

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
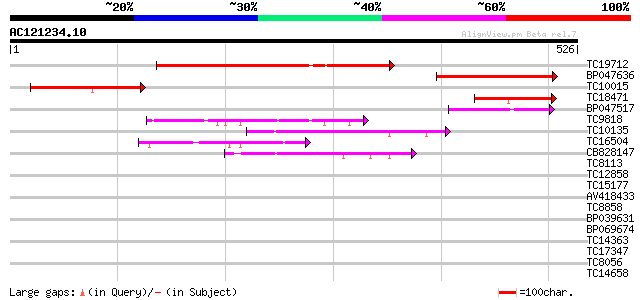
Query= AC121234.10 + phase: 0
(526 letters)
Database: LJGI
28,460 sequences; 14,692,800 total letters
Searching..................................................done
Score E
Sequences producing significant alignments: (bits) Value
TC19712 weakly similar to UP|Q9SA38 (Q9SA38) F3O9.19 protein (At... 232 1e-61
BP047636 175 2e-44
TC10015 similar to UP|Q9SA38 (Q9SA38) F3O9.19 protein (At1g16390... 117 6e-27
TC18471 weakly similar to UP|Q9SA38 (Q9SA38) F3O9.19 protein (At... 73 1e-13
BP047517 60 9e-10
TC9818 similar to UP|Q7XA50 (Q7XA50) Sorbitol-like transporter, ... 60 1e-09
TC10135 52 2e-07
TC16504 weakly similar to UP|Q84KI7 (Q84KI7) Sorbitol transporte... 48 5e-06
CB828147 47 6e-06
TC8113 weakly similar to UP|O23213 (O23213) Sugar transporter li... 38 0.004
TC12858 weakly similar to UP|Q9ZS76 (Q9ZS76) Hexose transporter,... 35 0.023
TC15177 35 0.030
AV418433 30 0.74
TC8858 homologue to UP|Q84N37 (Q84N37) Potyvirus VPg interacting... 30 1.3
BP039631 29 1.7
BP069674 29 2.2
TC14363 homologue to UP|CAT4_SOYBN (O48561) Catalase 4 , partia... 29 2.2
TC17347 similar to UP|Q9FMX3 (Q9FMX3) Monosaccharide transporter... 28 2.8
TC8056 28 3.7
TC14658 similar to UP|Q9SLF1 (Q9SLF1) Nodulin-like protein (At2g... 28 3.7
>TC19712 weakly similar to UP|Q9SA38 (Q9SA38) F3O9.19 protein
(At1g16390/F3O9_19), partial (35%)
Length = 655
Score = 232 bits (592), Expect = 1e-61
Identities = 111/222 (50%), Positives = 165/222 (74%), Gaps = 1/222 (0%)
Frame = +2
Query: 137 LADSSIGRKNMLIFSCVSMSITSMLIIFSTNVWIYSALKFLIGFWRSSIGTCVLVLLTEK 196
LAD+S+GRKNML FSC+ MS++S+L FS+N+WIYS+LKFL GF R++IGT LVL +E
Sbjct: 2 LADTSLGRKNMLFFSCLMMSLSSLLASFSSNIWIYSSLKFLSGFGRATIGTSSLVLASEL 181
Query: 197 VSAEWRFRVGIVEYFTFTMGYMSLPGFAYINRNSSWKSLYIWSSVPAICYSVIAYLFVTE 256
V + R ++ + +F FT+G++SLP AYINRNSSW++LY+W+S+P I Y ++ LFV E
Sbjct: 182 VGKKRRAQISVCGFFLFTIGFLSLPAMAYINRNSSWRNLYLWTSIPTILYCILVKLFVQE 361
Query: 257 SPRWLVMQGREKEILKMLKRVSSEESADDDSVNLASNLPILPPKEKVSF-FQLYSSIGEL 315
SPRWL+++G+++E L LK ++ S ++NLA N I P +E+ + L+S++ L
Sbjct: 362 SPRWLLVRGKKEEALASLKIIT---SITQSNLNLAIN-SIFPQQEEENLNVDLFSALNIL 529
Query: 316 FHKRWAVIRMIAVMILGIGLGMVYFGMPLAVGNLGFNIYLAV 357
K+W+ R++AVM +G G+G+VY+GMPL +GNL FN+YL+V
Sbjct: 530 LRKKWSCTRLLAVMSMGFGIGLVYYGMPLGLGNLSFNLYLSV 655
