

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
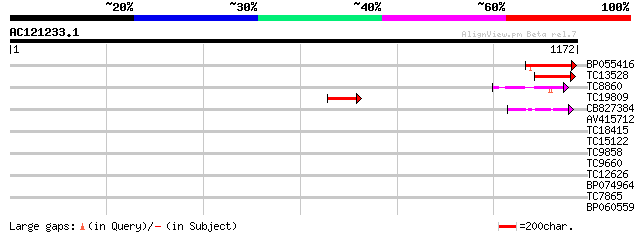
Query= AC121233.1 - phase: 0 /pseudo
(1172 letters)
Database: LJGI
28,460 sequences; 14,692,800 total letters
Searching..................................................done
Score E
Sequences producing significant alignments: (bits) Value
BP055416 173 2e-43
TC13528 similar to UP|Q9SXL4 (Q9SXL4) Histidine kinase 1, partia... 144 1e-34
TC8860 similar to UP|Q9C5T8 (Q9C5T8) Cytokinin receptor CRE1b, p... 78 7e-15
TC19809 similar to UP|Q9C5T8 (Q9C5T8) Cytokinin receptor CRE1b, ... 63 2e-10
CB827384 54 1e-07
AV415712 35 0.090
TC18415 weakly similar to UP|Q9XET9 (Q9XET9) Ethylene receptor h... 34 0.12
TC15122 similar to UP|Q94AY5 (Q94AY5) At1g19950/T20H2_25, partia... 30 2.2
TC9858 similar to UP|Q8L466 (Q8L466) At1g24050/T23E23_11, partia... 29 3.8
TC9660 similar to UP|O49483 (O49483) Receptor protein kinase - l... 29 3.8
TC12626 similar to GB|AAH41272.1|27371281|BC041272 mg:db03h09-pr... 29 3.8
BP074964 29 5.0
TC7865 homologue to UP|SAHH_ARATH (O23255) Adenosylhomocysteinas... 28 6.5
BP060559 28 8.5
>BP055416
Length = 500
Score = 173 bits (438), Expect = 2e-43
Identities = 89/115 (77%), Positives = 95/115 (82%), Gaps = 10/115 (8%)
Frame = -2
Query: 1066 DALNGMPGVE----------RNTITSQTEILSFPSYDLILMDCQMPKMDGYEATKEIRKS 1115
DALN M GVE RN +SQTEIL+ P YDL+LMDCQMPKMDGYEATKEIRKS
Sbjct: 499 DALNAMLGVEDGRRESLLKERNIRSSQTEILNCPPYDLVLMDCQMPKMDGYEATKEIRKS 320
Query: 1116 EIGTSFHIPIVALTAHAMSCDEAKCLEVGMDAYLTKPIDFKLMESTILSLTKRET 1170
E+GT HIPIVALTAHAMSCDEAKCL VGMDAYLTKPIDFKLMESTILS+T+R T
Sbjct: 319 EVGTGLHIPIVALTAHAMSCDEAKCLGVGMDAYLTKPIDFKLMESTILSVTRRTT 155
>TC13528 similar to UP|Q9SXL4 (Q9SXL4) Histidine kinase 1, partial (6%)
Length = 520
Score = 144 bits (362), Expect = 1e-34
Identities = 71/84 (84%), Positives = 75/84 (88%)
Frame = +3
Query: 1085 ILSFPSYDLILMDCQMPKMDGYEATKEIRKSEIGTSFHIPIVALTAHAMSCDEAKCLEVG 1144
I S YDLILMDCQMPKMDGYEATK IRKSE GT+ HIPIVALTAHAMSCDEA+CL+VG
Sbjct: 3 IFSCHPYDLILMDCQMPKMDGYEATKAIRKSEEGTALHIPIVALTAHAMSCDEAQCLQVG 182
Query: 1145 MDAYLTKPIDFKLMESTILSLTKR 1168
MDAYLTKPIDFK M STI+SLTKR
Sbjct: 183 MDAYLTKPIDFKKMVSTIISLTKR 254
>TC8860 similar to UP|Q9C5T8 (Q9C5T8) Cytokinin receptor CRE1b, partial (13%)
Length = 1012
Score = 78.2 bits (191), Expect = 7e-15
Identities = 58/174 (33%), Positives = 81/174 (46%), Gaps = 17/174 (9%)
Frame = +2
Query: 998 TRAKGEDSKGGETSRVSGSSKAMNGKKSLEGLRILLAEDTAVIQRVATIMLEKMGATVVA 1057
TR G+D G + R L G +IL+ +D V +RVA L+ GA V
