

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC121237.3 - phase: 1
(455 letters)
Database: LJGI
28,460 sequences; 14,692,800 total letters
Searching..................................................done
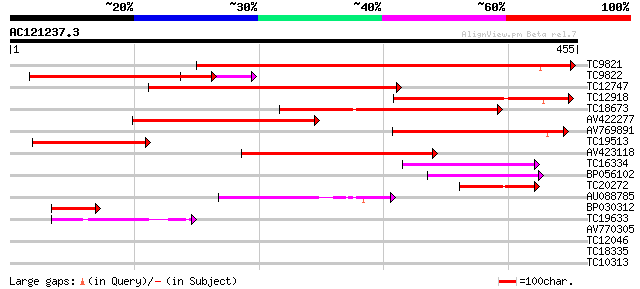
Score E
Sequences producing significant alignments: (bits) Value
TC9821 similar to UP|Q84LI7 (Q84LI7) Polygalacturonase-like prot... 432 e-122
TC9822 similar to UP|Q84LI7 (Q84LI7) Polygalacturonase-like prot... 224 2e-59
TC12747 weakly similar to UP|Q8LAH2 (Q8LAH2) Polygalacturonase-l... 217 3e-57
TC12918 216 6e-57
TC18673 similar to UP|Q9LWY0 (Q9LWY0) ESTs AU029388(E30287), par... 192 1e-49
AV422277 187 2e-48
AV769891 187 4e-48
TC19513 similar to UP|Q84LI7 (Q84LI7) Polygalacturonase-like pro... 146 8e-36
AV423118 136 8e-33
TC16334 100 8e-22
BP056102 87 4e-18
TC20272 weakly similar to UP|Q9LRY8 (Q9LRY8) Polygalacturonase-l... 63 9e-11
AU088785 62 3e-10
BP030312 53 9e-08
TC19633 similar to UP|Q7NET9 (Q7NET9) Glr3789 protein, partial (4%) 46 1e-05
AV770305 32 0.28
TC12046 28 2.4
TC18335 similar to UP|Q7XJ13 (Q7XJ13) Chloroplast nucleoid DNA-b... 28 2.4
TC10313 similar to UP|Q8LB12 (Q8LB12) Seh1-like protein, partial... 28 4.1
>TC9821 similar to UP|Q84LI7 (Q84LI7) Polygalacturonase-like protein,
partial (61%)
Length = 1080
Score = 432 bits (1112), Expect = e-122
Identities = 196/308 (63%), Positives = 250/308 (80%), Gaps = 4/308 (1%)
Frame = +3
Query: 151 GSTWWDKFQKKQLKITRPYMIEIMYSDQIQISNLTLVNSPSWFVHPVYSSNIIINGLTIL 210
G+ WW KF KQL+ TRPY+IEIMYSD +QISNLTLVNSPSW +HPVY SN+I+ G+TIL
Sbjct: 15 GALWWKKFHNKQLQYTRPYLIEIMYSDNVQISNLTLVNSPSWNIHPVYCSNVIVQGITIL 194
Query: 211 APVDVPNTDGIDPDSSTNVLIEDNYIVSGDDCIAIKSGWDEYGIKVGKPSQNIIVRRLTC 270
APV PNTDGI+PDS TN IED YIVSGDDC+A+KSGWDEYGI G P++++++RRLTC
Sbjct: 195 APVTSPNTDGINPDSCTNTRIEDCYIVSGDDCVAVKSGWDEYGISYGMPTKHLVIRRLTC 374
Query: 271 ISPKSALVALGSEMSGGIQDVRIEDVTAINTESAVRIKSAVGRGAFVKDIFVKGMDLNTL 330
ISP SA++ALGSEMSGGI+DVR ED+ AIN+ES VRIK+AVGRG +V+DI+V+ M + T+
Sbjct: 375 ISPTSAVIALGSEMSGGIEDVRAEDILAINSESGVRIKTAVGRGGYVRDIYVRRMTMKTM 554
Query: 331 KYVFWMTGSYGDHPDNGFDPNALPKISGINYRDVTAKNVTIAGKVEGISNDPFTGICVSN 390
K+VFWMTG YG H DN +DPNA+P I INYRD+ A+NVT A ++EGIS PFTGIC+S+
Sbjct: 555 KWVFWMTGDYGSHADNNYDPNAIPVIENINYRDMVAENVTTAPRLEGISGAPFTGICISH 734
