

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
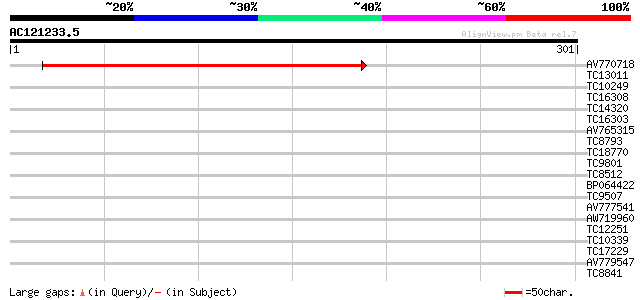
Query= AC121233.5 - phase: 0
(301 letters)
Database: LJGI
28,460 sequences; 14,692,800 total letters
Searching..................................................done
Score E
Sequences producing significant alignments: (bits) Value
AV770718 288 1e-78
TC13011 similar to UP|Q9JM93 (Q9JM93) SRp25 nuclear protein, par... 33 0.059
TC10249 similar to UP|Q84LL9 (Q84LL9) Salt tolerance protein 3, ... 31 0.29
TC16308 similar to GB|AAM16172.1|20334832|AY094016 At2g39720/T5I... 30 0.38
TC14320 similar to UP|Q41111 (Q41111) Dehydrin, partial (56%) 30 0.38
TC16303 weakly similar to UP|AGA1_YEAST (P32323) A-agglutinin at... 30 0.50
AV765315 30 0.50
TC8793 homologue to UP|ENPL_CATRO (P35016) Endoplasmin homolog p... 30 0.50
TC18770 similar to UP|Q7Y0Q8 (Q7Y0Q8) Rac GDP-dissociation inhib... 30 0.50
TC9801 homologue to UP|Q8LAC6 (Q8LAC6) Glycine rich protein-like... 30 0.65
TC8512 similar to UP|O19470 (O19470) MHC class II H2-I-A-beta ge... 30 0.65
BP064422 30 0.65
TC9507 similar to UP|Q8VZB6 (Q8VZB6) DnaJ-like protein, partial ... 29 0.85
AV777541 29 0.85
AW719960 29 1.1
TC12251 28 1.9
TC10339 homologue to UP|RL24_CICAR (O65743) 60S ribosomal protei... 28 1.9
TC17229 similar to UP|O49595 (O49595) HMG protein (AT3G51880/ORF... 28 2.5
AV779547 27 3.2
TC8841 similar to UP|Q9M3Z0 (Q9M3Z0) 60S ribosomal protein L6, c... 27 3.2
>AV770718
Length = 547
Score = 288 bits (736), Expect = 1e-78
Identities = 146/172 (84%), Positives = 157/172 (90%)
Frame = +2
Query: 18 LVNDDTMIDGEAEYLSEEMSPSESESEEDVKLAEPSKTAVNNRDALLDKLGDISWPENVE 77
LV+DDTM+DGE E +EEMS SES+SEEDVKLAEPSKTAV NRD LLDKLGDISWPENVE
Sbjct: 32 LVSDDTMVDGETEDRNEEMSQSESDSEEDVKLAEPSKTAVYNRDGLLDKLGDISWPENVE 211
Query: 78 WIHKLSIDIDQEQEVDVNDDLARELAFYTQALEGTRQAFEKLQSIGLPFLRPADYYAEMV 137
WIHKLSIDIDQE+EVDVNDDL RELAFYTQA EGTRQA EKLQS+GLPFLRPADYYAEMV
Sbjct: 212 WIHKLSIDIDQEEEVDVNDDLKRELAFYTQAWEGTRQALEKLQSMGLPFLRPADYYAEMV 391
Query: 138 KTDSHMEKVKSRLLAEKQKMVEAEERRKAREAKRLSKEVQSQKLKERAKQKK 189
KTDSHM KVK R+ EK+KM EA+ERRKAREAKR +KEVQ+QKLKERAKQKK
Sbjct: 392 KTDSHMVKVKGRI*TEKRKMEEADERRKAREAKRSAKEVQAQKLKERAKQKK 547
>TC13011 similar to UP|Q9JM93 (Q9JM93) SRp25 nuclear protein, partial (10%)
Length = 593
Score = 33.1 bits (74), Expect = 0.059
Identities = 31/113 (27%), Positives = 47/113 (41%), Gaps = 2/113 (1%)
