

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
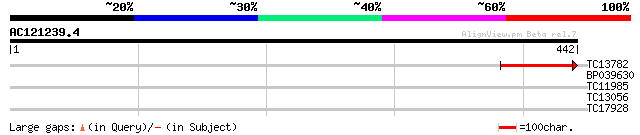
Query= AC121239.4 - phase: 0
(442 letters)
Database: LJGI
28,460 sequences; 14,692,800 total letters
Searching..................................................done
Score E
Sequences producing significant alignments: (bits) Value
TC13782 similar to UP|Q9ST97 (Q9ST97) Alpha-1,3-mannosyl-glycopr... 116 8e-27
BP039630 32 0.27
TC11985 similar to GB|AAF40452.1|7211981|F13M7 ESTs gb|N65605, g... 30 0.61
TC13056 similar to UP|IF38_MEDTR (Q9XHM1) Eukaryotic translation... 30 0.80
TC17928 27 8.8
>TC13782 similar to UP|Q9ST97 (Q9ST97) Alpha-1,3-mannosyl-glycoprotein
beta-1,2-N-acetylglucosaminyltransferase , partial
(14%)
Length = 522
Score = 116 bits (290), Expect = 8e-27
Identities = 56/60 (93%), Positives = 58/60 (96%)
Frame = +3
Query: 383 RIKYKDQWDFENIAQQFGIFQEWKDGVPRTAYKGVVVFRYQTTKRIFLVGPESLKLLQIE 442
RIKYKDQ DFE+IA QFGIFQEWKDGVPRTAYKGVVVFRYQ+TKRIFLVGPESLKLLQIE
Sbjct: 3 RIKYKDQSDFESIASQFGIFQEWKDGVPRTAYKGVVVFRYQSTKRIFLVGPESLKLLQIE 182
>BP039630
Length = 550
Score = 31.6 bits (70), Expect = 0.27
Identities = 23/87 (26%), Positives = 41/87 (46%)
Frame = +3
Query: 133 ISSRFPLFVSQDGSNSDVKRKALSYDELSHMQHLDFEPVQTERPGELIAYYKIARHYKWA 192
+ F F+SQ S + R LS++ H+D + + RPGEL+ ++ + K A
Sbjct: 261 LEEAFCKFLSQPVSVTCSSRTDAGVHALSNVCHVDIQRISKRRPGELLTPHEPSVVRK-A 437
Query: 193 LGQLFDKHNFNRVIILEDDMEIAPDFF 219
+ KH+ + ++I D+ P F
Sbjct: 438 VNHFLQKHDNDLMVI---DVRCVPSDF 509
>TC11985 similar to GB|AAF40452.1|7211981|F13M7 ESTs gb|N65605, gb|N38087,
gb|T20485, gb|T13726, gb|N38339, gb|F15440 and gb|N97201
come from this gene. {Arabidopsis thaliana;}, partial
(88%)
Length = 1334
Score = 30.4 bits (67), Expect = 0.61
Identities = 28/122 (22%), Positives = 51/122 (40%), Gaps = 2/122 (1%)
Frame = +3
Query: 266 GWMLARSTWDELSPKWPKAYWDDWLRLKENHKGRQFIRPEVCRTYNFG--EHGSSLGQFF 323
G+ LA+S W + K K D WLR K+ + R+ + + R++ G++L
Sbjct: 555 GFELAKSCWKQRGMKKTKRENDGWLREKQRKRRREGLERKSSRSWKKTRLREGANLDCLR 734
Query: 324 KQYLEPIKLNEVQVDWKSMDLSYLLEDKYVIHFANIIKKAKPVSGADFAQKARHIDGDVR 383
+ + +P +LS ++E+K KA + + K H + D R
Sbjct: 735 RSHQQP-------------NLSTVVEEKKSCLPVRPATKADQMRECLRSLKQNHKEDDAR 875
Query: 384 IK 385
+K
Sbjct: 876 VK 881
>TC13056 similar to UP|IF38_MEDTR (Q9XHM1) Eukaryotic translation initiation
factor 3 subunit 8 (eIF3 p110) (eIF3c), partial (12%)
Length = 607
Score = 30.0 bits (66), Expect = 0.80
Identities = 19/59 (32%), Positives = 29/59 (48%)
Frame = -1
Query: 152 RKALSYDELSHMQHLDFEPVQTERPGELIAYYKIARHYKWALGQLFDKHNFNRVIILED 210
R++LS + L H + LD P RPG+ K+ + AL Q NF+ ++ L D
Sbjct: 514 RQSLSQEILQHHKSLDVNPGSLVRPGDTHDLLKLLVNLVKALLQTHPVINFDSILHLVD 338
>TC17928
Length = 555
Score = 26.6 bits (57), Expect = 8.8
Identities = 15/50 (30%), Positives = 25/50 (50%), Gaps = 2/50 (4%)
Frame = +3
Query: 355 HFANIIKKAKPVSGADFAQKARHIDGDVRIKYK--DQWDFENIAQQFGIF 402
H I+K+ +SG + KA D ++ + D+ FEN +Q G+F
Sbjct: 165 HNLEIVKELGTLSGCTLSYKALGGRNDYKVALETLDKLQFENPLEQLGLF 314
Database: LJGI
Posted date: Jul 30, 2004 11:16 AM
Number of letters in database: 14,692,800
Number of sequences in database: 28,460
Lambda K H
0.324 0.139 0.424
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 7,280,237
Number of Sequences: 28460
Number of extensions: 95219
Number of successful extensions: 402
Number of sequences better than 10.0: 10
Number of HSP's better than 10.0 without gapping: 402
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 402
length of query: 442
length of database: 4,897,600
effective HSP length: 93
effective length of query: 349
effective length of database: 2,250,820
effective search space: 785536180
effective search space used: 785536180
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 15 ( 7.0 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (22.0 bits)
S2: 57 (26.6 bits)
Medicago: description of AC121239.4


