbjct: 103 YAYGS-PRYRRPMRYRPY 153
>BG662132
Length = 409
Score = 39.7 bits (91), Expect = 4e-04
Identities = 22/85 (25%), Positives = 40/85 (46%), Gaps = 1/85 (1%)
Frame = +3
Query: 67 VEKEMGSVQDPANASASQINREEVDARSIFVGNVDYACTPEDVQQHFQSCGTVNRVTIRT 126
+ E + D + + + + D +FVG + T E ++ HF G V+ TI
Sbjct: 3 IRHEESEIGDLGFCTVTMNWKMDSDRAKLFVGGISRDTTEEILKHHFTKYGNVSDSTISI 182
Query: 127 DKFGQ-PKGYAYVEFVEVEAVQEAL 150
D+ + P+G+ +V F ++ A AL
Sbjct: 183 DRTTRSPRGFGFVTFFDLSAAHRAL 257
>TC19344 weakly similar to GB|AAA34838.1|172092|YSCPABPG
polyadenylate-binding protein {Saccharomyces
cerevisiae;} , partial (18%)
Length = 518
Score = 38.5 bits (88), Expect = 0.001
Identities = 16/57 (28%), Positives = 33/57 (57%)
Frame = +3
Query: 94 SIFVGNVDYACTPEDVQQHFQSCGTVNRVTIRTDKFGQPKGYAYVEFVEVEAVQEAL 150
+I+V N+D + E+++ HF +CG++ + D+ G +G+ +V F E +A+
Sbjct: 156 NIYVKNIDDNVSDEELRGHFSACGSITYAKVMKDEKGISQGFGFVCFSTPEEANKAV 326
>TC12138 similar to GB|AAP81806.1|32441262|BT009688 At5g46840 {Arabidopsis
thaliana;}, partial (45%)
Length = 874
Score = 38.5 bits (88), Expect = 0.001
Identities = 34/123 (27%), Positives = 51/123 (40%), Gaps = 15/123 (12%)
Frame = +2
Query: 93 RSIFVGNVDYACTPEDVQQHFQSCGTVNRVT-------IRTDKFGQPKGYAYVEFVEVEA 145
R++FVGN+ + E++ Q F CG N + +R KG AYV F EA
Sbjct: 428 RTVFVGNLPFDVKDEELYQLF--CGIHNLSSSIEAVRVVRDPHLNVGKGIAYVLFKTREA 601
Query: 146 VQEALLLNESELHGRQLKVT--------AKRTNVPGMKQFRPRRPSNPYMGFRGRTPYAP 197
A+ +L R+L+++ +KR N P + P + N TP
Sbjct: 602 ANFAVKRRYLKLRDRELRLSHAKADATPSKRLNTPSKRLETPSKRPNSSPAQAPSTPAKK 781
Query: 198 PFA 200
P A
Sbjct: 782 PVA 790
Score = 30.8 bits (68), Expect = 0.20
Identities = 31/121 (25%), Positives = 53/121 (43%), Gaps = 24/121 (19%)
Frame = +2
Query: 68 EKEMGS----VQDPANASASQINREEVDA--RSIFVGNVDYACTPEDVQQHFQSCGTVNR 121
+K +GS + +PA+ S+ ++ D R+IFVGN+ + + F+ G V
Sbjct: 2 KKTVGSKRKVLDNPADMLVSKEGFDDEDKLLRTIFVGNLPLKVKKKTLMTEFKKFGEVES 181
Query: 122 VTIRT----DKFGQPKG--------------YAYVEFVEVEAVQEALLLNESELHGRQLK 163
V IR+ D KG AY+ F + +A Q +L N + + G ++
Sbjct: 182 VRIRSVPLQDTKKPRKGAILAKKIDESADSVNAYIVFKDEQAAQASLSHNMTVVEGNHIR 361
Query: 164 V 164
V
Sbjct: 362 V 364
>TC14530 similar to UP|P82277 (P82277) Plastid-specific 30S ribosomal
protein 2, chloroplast precursor (PSRP-2), partial (68%)
Length = 1104
Score = 38.1 bits (87), Expect = 0.001
Identities = 25/89 (28%), Positives = 46/89 (51%), Gaps = 2/89 (2%)
Frame = +2