Frame = -3
Query: 182 KERAKQKKDDIESVKKWRKQRQQSGFADDGADKALDFEDGKVFERSKKKRPGVSPGDRSG 241
K K K + KK +K ++ S ++ + ED + + S+ + G + RS
Sbjct: 582 KPAKKGAKKGADHSKKRKKAKEPSDSEEE------EEEDAEESDHSEDEVNGEAEVKRSR 421
Query: 242 GKAKQAFGKGKKPKK--GDVKNSKFGFGGKKGLKKQNTADTTNDFGGFSKKGA 292
G G+GK PKK +VK S G GG A N G K+G+
Sbjct: 420 GGKTSGKGRGKAPKKSGAEVKGSGRGRGG--------AAAAANKSGSSGKRGS 286
>TC10249 similar to UP|Q84LL9 (Q84LL9) Salt tolerance protein 3, partial
(35%)
Length = 1096
Score = 30.8 bits (68), Expect = 0.29
Identities = 36/155 (23%), Positives = 62/155 (39%), Gaps = 8/155 (5%)
Frame = +1
Query: 108 ALEGTRQAFEKLQSIGLPFLRPADYYAEMVKTDSHMEKVKSRLLAEKQKMVEAEERRKAR 167
A E T+ Q +G +R D+ +E +T + RL E+ EAEERRK +
Sbjct: 49 AAERTQSHVTGKQHVGYGMVR--DFISEF-QTAKEKAIEEERLARER----EAEERRKRK 207
Query: 168 EAK--------RLSKEVQSQKLKERAKQKKDDIESVKKWRKQRQQSGFADDGADKALDFE 219
E + +E K ++R + + D +S ++ ++R G+ D G
Sbjct: 208 EQEYERKRRGDSSDREKHRDKDRDRERDRYRDRDSDRERSRERDVRGYRDGGRG-----V 372
Query: 220 DGKVFERSKKKRPGVSPGDRSGGKAKQAFGKGKKP 254
D ++ R R RS K ++ + P
Sbjct: 373 DSRLGNRRNGGRDRYRDRSRSRSPVKNSYRRS*NP 477
>TC16308 similar to GB|AAM16172.1|20334832|AY094016 At2g39720/T5I7.2
{Arabidopsis thaliana;}, partial (13%)
Length = 640
Score = 30.4 bits (67), Expect = 0.38
Identities = 16/44 (36%), Positives = 25/44 (56%), Gaps = 3/44 (6%)
Frame = -2
Query: 230 KRPGVSPGDRSGGKAKQAFGKGKKP---KKGDVKNSKFGFGGKK 270
+R GV G GGK +G+ P ++G ++ S+FG GGK+
Sbjct: 285 RRCGVYGGSLVGGKRGCRRRRGRPPGSGRRGSLRRSRFGSGGKR 154
>TC14320 similar to UP|Q41111 (Q41111) Dehydrin, partial (56%)
Length = 1231
Score = 30.4 bits (67), Expect = 0.38
Identities = 20/73 (27%), Positives = 36/73 (48%), Gaps = 2/73 (2%)
Frame = +3
Query: 161 EERRKAREAKRLSKEVQSQKLKERAKQKKDDI--ESVKKWRKQRQQSGFADDGADKALDF 218
EE + E + E + K+ E ++KK+++ E KK ++ +D + + D
Sbjct: 450 EEEKPQEEV--IVTEFEKVKVSEEPEKKKEEVDHEGEKKHSTLLEKLHRSDSSSSSSSDE 623
Query: 219 EDGKVFERSKKKR 231
E+GK E KKK+
Sbjct: 624 EEGKEGEEKKKKK 662
>TC16303 weakly similar to UP|AGA1_YEAST (P32323) A-agglutinin attachment
subunit precursor, partial (3%)
Length = 1235
Score = 30.0 bits (66), Expect = 0.50
Identities = 34/118 (28%), Positives = 47/118 (39%), Gaps = 1/118 (0%)
Frame = +1
Query: 182 KERAKQKKDDIESVK-KWRKQRQQSGFADDGADKALDFEDGKVFERSKKKRPGVSPGDRS 240
+ER++ D +S K K RK+ + G A D ED +V E R G +
Sbjct: 247 RERSQTPVYDNDSSKSKPRKRLIKKGGGGGSQSVAPDLED-EVEEEDFPVRSDEGEGRKR 423