>BP047636
Length = 534
Score = 175 bits (443), Expect = 2e-44
Identities = 84/112 (75%), Positives = 97/112 (86%)
Frame = -3
Query: 397 CAVLENRVPAAKVVLAMVAFFGACTAYNVFLIYIIELFPTCVRNTTTSLVRQAIVFGCIF 456
C + + AKV LAM +FF ACTAY+VFLIYIIELFPTCVRNTTTSLVRQA+ FGCIF
Sbjct: 532 CVFVGDESQVAKVGLAMASFFFACTAYSVFLIYIIELFPTCVRNTTTSLVRQAVTFGCIF 353
Query: 457 CPFLISAGRKNNIYSYGVFGVVIMLSNFTLFFLPETIGIVLCDTMDQQEKKE 508
PFLIS GRKN+++SYGVFG+VI+LS+ TLF LPET+GIVLCDTMD+QEKKE
Sbjct: 352 IPFLISVGRKNHLFSYGVFGIVIILSHLTLFGLPETMGIVLCDTMDEQEKKE 197
>TC10015 similar to UP|Q9SA38 (Q9SA38) F3O9.19 protein (At1g16390/F3O9_19),
partial (14%)
Length = 436
Score = 117 bits (292), Expect = 6e-27
Identities = 54/114 (47%), Positives = 77/114 (67%), Gaps = 7/114 (6%)
Frame = +3
Query: 20 KKPILSWDVIIEKSLSNFGWMDFLQAVLVAVAMFFDAQQSFISIYTDNYPKWHCTN---- 75
K+ + S IEK + +F W FLQA+LV+ A FDAQQ+FI+++TD +P WHCT
Sbjct: 93 KQHLPSLGSTIEKVIGDFNWTQFLQALLVSSAWVFDAQQTFITVFTDAHPPWHCTQQQDA 272
Query: 76 ---SICTSSSDICKLPRSSWSWDTHPSNTIISHWNLECASTFITGLPQSSFFIG 126
C++S+++C L ++SW+WD S++IIS W LECAS +TGLP S FF+G
Sbjct: 273 GAAESCSTSNNMCTLEKTSWAWDRPTSSSIISEWGLECASPVMTGLPASLFFMG 434
>TC18471 weakly similar to UP|Q9SA38 (Q9SA38) F3O9.19 protein
(At1g16390/F3O9_19), partial (12%)
Length = 385
Score = 72.8 bits (177), Expect = 1e-13
Identities = 37/78 (47%), Positives = 48/78 (61%), Gaps = 2/78 (2%)
Frame = +2
Query: 432 ELFPTCVRNTTTSLVRQAIVFGCIFCPFLI--SAGRKNNIYSYGVFGVVIMLSNFTLFFL 489
ELFPTCVRN+ S+ R A+VFG +F P L+ G + + YGVFG+ I S FL
Sbjct: 2 ELFPTCVRNSALSMARLAVVFGGVFSPLLVGTGGGGGSEVVCYGVFGLAIGCSGVFGVFL 181
Query: 490 PETIGIVLCDTMDQQEKK 507
PET G + D MD++E K
Sbjct: 182 PETRGRAMSDPMDEEENK 235
>BP047517
Length = 495
Score = 60.1 bits (144), Expect = 9e-10
Identities = 33/98 (33%), Positives = 46/98 (46%)
Frame = -1
Query: 408 KVVLAMVAFFGACTAYNVFLIYIIELFPTCVRNTTTSLVRQAIVFGCIFCPFLISAGRKN 467
++V ++ FG YN+ IY ELFPT VRN QA G I P ++ G
Sbjct: 432 RMVCGVLGIFGMAGTYNLLFIYTSELFPTVVRNAALGCATQASQMGAILAPVVVVMG--- 262
Query: 468 NIYSYGVFGVVIMLSNFTLFFLPETIGIVLCDTMDQQE 505
+ + VF M F+LPET+ + L DT + E
Sbjct: 261 GAWPFVVFAACGMFGGVFAFYLPETLNLPLYDTFNGLE 148
>TC9818 similar to UP|Q7XA50 (Q7XA50) Sorbitol-like transporter, partial
(62%)
Length = 1162
Score = 59.7 bits (143), Expect = 1e-09
Identities = 60/221 (27%), Positives = 104/221 (46%), Gaps = 15/221 (6%)
Frame = +2
