

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC121241.10 + phase: 0 /pseudo
(330 letters)
Database: LJGI
28,460 sequences; 14,692,800 total letters
Searching..................................................done
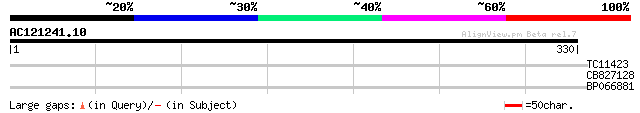
Score E
Sequences producing significant alignments: (bits) Value
TC11423 34 0.030
CB827128 28 2.8
BP066881 27 6.2
>TC11423
Length = 563
Score = 34.3 bits (77), Expect = 0.030
Identities = 12/24 (50%), Positives = 16/24 (66%)
Frame = +1
Query: 3 DSSNFVHSIIPKFDGHYDHWAMFM 26
+S NF +P FDG YDHW++ M
Sbjct: 490 ESGNFTTPCVPIFDGDYDHWSLVM 561
>CB827128
Length = 556
Score = 27.7 bits (60), Expect = 2.8
Identities = 26/101 (25%), Positives = 51/101 (49%), Gaps = 1/101 (0%)
Frame = +1
Query: 110 IRAQLQ-ALRRDFEILHMKEGETINTYFSRTLTIANKMKAHSEIMSQTIITEKILRSMVS 168
+ +QLQ AL+R E KE E + + + ++K H E + + + +S +
Sbjct: 151 VESQLQEALQRHAE----KEAEAKDLN-EKLNALEGQIKLHEEQAREAVAVSETHKSGLE 315
Query: 169 KIDYVVCSIEESNNLDTMTIDELQSSLVHEQRIKSNGEEEN 209
+ S+ + +L+T+ +DELQ+ L+H ++ + EEN
Sbjct: 316 E------SLLKLKHLETV-VDELQNKLLHHEKETAGLNEEN 417
>BP066881
Length = 449
Score = 26.6 bits (57), Expect = 6.2
Identities = 17/55 (30%), Positives = 31/55 (55%), Gaps = 2/55 (3%)
Frame = -1
Query: 157 IITEKILRSMVSKIDYVVCSIEESNN--LDTMTIDELQSSLVHEQRIKSNGEEEN 209
I T KI + VS + ++ I+ S + LD+ +E + L+H+ I+ NGE ++
Sbjct: 392 ISTVKIAQVHVSAAECLLEIIKLSRDVTLDSTINEEFKEELLHQYEIEKNGEAKS 228
Database: LJGI
Posted date: Jul 30, 2004 11:16 AM
Number of letters in database: 14,692,800
Number of sequences in database: 28,460
Lambda K H
0.334 0.144 0.439
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 4,924,165
Number of Sequences: 28460
Number of extensions: 52616
Number of successful extensions: 451
Number of sequences better than 10.0: 6
Number of HSP's better than 10.0 without gapping: 450
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 451
length of query: 330
length of database: 4,897,600
effective HSP length: 91
effective length of query: 239
effective length of database: 2,307,740
effective search space: 551549860
effective search space used: 551549860
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 15 ( 7.2 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 39 (21.6 bits)
S2: 55 (25.8 bits)
Medicago: description of AC121241.10