Query: 241 GGKAKQAFGKGKKPKKGDVKNSKFGFGGKKGLKKQNTADTTNDFGGFSKKGAVGGSKK 298
+ GK +K KGD + GGK G K FGG + K G K+
Sbjct: 424 KKEKDGGSGKKEKRLKGDTRFGSSSSGGKGGSK---FGSAKKGFGGKAGKDHDGEVKE 588
>AV765315
Length = 510
Score = 30.0 bits (66), Expect = 0.50
Identities = 17/63 (26%), Positives = 35/63 (54%)
Frame = -1
Query: 152 AEKQKMVEAEERRKAREAKRLSKEVQSQKLKERAKQKKDDIESVKKWRKQRQQSGFADDG 211
AEK+ EAE+ RKARE + + +++ E+ ++++D ++ R++ + DD
Sbjct: 477 AEKEAFEEAEKERKAREEEEAQR---AREEGEKRRRERDHKYGDRRRRRRPDPHDYMDDY 307
Query: 212 ADK 214
D+
Sbjct: 306 HDE 298
>TC8793 homologue to UP|ENPL_CATRO (P35016) Endoplasmin homolog precursor
(GRP94 homolog), partial (12%)
Length = 576
Score = 30.0 bits (66), Expect = 0.50
Identities = 18/52 (34%), Positives = 29/52 (55%), Gaps = 1/52 (1%)
Frame = +3
Query: 26 DGEAEYLSEEMSPSESESEEDVKLAEPSKTAVNNRDA-LLDKLGDISWPENV 76
D E + +ESE++ED +EP+ AVN+ DA + D+L S+P +
Sbjct: 258 DATVEEEDDTEVEAESETKEDASTSEPAGDAVNDADADVKDEL*IASFPSTL 413
>TC18770 similar to UP|Q7Y0Q8 (Q7Y0Q8) Rac GDP-dissociation inhibitor 1,
partial (66%)
Length = 948
Score = 30.0 bits (66), Expect = 0.50
Identities = 17/43 (39%), Positives = 27/43 (62%), Gaps = 1/43 (2%)
Frame = +1
Query: 160 AEERRKAREAKRLSKEVQSQ-KLKERAKQKKDDIESVKKWRKQ 201
AEE E +L ++ Q LKE+ ++ KDD ES++KW++Q
Sbjct: 268 AEETELEEEDSKLELDLGPQFSLKEQLEKDKDD-ESLRKWKEQ 393
>TC9801 homologue to UP|Q8LAC6 (Q8LAC6) Glycine rich protein-like, complete
Length = 981
Score = 29.6 bits (65), Expect = 0.65
Identities = 20/67 (29%), Positives = 29/67 (42%), Gaps = 8/67 (11%)
Frame = +2
Query: 213 DKALDFEDGKVFERSKKKRPGVSPGDRSGGKAKQAFGKGKK--------PKKGDVKNSKF 264
D+ +D + R+ +K PGV G GG+ + A G+ K KG +
Sbjct: 314 DEVIDKVQEETKSRTDRKPPGVGRGRGRGGREEGAGGRPAKGPGRGFDDGAKGPGGRGRG 493
Query: 265 GFGGKKG 271
G GGK G
Sbjct: 494 GLGGKPG 514
>TC8512 similar to UP|O19470 (O19470) MHC class II H2-I-A-beta gene (K
haplotype), 3' end (Fragment), partial (6%)
Length = 604
Score = 29.6 bits (65), Expect = 0.65
Identities = 16/53 (30%), Positives = 29/53 (54%)
Frame = +2
Query: 144 EKVKSRLLAEKQKMVEAEERRKAREAKRLSKEVQSQKLKERAKQKKDDIESVK 196
++ R AE++KM+E E + EA ++ ++ KERA+Q K + +K
Sbjct: 107 QEAMQRKKAERKKMIEEAEAKLGAEA------IRKKEAKERARQMKKSMPRMK 247
>BP064422
Length = 529
Score = 29.6 bits (65), Expect = 0.65
Identities = 35/134 (26%), Positives = 51/134 (37%), Gaps = 18/134 (13%)
Frame = -1