Query: 391 VTIEMSAHKKKLPWNCTDISGVTSNVVPKPCELL----KEKEIECPFPRDKLPIENVQFK 446
VT+E++ KK+PW CTD+SG++S V P+PCELL +EK C FP D L +E+VQ +
Sbjct: 735 VTLELAQKAKKVPWTCTDVSGISSGVTPEPCELLPGQAEEKFGACSFPEDHLALEDVQVQ 914
Query: 447 TCNFQSSV 454
TC ++ ++
Sbjct: 915 TCTYRRNL 938
>TC9822 similar to UP|Q84LI7 (Q84LI7) Polygalacturonase-like protein,
partial (40%)
Length = 915
Score = 224 bits (572), Expect = 2e-59
Identities = 104/150 (69%), Positives = 124/150 (82%)
Frame = +1
Query: 17 LNNFDYPAINCRKHSAVLTDFGGVGDGKTLNTKAFNSAITNLSQYANDGGAQLIVPPGKW 76
L NF+Y A+NCR +S +TDFGGVGDG TLNT+AF AI +LSQY+++GG+QL VPPG+W
Sbjct: 112 LENFEYCAVNCRAYSPSVTDFGGVGDGTTLNTQAFRKAIEHLSQYSSNGGSQLYVPPGRW 291
Query: 77 LTGSFNLTSHFTLFLQKDAVILASQYESDWPQLPALPSYGRGREKPGGRFSSLIFGTNLI 136
LTGSFNLTSHFTLF+ KDAVIL SQ E++WP + LPSYGRGR+ GGRFSSLI GT+L
Sbjct: 292 LTGSFNLTSHFTLFIHKDAVILGSQDENEWPVIDPLPSYGRGRDTQGGRFSSLICGTHLT 471
Query: 137 DVIITGNNGTIDGQGSTWWDKFQKKQLKIT 166
DVIITG+NGT+DGQG WW KF KQL+ T
Sbjct: 472 DVIITGDNGTLDGQGDLWWKKFHNKQLQYT 561
Score = 65.1 bits (157), Expect = 2e-11
Identities = 34/61 (55%), Positives = 37/61 (59%)
Frame = +2
Query: 138 VIITGNNGTIDGQGSTWWDKFQKKQLKITRPYMIEIMYSDQIQISNLTLVNSPSWFVHPV 197
VI G N TI I RPY+IEIMYSD +QISNLTLVNSPSW +HPV
Sbjct: 515 VICGGKNSTISSYN-------------IPRPYLIEIMYSDNVQISNLTLVNSPSWNIHPV 655
Query: 198 Y 198
Y
Sbjct: 656 Y 658
>TC12747 weakly similar to UP|Q8LAH2 (Q8LAH2) Polygalacturonase-like
protein, partial (43%)
Length = 624
Score = 217 bits (553), Expect = 3e-57
Identities = 104/203 (51%), Positives = 141/203 (69%)
Frame = +2
Query: 112 LPSYGRGREKPGGRFSSLIFGTNLIDVIITGNNGTIDGQGSTWWDKFQKKQLKITRPYMI 171
LPSYGRG + PGGR+ SLI+G NL DV+ITG +GTIDGQGS WWD L +RP++I
Sbjct: 11 LPSYGRGIDIPGGRYRSLIYGNNLTDVVITGYDGTIDGQGSVWWDLISTGDLNYSRPHII 190
Query: 172 EIMYSDQIQISNLTLVNSPSWFVHPVYSSNIIINGLTILAPVDVPNTDGIDPDSSTNVLI 231
E++ SD I ISNLT++NSPSW +HPVY + I+ +T+ AP + P+T GI PDSS + I
Sbjct: 191 ELIGSDNITISNLTILNSPSWGIHPVYCRYVQIHNITVHAPPESPHTSGIVPDSSEYICI 370
Query: 232 EDNYIVSGDDCIAIKSGWDEYGIKVGKPSQNIIVRRLTCISPKSALVALGSEMSGGIQDV 291
+ + I +G D IA+KSGWDEYG+ GKP+ N+ + + S A +A GSEMSGGI ++
Sbjct: 371 DKSNISTGHDAIALKSGWDEYGVAYGKPTLNVHISSVYLQSSSGAGLAFGSEMSGGISNI 550
Query: 292 RIEDVTAINTESAVRIKSAVGRG 314
E V IN++ + +K+ GRG
Sbjct: 551 IAEQVHIINSKIGIELKTTKGRG 619
>TC12918
Length = 545
Score = 216 bits (550), Expect = 6e-57
