

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
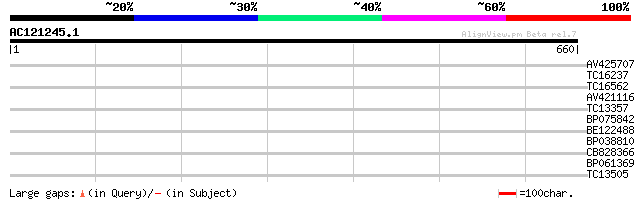
Query= AC121245.1 - phase: 0 /pseudo
(660 letters)
Database: LJGI
28,460 sequences; 14,692,800 total letters
Searching..................................................done
Score E
Sequences producing significant alignments: (bits) Value
AV425707 30 1.6
TC16237 weakly similar to UP|Q84TV1 (Q84TV1) Nodulin-like protei... 30 1.6
TC16562 similar to GB|AAM78073.1|21928089|AY125563 AT4g24660/F22... 30 1.6
AV421116 30 1.6
TC13357 similar to UP|Q9CB00 (Q9CB00) Ara4-interacting protein, ... 28 3.6
BP075842 28 3.6
BE122488 28 4.7
BP038810 27 8.0
CB828366 27 8.0
BP061369 27 8.0
TC13505 similar to UP|Q9T0K5 (Q9T0K5) Extensin-like protein, par... 27 8.0
>AV425707
Length = 252
Score = 29.6 bits (65), Expect = 1.6
Identities = 18/49 (36%), Positives = 28/49 (56%), Gaps = 3/49 (6%)
Frame = -2
Query: 172 IDKMLVQMLNEDPSFKPQGGRLGESDMDYDTDEKSMASL---RRRLRRA 217
++K+L+ L S +P R+ D+D DT+E+ L RRR+RRA
Sbjct: 188 VNKVLMGSLAGPESPRPWSHRMKLRDIDMDTEEEEEEGLLGNRRRMRRA 42
>TC16237 weakly similar to UP|Q84TV1 (Q84TV1) Nodulin-like protein,
partial (11%)
Length = 695
Score = 29.6 bits (65), Expect = 1.6
Identities = 18/68 (26%), Positives = 36/68 (52%)
Frame = +3
Query: 6 VNVTISFLWSLSYWSTFLLTWAVVPLLQGYEDAGDFTVKARLRTSLHGNLVFYLSLGSVA 65
+ ++ F+W S+W+T++LT A V L+G A F + ++ G+L L +
Sbjct: 18 IKFSLQFMWVCSFWNTWVLT-AWVLRLKGPLYASSFNPLFLVIVAIAGSLFLDEKLYLGS 194
Query: 66 LFGLILLI 73
+ G +L++
Sbjct: 195VIGALLIV 218
>TC16562 similar to GB|AAM78073.1|21928089|AY125563 AT4g24660/F22K18_140
{Arabidopsis thaliana;}, partial (29%)
Length = 621
Score = 29.6 bits (65), Expect = 1.6
Identities = 18/61 (29%), Positives = 30/61 (48%)
Frame = -2
Query: 96 LVTGAFLLGFGMSEIPKGIWLNADWAIQQKFLSHKVAKMAVKLDDAHQDFSNAIVITQAT 155
LV G F G E+P + AD+ +Q+ FLS K+A +D + +++T +
Sbjct: 530 LVGGDFAAGLLAVEVPVAVTGGADYGLQRTFLSGG-EKLAAAIDGVAANTDGVVLLTFSV 354
Query: 156 S 156
S
Sbjct: 353 S 351
>AV421116
Length = 421
Score = 29.6 bits (65), Expect = 1.6
Identities = 19/87 (21%), Positives = 37/87 (41%)
Frame = +1
Query: 137 KLDDAHQDFSNAIVITQATSKQMSKRDSLRPYMNIIDKMLVQMLNEDPSFKPQGGRLGES 196
+++ AH D N ++ + + + N + L+ N S +GGR
Sbjct: 52 EIEKAHPDVFNMMLQILEDGRLTDSKGRTVDFKNTL---LIMTSNVGSSVIEKGGRKIGF 222
Query: 197 DMDYDTDEKSMASLRRRLRRAREQYYR 223
D+DYD + S ++ + +QY+R
Sbjct: 223 DLDYDEKDSSYNRIKSLVTEELKQYFR 303
>TC13357 similar to UP|Q9CB00 (Q9CB00) Ara4-interacting protein, partial
(10%)
Length = 424
Score = 28.5 bits (62), Expect = 3.6
Identities = 24/107 (22%), Positives = 49/107 (45%), Gaps = 19/107 (17%)
Frame = +1
Query: 272 DTIEFL--WRCILRKQ-----------------LEKSLAVILGFMSFAILLAEATILPSG 312
++IEF WRC++RKQ +S+ VIL +S IL+ E +