Query: 158 VEAEERRKAREAKRLSKEVQSQKLKERAKQ--------------KKDDIESV-KKWRKQR 202
+ A +K EA K VQ +KL++ K+ K IE + +K +R
Sbjct: 526 INARSSKKVXEAXARKKRVQMRKLEKVRKKANAISDQTEISDFSKTKPIEKLYRKADPKR 347
Query: 203 QQSGFADDGADKALDFEDGKVFERSKKKRPGVSPGDRSGGKAKQAFGKGKKPK-KGDVKN 261
+ + + GKV + K+ G GK GKGK PK +G K
Sbjct: 346 PKKEYVVAKKGVPVRTGKGKVLVDRRMKKDARKHGMNQTGKGGSQNGKGKAPKGQGSSKG 167
Query: 262 SKFGFG--GKKGLK 273
S G+KG K
Sbjct: 166 SSKASAQKGRKGNK 125
>TC9507 similar to UP|Q8VZB6 (Q8VZB6) DnaJ-like protein, partial (40%)
Length = 905
Score = 29.3 bits (64), Expect = 0.85
Identities = 42/160 (26%), Positives = 60/160 (37%), Gaps = 1/160 (0%)
Frame = +1
Query: 130 ADYYAEMVKTDSHMEKVKSRL-LAEKQKMVEAEERRKAREAKRLSKEVQSQKLKERAKQK 188
A Y E V +++ K+ + +K E+ RK+ ++LS +V K K +
Sbjct: 463 ASYTTEQVSIIREIKRKKNYYEILGLEKSCTVEDVRKSY--RKLSLKVHPDKNKAPGAE- 633
Query: 189 KDDIESVKKWRKQRQQSGFADDGADKALDFEDGKVFERSKKKRPGVSPGDRSGGKAKQAF 248
E+ K K Q G + L ED V+ER RPG G A+
Sbjct: 634 ----EAFKAVSKAFQCLGNEESKRKYDLSGEDESVYERRAAARPG-------PGAARAYN 780
Query: 249 GKGKKPKKGDVKNSKFGFGGKKGLKKQNTADTTNDFGGFS 288
G + D F FGG + T FGGFS
Sbjct: 781 GYYEADFDADEIFRNFFFGGMR-------PTATASFGGFS 879
>AV777541
Length = 466
Score = 29.3 bits (64), Expect = 0.85
Identities = 19/57 (33%), Positives = 32/57 (55%), Gaps = 6/57 (10%)
Frame = +1
Query: 143 MEKVKSRLLAEKQK-----MVEAEERRKAREAKRLSKEVQSQKLKE-RAKQKKDDIE 193
+E K + LAE Q+ + EE RKA A R ++ ++ + KE RAK ++++ E
Sbjct: 130 IEAEKLQRLAEMQRKKEEAQIRREEERKASSAAREARAIEQLRRKEERAKAQQEEAE 300
>AW719960
Length = 449
Score = 28.9 bits (63), Expect = 1.1
Identities = 16/40 (40%), Positives = 22/40 (55%)
Frame = +3
Query: 232 PGVSPGDRSGGKAKQAFGKGKKPKKGDVKNSKFGFGGKKG 271
PG +PG + G + + +GKK K+G K G GKKG
Sbjct: 300 PGGAPGYGTPGHSAGSGKEGKKGKEGKDGKKKEGKEGKKG 419
>TC12251
Length = 412
Score = 28.1 bits (61), Expect = 1.9
Identities = 17/58 (29%), Positives = 29/58 (49%), Gaps = 1/58 (1%)
Frame = -2
Query: 199 RKQRQQSGFADDGADKALDFEDGKVFERSKKKRPGVSPGDRSGG-KAKQAFGKGKKPK 255
R+++ SGF+ DGAD + + S+ + P GG +A++ G G+K K
Sbjct: 324 RRKQHXSGFSGDGADPEI-----RHHPASRARNPSSDEAANCGGVRARKTHGIGRKNK 166
>TC10339 homologue to UP|RL24_CICAR (O65743) 60S ribosomal protein L24,
complete
Length = 677
Score = 28.1 bits (61), Expect = 1.9
Identities = 20/74 (27%), Positives = 35/74 (47%), Gaps = 10/74 (13%)
Frame = +2
Query: 133 YAEMVKTDSHMEKVKSRLLAEKQKM----------VEAEERRKAREAKRLSKEVQSQKLK 182