Query: 128 LLGSFFLAALADSSIGRKNMLIFSCVSMSITSMLIIFSTNVWIYSALKFLIGFWRSSIGT 187
L+GS LA IGR+ ++F+ + ++L+ FS N W +F+ G IG
Sbjct: 389 LIGSG-LAGRTSDWIGRRYTIVFAGAIFFVGALLMGFSPNYWFLMFGRFIAGI---GIGY 556
Query: 188 CVLV--LLTEKVS--AEWRFRVGIVEYFT---FTMGYMSLPGFAYINRNSSWKSLYIWSS 240
+++ + T +VS + F E F +GY+S F+ ++ W+ + +
Sbjct: 557 ALMIAPVYTAEVSPASSRGFLTSFPEVFINGGILLGYISNFAFSKLSLKVGWRMMLGVGA 736
Query: 241 VPAICYSVIAYLFVTESPRWLVMQGREKEILKMLKRVSSEESADDDSVNL-----ASNLP 295
+P++ V L + ESPRWLVM+GR + +K+L + S +S ++ + L A+ +P
Sbjct: 737 LPSVILGV-GVLAMPESPRWLVMRGRLGDAIKVLNKTS--DSPEEAQLRLADIKRAAGIP 907
Query: 296 ILPPKEKVSFFQLYSSIG---ELFHKRWAVIRMIAVMILGI 333
+ V + + G ELF IR I + LGI
Sbjct: 908 ESCTDDVVEVSKRSTGEGVWKELFLYPTPAIRHIVIAALGI 1030
>TC10135
Length = 1501
Score = 52.4 bits (124), Expect = 2e-07
Identities = 46/198 (23%), Positives = 88/198 (44%), Gaps = 8/198 (4%)
Frame = +2
Query: 220 LPGFAYINRNSSWKSLYIWSSVPAICYSVIAYLFVTESPRWLVMQGREKEILKMLKRVSS 279
L A+ N +W+ + ++ PAI ++ L + ESPRWL +GRE+E +LK++ +
Sbjct: 59 LINLAFTNAPGTWRWMLGVAAAPAII-QIVLMLSLPESPRWLYRKGREEESKSILKKIYA 235
Query: 280 EESADDDSVNLASNLPILPPKEKVSFFQLYSSIGELFHKRWAVIRM-IAVMILGIGLGMV 338
E D + L ++ + K S + + + +R M + +G+ V
Sbjct: 236 PEDVDAEIEALKESVESEIEESKTSKISMMTLLKTTTVRRGLYAGMGLQFFQQFVGINTV 415
Query: 339 YFGMPLAVGNLGF-----NIYLAVVFSASMELPSCVATYFLENLRRKPSILV--FSILGG 391
+ P V GF + L+++ S S ++ YF++ RK L+ ++
Sbjct: 416 MYYSPTIVQLAGFASNRTALLLSLITSGLNAFGSILSIYFIDKTGRKKLALISLCGVVLS 595
Query: 392 ICCVTCAVLENRVPAAKV 409
+ +T A E+ + A V
Sbjct: 596 LALLTVAFRESELHAPTV 649
>TC16504 weakly similar to UP|Q84KI7 (Q84KI7) Sorbitol transporter, partial
(37%)
Length = 903
Score = 47.8 bits (112), Expect = 5e-06
Identities = 47/173 (27%), Positives = 77/173 (44%), Gaps = 13/173 (7%)
Frame = +2
Query: 120 QSSFFIGCL----LGSFFLAALADSSIGRKNMLIFSCVSMSITSMLIIFSTNVWIYSALK 175
Q F +G L L + A IGR+ +I + + + S+L+ + + I
Sbjct: 200 QVQFSVGLLHMSALPACMTAGRIADYIGRRYTIILTAIIFLVGSVLMGYGPSYPI----- 364
Query: 176 FLIGFWRSSIGTCVLVLLTEKVSAEWR------FRVGIVEYFT---FTMGYMSLPGFAYI 226
+IG S IG +++ AE F +E T +GY+S F +
Sbjct: 365 LMIGLSISGIGLGFSLIVAPVYCAEISPPSHRGFLTSFLEVSTNAGIALGYVSNYFFGKL 544
Query: 227 NRNSSWKSLYIWSSVPAICYSVIAYLFVTESPRWLVMQGREKEILKMLKRVSS 279
+ W+ + + +VP++ + +V ESPRWLVMQGR E K+L VS+
Sbjct: 545 SIRLGWRLMLVVHAVPSLALVALMLNWV-ESPRWLVMQGRVGEARKVLLLVSN 700
>CB828147
Length = 570