Sbjct: 55 NSIEFRCQWRCLIRKQSTPSSTSPAYLNQSHSRSSRSMGVILMKLSMRILVKETEVY--- 225
Query: 313 VDLSLFSILVHAAGHTEVLVQLAAFVPLMYMCVCTYYSLFKMGMFMF 359
+ + + +L+ + + +F+ + + +C +Y L ++ F+F
Sbjct: 226 LPVCIIPLLLLLLCKMISWI*MMSFMLMNFADLCLFYLLLELIPFLF 366
>BP075842
Length = 434
Score = 28.5 bits (62), Expect = 3.6
Identities = 15/46 (32%), Positives = 26/46 (55%), Gaps = 1/46 (2%)
Frame = +1
Query: 8 VTISFLWSLSYWSTFLLTWAVVPLLQGYEDAG-DFTVKARLRTSLH 52
+ +S L+SLS W T +TW VP ++ G + + AR++ + H
Sbjct: 70 INMSSLFSLSVWDTSFITWIRVPGPCHHKKKGFESLLHARVKLTCH 207
>BE122488
Length = 346
Score = 28.1 bits (61), Expect = 4.7
Identities = 9/12 (75%), Positives = 10/12 (83%)
Frame = +3
Query: 546 HEQKNWWQKVFS 557
HE NWWQK+FS
Sbjct: 9 HETLNWWQKIFS 44
>BP038810
Length = 268
Score = 27.3 bits (59), Expect = 8.0
Identities = 13/30 (43%), Positives = 15/30 (49%)
Frame = -3
Query: 79 WSGSVRGFAMACSNTFGLVTGAFLLGFGMS 108
WSG+ RGF CS F F L F +S
Sbjct: 209 WSGTFRGFLTHCSRLFSQARLRFSLSFLVS 120
>CB828366
Length = 568
Score = 27.3 bits (59), Expect = 8.0
Identities = 16/45 (35%), Positives = 23/45 (50%), Gaps = 10/45 (22%)
Frame = +2
Query: 310 PSGVDLSLFSILVHAAGHTEVLVQLAAFV----------PLMYMC 344
PS ++SLF I+VH+ G E V +A F P+ Y+C
Sbjct: 392 PSSGNISLFLIVVHSPGKYESEVSVAVFSCSFNFCNERNPIHYIC 526
>BP061369
Length = 364
Score = 27.3 bits (59), Expect = 8.0
Identities = 13/29 (44%), Positives = 19/29 (64%)
Frame = +1
Query: 535 YHGRDQK*RS*HEQKNWWQKVFSLKDKPE 563
+HG+++K S H N WQ+ F +KDK E
Sbjct: 157 HHGKEEKHISNHS--NTWQRWFYIKDKRE 237
>TC13505 similar to UP|Q9T0K5 (Q9T0K5) Extensin-like protein, partial (7%)
Length = 359
Score = 27.3 bits (59), Expect = 8.0
Identities = 11/23 (47%), Positives = 17/23 (73%)
Frame = -3
Query: 211 RRRLRRAREQYYRYRSEYTKFVL 233
RRRLRR R +++R R ++T+ L
Sbjct: 204 RRRLRRRRRRWWRRRGKFTRCAL 136
Database: LJGI
Posted date: Jul 30, 2004 11:16 AM
Number of letters in database: 14,692,800
Number of sequences in database: 28,460
Lambda K H
0.348 0.152 0.521
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 11,802,540
Number of Sequences: 28460
Number of extensions: 170132
Number of successful extensions: 1665
Number of sequences better than 10.0: 22
Number of HSP's better than 10.0 without gapping: 1649
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 1664
length of query: 660
length of database: 4,897,600
effective HSP length: 96
effective length of query: 564
effective length of database: 2,165,440
effective search space: 1221308160
effective search space used: 1221308160
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 14 ( 7.0 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 38 (21.8 bits)
S2: 58 (26.9 bits)
Medicago: description of AC121245.1