Identities = 103/146 (70%), Positives = 120/146 (81%), Gaps = 2/146 (1%)
Frame = +1
Query: 309 SAVGRGAFVKDIFVKGMDLNTLKYVFWMTGSYGDHPDNGFDPNALPKISGINYRDVTAKN 368
+AVGRGA+VK+IFVKGM+L T+KYVFWMTGSYG HPD GFDP ALP I+GINYRDV AKN
Sbjct: 1 TAVGRGAYVKNIFVKGMNLFTMKYVFWMTGSYGSHPDTGFDPKALPTITGINYRDVIAKN 180
Query: 369 VTIAGKVEGISNDPFTGICVSNVTIEMSAHKKKLPWNCTDISGVTSNVVPKPCELLKEK- 427
VT K+EGI+NDPFTGIC+SN IE KKL WNCTD+ GVTSNV P+PC LL+EK
Sbjct: 181 VTYPAKLEGIANDPFTGICISNANIEKVG--KKLAWNCTDVHGVTSNVSPEPCALLQEKP 354
Query: 428 -EIECPFPRDKLPIENVQFKTCNFQS 452
+ +CPFP DKL IE+VQ KTC+F+S
Sbjct: 355 GKFDCPFPSDKLLIESVQLKTCSFKS 432
>TC18673 similar to UP|Q9LWY0 (Q9LWY0) ESTs AU029388(E30287), partial (23%)
Length = 560
Score = 192 bits (488), Expect = 1e-49
Identities = 91/179 (50%), Positives = 127/179 (70%)
Frame = +2
Query: 217 NTDGIDPDSSTNVLIEDNYIVSGDDCIAIKSGWDEYGIKVGKPSQNIIVRRLTCISPKSA 276
NTDGIDPDSS +V IED YI +GDD IAIKSGWDEYGI G+PS NI++RRL + S
Sbjct: 17 NTDGIDPDSSDDVCIEDCYISTGDDLIAIKSGWDEYGIAYGRPSTNIVIRRLVGKTQTSG 196
Query: 277 LVALGSEMSGGIQDVRIEDVTAINTESAVRIKSAVGRGAFVKDIFVKGMDLNTLKYVFWM 336
+A+GSEMSGG+ +V ED+ ++ +A+RIK++ GRG +V++I++ M L ++Y
Sbjct: 197 -IAIGSEMSGGVSEVHAEDIHFYDSYNAIRIKTSRGRGGYVRNIYISNMTLVNIEYAITF 373
Query: 337 TGSYGDHPDNGFDPNALPKISGINYRDVTAKNVTIAGKVEGISNDPFTGICVSNVTIEM 395
G YG+HPD+ +DPNALP I I +DV +N+ AG +EGI D F IC+SN+++ +
Sbjct: 374 NGLYGEHPDDAYDPNALPVIEKITIKDVIGENIKQAGILEGIEGDNFVNICLSNISLNV 550
>AV422277
Length = 454
Score = 187 bits (476), Expect = 2e-48
Identities = 84/150 (56%), Positives = 113/150 (75%)
Frame = +2
Query: 99 ASQYESDWPQLPALPSYGRGREKPGGRFSSLIFGTNLIDVIITGNNGTIDGQGSTWWDKF 158
A+Q +WP + LPSYGRGRE+PGGR+ S I G + DV+ITG NGTIDGQG WW+ +
Sbjct: 5 ATQELGNWPLIAPLPSYGRGRERPGGRYVSFIHGDGVRDVVITGENGTIDGQGDVWWNMW 184
Query: 159 QKKQLKITRPYMIEIMYSDQIQISNLTLVNSPSWFVHPVYSSNIIINGLTILAPVDVPNT 218
+++ L+ TRP ++E + S I ISN+ +SP W +HPVY SN+++ +TILAP D PNT
Sbjct: 185 RQRTLQFTRPNLVEFVNSRNIIISNVIFKDSPFWNIHPVYCSNVVVRYVTILAPRDSPNT 364
Query: 219 DGIDPDSSTNVLIEDNYIVSGDDCIAIKSG 248
DG+DPDSS+NV IED+YI +GDD +A+KSG
Sbjct: 365 DGVDPDSSSNVCIEDSYISTGDDLVAVKSG 454
>AV769891
Length = 526
Score = 187 bits (474), Expect = 4e-48
Identities = 83/146 (56%), Positives = 109/146 (73%), Gaps = 5/146 (3%)
Frame = -3
Query: 308 KSAVGRGAFVKDIFVKGMDLNTLKYVFWMTGSYGDHPDNGFDPNALPKISGINYRDVTAK 367