Query: 78 ANASASQINREEVDARSIFVGNVDYACTPEDVQQHFQSCGTVNRVTIRTDKF-GQPKGYA 136
A A + + E D R ++VGN+ T +++ Q V + + DK+ G+ + +A
Sbjct: 188 AVAEQAAATQSEADRR-LYVGNIPRTLTNDELANIVQEHAAVEKAEVMYDKYSGRSRRFA 364
Query: 137 YVEFVEVEAVQEAL-LLNESELHGRQLKV 164
+V VE A+ LN +++ GR++KV
Sbjct: 365 FVTVKTVEDANAAIEKLNGTQIGGREIKV 451
>TC14791 similar to UP|Q9LJ04 (Q9LJ04) ESTs AU082210(C53655), partial (37%)
Length = 1779
Score = 37.7 bits (86), Expect = 0.002
Identities = 35/114 (30%), Positives = 46/114 (39%), Gaps = 10/114 (8%)
Frame = +2
Query: 59 ALKEMQAKVEKEMGSVQ---------DPANASASQINREEVDARSIFVGNVDYACTPEDV 109
ALK Q K+ S Q P NA E R IFV NV P +
Sbjct: 779 ALKNPQKKIGNRTTSSQLASAGPVPATPPNAPPVS----EYTQRKIFVSNVSSDIEPLKL 946
Query: 110 QQHFQSCGTVNRVTIRTDK-FGQPKGYAYVEFVEVEAVQEALLLNESELHGRQL 162
+ F+ G V + DK G+PKG+A + VE+ ++AL E G L
Sbjct: 947 LEFFKQFGEVEDGPLGLDKNTGRPKGFALFVYRSVESAKKALEEPNKEYEGHTL 1108
Score = 36.2 bits (82), Expect = 0.005
Identities = 35/159 (22%), Positives = 63/159 (39%), Gaps = 12/159 (7%)
Frame = +2
Query: 35 GADNNEADLDDMKKRLKEMEDEAAA---LKEMQAKVEKEMGSVQDPANASASQINREEVD 91
G++NNE + ++ L+E E +E + K+ + Q+ +
Sbjct: 425 GSNNNEEEEEEEDLELEEEPVERLLEPFTREQLHSLVKQAAEKYPDFVENVRQLADVDPA 604
Query: 92 ARSIFVGNVDYACTPEDVQQHFQSCGTVNRVTIRTDKF-GQPKGYAYVEFVEVEAVQEAL 150
R IFV + + T E + F G + TDK G+ KGYA++ F + + AL
Sbjct: 605 HRKIFVHGLGWDTTAETLTSVFSKYGEIEDCKAVTDKVTGKSKGYAFILFKHRDGCRRAL 784
Query: 151 LLNESELHGRQLK--------VTAKRTNVPGMKQFRPRR 181
+ ++ R V A N P + ++ R+
Sbjct: 785 KNPQKKIGNRTTSSQLASAGPVPATPPNAPPVSEYTQRK 901
Database: LJGI
Posted date: Jul 30, 2004 11:16 AM
Number of letters in database: 14,692,800
Number of sequences in database: 28,460
Lambda K H
0.319 0.137 0.408
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 3,184,055
Number of Sequences: 28460
Number of extensions: 39211
Number of successful extensions: 266
Number of sequences better than 10.0: 66
Number of HSP's better than 10.0 without gapping: 255
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 263
length of query: 223
length of database: 4,897,600
effective HSP length: 87
effective length of query: 136
effective length of database: 2,421,580
effective search space: 329334880
effective search space used: 329334880
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 16 ( 7.4 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.8 bits)
S2: 53 (25.0 bits)
Medicago: description of AC121245.9