Score = 47.4 bits (111), Expect = 6e-06
Identities = 45/197 (22%), Positives = 83/197 (41%), Gaps = 19/197 (9%)
Frame = +1
Query: 200 EWRFRVGIVEYFTFTMGYMSLPGFAYINRNSSWKSLYIWSSVPAICYSVIAYLFVTESPR 259
E+ +GI+ +GYMS F + N W+ + + P+I + L + ESPR
Sbjct: 1 EFSINIGIL------LGYMSTFFFEKLRLNLGWRMMLAVPAAPSIAL-IFLMLKLGESPR 159
Query: 260 WLVMQGREKEILKMLKRVSSEESADDDSVNLASNLPILPPKEKVSFFQL-------YSSI 312
WL+MQGR + K+L V + + + + + P Q+ ++
Sbjct: 160 WLIMQGRVGDAKKVLLLVFNTKQEAEQRLAEIKTAVGIEPNNTQEIVQVPKKTRHGGGAL 339
Query: 313 GELFHKRWAVIRMIAVMILGI-------GLGMVYFGMPLAVGNLGF----NIYLAVVFSA 361
ELF K ++R I V +G+ G+G++ P G + L +
Sbjct: 340 KELFCKPTPLVRRIMVAAVGLHVFMHLGGIGVILLYSPRIFEKTGITSKSKLLLCTIGMG 519
Query: 362 SMELP-SCVATYFLENL 377
M+L S ++ +F++ +
Sbjct: 520 VMKLVFSFISIFFMDGV 570
>TC8113 weakly similar to UP|O23213 (O23213) Sugar transporter like
protein, partial (93%)
Length = 1865
Score = 38.1 bits (87), Expect = 0.004
Identities = 40/169 (23%), Positives = 70/169 (40%), Gaps = 4/169 (2%)
Frame = +2
Query: 115 ITGLPQSSFFIGCLLGSFFLAALADSSIGRKNMLIFSCVSMSITSMLIIFSTNVWIYSAL 174
+ G+ +GCL A IGR+ + + + + ++ + + N I
Sbjct: 242 LAGILNICALVGCLA-----AGKTSDYIGRRYTIFLASILFLVGAVFMGYGPNFAILMFG 406
Query: 175 KFLIGFWRSSIGTCVLVLLTEKVSAEWR-FRVGIVEY---FTFTMGYMSLPGFAYINRNS 230
+ + G T V E SA R F + E +GY+S +
Sbjct: 407 RCVCGLGVGFALTTAPVYSAELSSASTRGFLTSLPEVCIGLGIFIGYISNYFLGKLALTL 586
Query: 231 SWKSLYIWSSVPAICYSVIAYLFVTESPRWLVMQGREKEILKMLKRVSS 279
W+ + +++P++ + + L + ESPRWLVMQGR K+L VS+
Sbjct: 587 GWRLMLGLAAIPSLGLA-LGILTMPESPRWLVMQGRLGCAKKVLLEVSN 730
>TC12858 weakly similar to UP|Q9ZS76 (Q9ZS76) Hexose transporter, partial
(51%)
Length = 882
Score = 35.4 bits (80), Expect = 0.023
Identities = 46/242 (19%), Positives = 96/242 (39%), Gaps = 9/242 (3%)
Frame = +3
Query: 168 VWIYSALKFLIGFWRSSIGTCVLVLLTEKVSAEWRFRVGIVEYFTFTMGYMSLPGFAY-- 225
+W+ + L+GF V + L+E ++R + + + T+G F Y
Sbjct: 111 IWMLIVGRLLLGFGIGCANQSVPIYLSEMAPYKYRGGLNMCFQLSITIGIFVANLFNYYF 290
Query: 226 --INRNSSWKSLYIWSSVPAICYSVIAYLFVTESPRWLVMQGREKEILKMLKRVSSEESA 283
I W+ +VPA+ + V+ L + +SP LV +GR + + L ++
Sbjct: 291 AKILNGQGWRLSLGLGAVPAVIF-VVGSLCLPDSPSSLVARGRHEAARQELVKI---RGT 458
Query: 284 DDDSVNLASNLPILPPKEKVSFFQLYSSIGELFHKRWAVIRM-IAVMILGIGLGMVYFGM 342
D L + E V + ++ E ++ V + I GL ++ F
Sbjct: 459 TDIEAELKDIITASEALENVK--HPWKTLLERKYRPQLVFAVCIPFFQQFTGLNVITFYA 632