K+AVGRG +VKDI+V+ M ++T+K+ FWMTG+YG H D +DPNALP+I GINYRD+ A
Sbjct: 524 KTAVGRGGYVKDIYVQRMTMHTIKWAFWMTGNYGSHADKNYDPNALPEIKGINYRDMVAD 345
Query: 368 NVTIAGKVEGISNDPFTGICVSNVTIEMSAHKKKLPWNCTDISGVTSNVVPKPCELLKEK 427
VT+AG +EGISND FTGIC++NVTI M+A KK PW C+D+ G+TS V PKPC LL ++
Sbjct: 344 EVTMAGNLEGISNDQFTGICIANVTISMAAKSKKQPWTCSDVEGITSGVTPKPCNLLPDQ 165
Query: 428 EIE-----CPFPRDKLPIENVQFKTC 448
E C FP + LPI+ ++ K C
Sbjct: 164 GPEKITTTCEFPTETLPIDMLELKKC 87
>TC19513 similar to UP|Q84LI7 (Q84LI7) Polygalacturonase-like protein,
partial (20%)
Length = 442
Score = 146 bits (368), Expect = 8e-36
Identities = 70/95 (73%), Positives = 80/95 (83%)
Frame = +2
Query: 19 NFDYPAINCRKHSAVLTDFGGVGDGKTLNTKAFNSAITNLSQYANDGGAQLIVPPGKWLT 78
+F Y AINCR HSA LTDFGG+GDGKT NTKAF SAI++LSQYA+ GGAQL VP GKWLT
Sbjct: 158 SFVYSAINCRAHSASLTDFGGIGDGKTSNTKAFQSAISHLSQYASKGGAQLYVPAGKWLT 337
Query: 79 GSFNLTSHFTLFLQKDAVILASQYESDWPQLPALP 113
GSF+LTSHFTL+L KDAV+LASQ S+WP + LP
Sbjct: 338 GSFSLTSHFTLYLNKDAVLLASQNISEWPVIEPLP 442
>AV423118
Length = 472
Score = 136 bits (342), Expect = 8e-33
Identities = 63/157 (40%), Positives = 100/157 (63%)
Frame = +2
Query: 187 VNSPSWFVHPVYSSNIIINGLTILAPVDVPNTDGIDPDSSTNVLIEDNYIVSGDDCIAIK 246
+N+P++ +HPVY S + I+ LTI AP + P+T G+ PDSS +V IED I +G D IA+K
Sbjct: 2 LNAPAYSIHPVYCSYVHIHNLTIFAPPESPDTVGLVPDSSDHVCIEDCVIATGYDAIALK 181
Query: 247 SGWDEYGIKVGKPSQNIIVRRLTCISPKSALVALGSEMSGGIQDVRIEDVTAINTESAVR 306
SGWDEYGI G+P++N+ +RR+ + + +A GS+MSGGI +V +E+ N+ +
Sbjct: 182 SGWDEYGIAYGRPTENVHIRRVDLQASSGSALAFGSDMSGGISNVLVENAHLHNSNGGIE 361
Query: 307 IKSAVGRGAFVKDIFVKGMDLNTLKYVFWMTGSYGDH 343
++ GRG ++KDI + +++ + G G H
Sbjct: 362 FRTTRGRGGYMKDIIISDVEMKNVYTAIAARGHCGSH 472
>TC16334
Length = 710
Score = 99.8 bits (247), Expect = 8e-22
Identities = 43/110 (39%), Positives = 64/110 (58%)
Frame = +1
Query: 316 FVKDIFVKGMDLNTLKYVFWMTGSYGDHPDNGFDPNALPKISGINYRDVTAKNVTIAGKV 375
++KDI + L + M G YG HPD+ +DPNA+P + I +++V NV+IAG
Sbjct: 4 YMKDILISDTKLENIHLGIGMIGDYGSHPDDKYDPNAMPVVGSITFKNVVGGNVSIAGNF 183
Query: 376 EGISNDPFTGICVSNVTIEMSAHKKKLPWNCTDISGVTSNVVPKPCELLK 425
GI + PF IC+SNVT +S+ W C+++ G + V PKPC L+
Sbjct: 184 SGIVDSPFAPICLSNVTFSISSESSSPSWFCSNVMGFSEEVFPKPCSDLQ 333
>BP056102
Length = 472
Score = 87.4 bits (215), Expect = 4e-18