Y + K D E VK R A K+ V ++R + E + ++E +++K
Sbjct: 242 YRKQHKKDIAQEAVKKRRRATKKPYSRSIVGATLEVIQKKRTEKPEVRDAAREAALREIK 421
Query: 183 ERAKQKKDDIESVK 196
ER K+ KD+ ++ K
Sbjct: 422 ERIKKTKDEKKAKK 463
>TC17229 similar to UP|O49595 (O49595) HMG protein (AT3G51880/ORF13),
partial (66%)
Length = 540
Score = 27.7 bits (60), Expect = 2.5
Identities = 21/69 (30%), Positives = 29/69 (41%)
Frame = +3
Query: 229 KKRPGVSPGDRSGGKAKQAFGKGKKPKKGDVKNSKFGFGGKKGLKKQNTADTTNDFGGFS 288
K++ V P + GKAK+A KPK+ F +K K +N + G S
Sbjct: 189 KRKAAVKPDKGTAGKAKKAKKDPNKPKRPPSAFFVFLEDFRKTFKAENP-----NVKGVS 353
Query: 289 KKGAVGGSK 297
G GG K
Sbjct: 354 AVGKAGGEK 380
>AV779547
Length = 556
Score = 27.3 bits (59), Expect = 3.2
Identities = 21/87 (24%), Positives = 40/87 (45%)
Frame = +1
Query: 114 QAFEKLQSIGLPFLRPADYYAEMVKTDSHMEKVKSRLLAEKQKMVEAEERRKAREAKRLS 173
QA E Q + + F + YYA + K+ ++ ++Q ++ E ++ S
Sbjct: 244 QAIEGSQILIVVFSK---YYASSTWCLQELAKIADGIVGKRQTVLPVVCDVTPSEVRKQS 414
Query: 174 KEVQSQKLKERAKQKKDDIESVKKWRK 200
LK ++ K+D+E V++WRK
Sbjct: 415 GNYGEAFLKHE-ERFKEDLEMVQRWRK 492
>TC8841 similar to UP|Q9M3Z0 (Q9M3Z0) 60S ribosomal protein L6, complete
Length = 856
Score = 27.3 bits (59), Expect = 3.2
Identities = 19/73 (26%), Positives = 33/73 (45%)
Frame = +2
Query: 123 GLPFLRPADYYAEMVKTDSHMEKVKSRLLAEKQKMVEAEERRKAREAKRLSKEVQSQKLK 182
G+P R Y T + VKS +K EA++++K E + E ++
Sbjct: 425 GVPLRRVNQSYVIATSTKVDVSAVKSDKFDDKYFSKEAQKKKKKGEGEFF--EADKEEKN 598
Query: 183 ERAKQKKDDIESV 195
++KKDD ++V
Sbjct: 599 VLPQEKKDDQKTV 637
Database: LJGI
Posted date: Jul 30, 2004 11:16 AM
Number of letters in database: 14,692,800
Number of sequences in database: 28,460
Lambda K H
0.310 0.130 0.355
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 3,378,001
Number of Sequences: 28460
Number of extensions: 32372
Number of successful extensions: 208
Number of sequences better than 10.0: 47
Number of HSP's better than 10.0 without gapping: 206
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 208
length of query: 301
length of database: 4,897,600
effective HSP length: 90
effective length of query: 211
effective length of database: 2,336,200
effective search space: 492938200
effective search space used: 492938200
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 16 ( 7.2 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 42 (21.7 bits)
S2: 55 (25.8 bits)
Medicago: description of AC121233.5