Query: 343 PLAVGNLGF----NIYLAVVFSASMELPSCVATYFLENLRRKPSILVFSILGGICCVTCA 398
P+ +GF ++ AV+ + + + ++ + ++ R+ L GG + C
Sbjct: 633 PILFRTIGFGPTASLMSAVIIGSFKPVSTLISIFVVDKFGRRTLFLE----GGAQMLICQ 800
Query: 399 VL 400
++
Sbjct: 801 II 806
>TC15177
Length = 592
Score = 35.0 bits (79), Expect = 0.030
Identities = 28/73 (38%), Positives = 36/73 (48%)
Frame = -2
Query: 135 AALADSSIGRKNMLIFSCVSMSITSMLIIFSTNVWIYSALKFLIGFWRSSIGTCVLVLLT 194
AAL +SIG + FS S S S L ST IY L FL G + G LVLL
Sbjct: 546 AALESTSIGLDSSSTFSAASFSAASFLCSSSTATGIY--LNFLRGGSSAGSGGASLVLLA 373
Query: 195 EKVSAEWRFRVGI 207
++A W + +G+
Sbjct: 372 --LAAFWLWVIGV 340
>AV418433
Length = 401
Score = 30.4 bits (67), Expect = 0.74
Identities = 15/61 (24%), Positives = 33/61 (53%), Gaps = 3/61 (4%)
Frame = +3
Query: 231 SWKSLYIWSSVPAICYSVIAYLFVTESPRWLVMQGREKEI---LKMLKRVSSEESADDDS 287
SW++L + +P ++ F+ ESPRWL +G ++ L++L+ ++ S + +
Sbjct: 219 SWRALALTGLIPTAVL-LLGLFFIPESPRWLAKRGCREDFEAALQILRGKDADISQEAEE 395
Query: 288 V 288
+
Sbjct: 396 I 398
>TC8858 homologue to UP|Q84N37 (Q84N37) Potyvirus VPg interacting protein
(Fragment), partial (65%)
Length = 1272
Score = 29.6 bits (65), Expect = 1.3
Identities = 30/92 (32%), Positives = 44/92 (47%), Gaps = 14/92 (15%)
Frame = -2
Query: 80 SSSDICKLPRSS-WSWDTHPSNTIISHWNLECASTF-ITGLPQSSF--FIGCLLGSFFLA 135
SSSD+ L R+ WS ++ +WN+ ++ F +T L SS F CL A
Sbjct: 791 SSSDLSDLARAILWSLSASRLASLAFNWNISASAFFSLTILSSSSICNFFFCLSVLSSPA 612
Query: 136 ALADSSIGRK----------NMLIFSCVSMSI 157
+LA S+ R N+LIFS ++SI
Sbjct: 611 SLALSANSRSQASRANRAFLNILIFSSATISI 516
>BP039631
Length = 554
Score = 29.3 bits (64), Expect = 1.7
Identities = 16/53 (30%), Positives = 29/53 (54%)
Frame = +1
Query: 359 FSASMELPSCVATYFLENLRRKPSILVFSILGGICCVTCAVLENRVPAAKVVL 411
F +ME + + FLE + +KP++L+ + +G + CV A N+ +VL
Sbjct: 130 FPYTMETWAELILDFLEEVIQKPTVLMGNSVGSLACVIAASDSNQTLVRGIVL 288
>BP069674
Length = 513
Score = 28.9 bits (63), Expect = 2.2
Identities = 17/67 (25%), Positives = 27/67 (39%), Gaps = 11/67 (16%)
Frame = -1
Query: 29 IIEKSLSNFGWMDFL-----------QAVLVAVAMFFDAQQSFISIYTDNYPKWHCTNSI 77
+ ++ S+ W D L Q L + +F QQ F +P W CT S+
Sbjct: 357 VFQRQFSHLSWEDHLLKVWWKALTPHQRKLALIHIFVLYQQKF-----QTHPTWMCTVSL 193
Query: 78 CTSSSDI 84
C +D+
Sbjct: 192 CQCYTDV 172
>TC14363 homologue to UP|CAT4_SOYBN (O48561) Catalase 4 , partial (63%)
Length = 1045
Score = 28.9 bits (63), Expect = 2.2