Identities = 37/93 (39%), Positives = 53/93 (56%)
Frame = -1
Query: 336 MTGSYGDHPDNGFDPNALPKISGINYRDVTAKNVTIAGKVEGISNDPFTGICVSNVTIEM 395
+ G GDHPD F+PNALP + GI +V V AG ++G+ N PFT +C+S + +
Sbjct: 433 IAGDVGDHPDEKFNPNALPVVKGITITNVWGVRVLQAGLIKGLKNSPFTDVCLSEINLHG 254
Query: 396 SAHKKKLPWNCTDISGVTSNVVPKPCELLKEKE 428
+ + PW C+D+SG V P PC L +E
Sbjct: 253 MSGPRSPPWKCSDVSGFAHQVSPWPCSQLSSQE 155
>TC20272 weakly similar to UP|Q9LRY8 (Q9LRY8) Polygalacturonase-like
protein, partial (5%)
Length = 494
Score = 63.2 bits (152), Expect = 9e-11
Identities = 25/64 (39%), Positives = 43/64 (67%)
Frame = +1
Query: 362 RDVTAKNVTIAGKVEGISNDPFTGICVSNVTIEMSAHKKKLPWNCTDISGVTSNVVPKPC 421
+D+T ++TIAG G+ + PFT IC+S++T+ M+ + W C+++SG + +V PKPC
Sbjct: 7 QDITGTDITIAGNFAGLQDSPFTNICLSHITLSMN-FASSITWECSNVSGYSDSVFPKPC 183
Query: 422 ELLK 425
L+
Sbjct: 184 PNLE 195
>AU088785
Length = 773
Score = 61.6 bits (148), Expect = 3e-10
Identities = 42/146 (28%), Positives = 70/146 (47%), Gaps = 4/146 (2%)
Frame = +2
Query: 168 PYMIEIMYSDQIQISNLTLVNSPSWFVHPVYSSNIIINGLTILAPVDVPNTDGIDPDSST 227
P + M + + N+ +N + + +N+ + L ++AP PNTDGI SS
Sbjct: 14 PSSLFFMDXSNVVVQNIKSLNPKGFHMFVTKCTNVRLRKLKLIAPETSPNTDGIHISSSI 193
Query: 228 NVLIEDNYIVSGDDCIAIKSGWDEYGIKVGKPSQNIIVRRLTCISPKSALVALGS----E 283
NV+I N I +GDDCI++ G S+N+ + RL C P +++GS
Sbjct: 194 NVIIARNTIQTGDDCISMIXG-----------SENVFINRLKC-GPGHG-ISIGSLGKYA 334
Query: 284 MSGGIQDVRIEDVTAINTESAVRIKS 309
++ + I + I T + +RIKS
Sbjct: 335 EXREVKGIXIXNSALIGTTNGLRIKS 412
>BP030312
Length = 418
Score = 53.1 bits (126), Expect = 9e-08
Identities = 25/40 (62%), Positives = 29/40 (72%)
Frame = +3
Query: 34 LTDFGGVGDGKTLNTKAFNSAITNLSQYANDGGAQLIVPP 73
LTDFGGVGDG TLNT AF A++ +S+ GG QL VPP
Sbjct: 297 LTDFGGVGDGVTLNTVAFERAVSAISKLREKGGGQLNVPP 416
>TC19633 similar to UP|Q7NET9 (Q7NET9) Glr3789 protein, partial (4%)
Length = 459
Score = 45.8 bits (107), Expect = 1e-05
Identities = 36/118 (30%), Positives = 53/118 (44%), Gaps = 1/118 (0%)
Frame = +3
Query: 34 LTDFGGVGDGKTLNTKAFNSAITNLSQYANDGGAQLIVP-PGKWLTGSFNLTSHFTLFLQ 92
++DFG GDG +T + SAI + ++I P PGK+LT + L S LF++
Sbjct: 159 VSDFGAAGDGVRYDTTSIQSAINSCP---GSSPCRVIFPAPGKYLTATVFLKSGVVLFVE 329
Query: 93 KDAVILASQYESDWPQLPALPSYGRGREKPGGRFSSLIFGTNLIDVIITGNNGTIDGQ 150
A IL D+P+ P+ ++ N DV I G G +DGQ
Sbjct: 330 PGATILGGPRLEDYPREPSR--------------WYVVLAENATDVGIDG-GGVVDGQ 458