Identities = 17/40 (42%), Positives = 22/40 (54%), Gaps = 6/40 (15%)
Frame = +2
Query: 81 SSDICKLPRSSWSWDTHPSNTIISH---W---NLECASTF 114
S D+C+ P S WS DT+ S + H W NLE +S F
Sbjct: 362 SPDMCRFPSSPWSSDTYHSPFLDCHS*AW*P*NLEGSSWF 481
>TC17347 similar to UP|Q9FMX3 (Q9FMX3) Monosaccharide transporter (STP11
protein), partial (30%)
Length = 663
Score = 28.5 bits (62), Expect = 2.8
Identities = 16/60 (26%), Positives = 31/60 (51%)
Frame = +3
Query: 121 SSFFIGCLLGSFFLAALADSSIGRKNMLIFSCVSMSITSMLIIFSTNVWIYSALKFLIGF 180
SS ++ L+ SFF A+ +GRK + + + ++L F+ N+ + + L+GF
Sbjct: 429 SSLYLAALIASFF-ASTTTRMLGRKTSMFAGGLFFLVGALLNGFAVNIEMLIIGRLLLGF 605
>TC8056
Length = 730
Score = 28.1 bits (61), Expect = 3.7
Identities = 16/50 (32%), Positives = 25/50 (50%), Gaps = 8/50 (16%)
Frame = -1
Query: 203 FRVGIVEYFTFTMGYMSLPG--------FAYINRNSSWKSLYIWSSVPAI 244
+ +G+ + T ++G+++L F I R S K IWSSVP I
Sbjct: 715 YNIGLTQPLTHSLGHLTLSMTCLSFITLFQSIKRKKSGKIT*IWSSVPEI 566
>TC14658 similar to UP|Q9SLF1 (Q9SLF1) Nodulin-like protein
(At2g16660/T24I21.7), partial (36%)
Length = 976
Score = 28.1 bits (61), Expect = 3.7
Identities = 25/86 (29%), Positives = 37/86 (42%), Gaps = 3/86 (3%)
Frame = +3
Query: 166 TNVWIYSALKFLIGFWRSSIGTCVLVLLTEKVSAE---WRFRVGIVEYFTFTMGYMSLPG 222
T+V ++ +L + GF+ I V L +K + W I+ + + M+LPG
Sbjct: 42 TDVSLFVSLTSIWGFFGRIISGSVSELFIKKAATPRPLWNAASQILMAVGYILLAMALPG 221
Query: 223 FAYINRNSSWKSLYIWSSVPAICYSV 248
SLYI S V ICY V
Sbjct: 222 -----------SLYIGSIVVGICYGV 266
Database: LJGI
Posted date: Jul 30, 2004 11:16 AM
Number of letters in database: 14,692,800
Number of sequences in database: 28,460
Lambda K H
0.327 0.138 0.431
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 10,760,953
Number of Sequences: 28460
Number of extensions: 174876
Number of successful extensions: 1395
Number of sequences better than 10.0: 49
Number of HSP's better than 10.0 without gapping: 1378
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 1389
length of query: 526
length of database: 4,897,600
effective HSP length: 94
effective length of query: 432
effective length of database: 2,222,360
effective search space: 960059520
effective search space used: 960059520
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 15 ( 7.1 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 40 (21.7 bits)
S2: 57 (26.6 bits)
Medicago: description of AC121234.10