>AV770305
Length = 442
Score = 31.6 bits (70), Expect = 0.28
Identities = 16/55 (29%), Positives = 27/55 (49%)
Frame = +2
Query: 136 IDVIITGNNGTIDGQGSTWWDKFQKKQLKITRPYMIEIMYSDQIQISNLTLVNSP 190
++ + G IDG GS+ W++ + RP + + + +SNL L NSP
Sbjct: 290 VNHLTMNGGGKIDGLGSSGWERCRT----CARPTSLSFYSGNGLTVSNLHLANSP 442
>TC12046
Length = 965
Score = 28.5 bits (62), Expect = 2.4
Identities = 17/43 (39%), Positives = 21/43 (48%), Gaps = 6/43 (13%)
Frame = -3
Query: 64 DGGAQLIVPPGK------WLTGSFNLTSHFTLFLQKDAVILAS 100
D G QL PGK WL F+L +F+QKD I+ S
Sbjct: 699 DYGVQLYKRPGKKGN*VPWLQQVFDLRQGSVVFIQKDYYIIGS 571
>TC18335 similar to UP|Q7XJ13 (Q7XJ13) Chloroplast nucleoid DNA-binding
protein-like protein, partial (19%)
Length = 588
Score = 28.5 bits (62), Expect = 2.4
Identities = 21/66 (31%), Positives = 26/66 (38%)
Frame = +3
Query: 91 LQKDAVILASQYESDWPQLPALPSYGRGREKPGGRFSSLIFGTNLIDVIITGNNGTIDGQ 150
L D V ++YE D P +P GRGR GR F NNG ++
Sbjct: 387 LPADQVKPLNEYEEDGEXSPXMPXXGRGR----GRGRGXXF----------YNNGGMEYG 524
Query: 151 GSTWWD 156
G WD
Sbjct: 525 GGDGWD 542
>TC10313 similar to UP|Q8LB12 (Q8LB12) Seh1-like protein, partial (34%)
Length = 774
Score = 27.7 bits (60), Expect = 4.1
Identities = 13/47 (27%), Positives = 25/47 (52%)
Frame = +2
Query: 220 GIDPDSSTNVLIEDNYIVSGDDCIAIKSGWDEYGIKVGKPSQNIIVR 266
G++PD + +E ++SG D + + WD G+ + Q+ +VR
Sbjct: 143 GLNPDHDGRLPVERVAVLSGHDGMVWQMEWDMSGMTLATTGQDGMVR 283
Database: LJGI
Posted date: Jul 30, 2004 11:16 AM
Number of letters in database: 14,692,800
Number of sequences in database: 28,460
Lambda K H
0.317 0.137 0.414
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 7,562,403
Number of Sequences: 28460
Number of extensions: 95503
Number of successful extensions: 494
Number of sequences better than 10.0: 43
Number of HSP's better than 10.0 without gapping: 483
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 486
length of query: 455
length of database: 4,897,600
effective HSP length: 93
effective length of query: 362
effective length of database: 2,250,820
effective search space: 814796840
effective search space used: 814796840
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 16 ( 7.3 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.6 bits)
S2: 57 (26.6 bits)
Medicago: description of AC121237.3