Sbjct: 11 TRQFGKDMSNGSSVR-----------SLLCGKKILVVDDNLVNRRVAAGALKNFGADVKC 157
Query: 1058 VGDGQQAVDALNGMPGVERNTITSQTEILSFPS-YDLILMDCQMPKMDGYEATKEIR--- 1113
G+ A+ E+L +P +D MD QMP+MDG+EAT+ IR
Sbjct: 158 ALSGKAAL------------------EMLQYPDDFDSCFMDIQMPEMDGFEATRRIRMME 283
Query: 1114 -------KSEIG------TSFHIPIVALTAHAMSCDEAKCLEVGMDAYLTKPID 1154
KSE G + FH+PI+A+TA + KCL GMD Y++KP +
Sbjct: 284 REASEQLKSESGEENGKKSEFHMPILAMTADVIHATYDKCLNCGMDGYVSKPFE 445
>TC19809 similar to UP|Q9C5T8 (Q9C5T8) Cytokinin receptor CRE1b, partial
(11%)
Length = 485
Score = 63.2 bits (152), Expect = 2e-10
Identities = 32/69 (46%), Positives = 44/69 (63%)
Frame = +1
Query: 658 KMILWFEVDDTGCGIDPSKWDSVFESFEQADLSTTRLHGGTGLGLCIVRSLVNKMGGEIK 717
++ L V+DTG GI S DS+F F QAD ST+R +GGTG+GL I + LV MGG+I
Sbjct: 52 QVTLMVSVEDTGIGISFSAQDSIFMPFVQADSSTSRNYGGTGIGLSISKCLVELMGGQIN 231
Query: 718 IVKKEGPGT 726
+ + G+
Sbjct: 232 FISRPQVGS 258
>CB827384
Length = 548
Score = 53.9 bits (128), Expect = 1e-07
Identities = 40/139 (28%), Positives = 69/139 (48%), Gaps = 3/139 (2%)
Frame = +2
Query: 1029 LRILLAEDTAVIQRVATIMLEKMGATVVAVGDGQQAVDALNGMPGVERNTITSQTEILSF 1088
L +L +D+ V ++V +L+ V V G +A+ L G+ G E ++I
Sbjct: 125 LHVLAVDDSLVDRKVIERLLKISSCKVTVVDSGTRALQYL-GLDGEENSSIG-----FDG 286
Query: 1089 PSYDLILMDCQMPKMDGYEATKEIRKSEIGTSFHIPIVALTAHAMSCDEAKCLEVGMDAY 1148
+LI+ D MP M GYE K+I++S IP+V +++ + +CLE G + +
Sbjct: 287 VKVNLIMTDYSMPGMTGYELLKKIKESSAFRE--IPVVVMSSENILTRIDRCLEEGAEDF 460
Query: 1149 LTKPI---DFKLMESTILS 1164
L KP+ D + + I+S
Sbjct: 461 LLKPVKLSDVRRLRDFIMS 517
>AV415712
Length = 353
Score = 34.7 bits (78), Expect = 0.090
Identities = 23/87 (26%), Positives = 40/87 (45%)
Frame = +2
Query: 1029 LRILLAEDTAVIQRVATIMLEKMGATVVAVGDGQQAVDALNGMPGVERNTITSQTEILSF 1088
+ +L +D+ + ++V +L+ V AV G +A+ L P +
Sbjct: 116 VHVLAVDDSLIDRKVIERLLKISSCKVTAVDSGIRALQFLGLEP--------ESDDFAPC 271
Query: 1089 PSYDLILMDCQMPKMDGYEATKEIRKS 1115
DL++ D MP M GYE K+I++S
Sbjct: 272 LKVDLVITDYCMPGMTGYELLKKIKES 352
>TC18415 weakly similar to UP|Q9XET9 (Q9XET9) Ethylene receptor homolog,
partial (9%)
Length = 289
Score = 34.3 bits (77), Expect = 0.12
Identities = 24/73 (32%), Positives = 36/73 (48%), Gaps = 4/73 (5%)
Frame = +2
Query: 1100 MPKMDGYEATKEIRKSEIGTSFHIP----IVALTAHAMSCDEAKCLEVGMDAYLTKPIDF 1155
MP+MDG+E I+ +F+ P IVA A A + KCL GM+ + KPI
Sbjct: 2 MPEMDGFEVAARIQ------NFNRPNWPLIVAFIASAEEHVKEKCLLAGMNGLIRKPIIL 163
Query: 1156 KLMESTILSLTKR 1168
+ + S+ +R
Sbjct: 164 HEIADELRSVLQR 202
>TC15122 similar to UP|Q94AY5 (Q94AY5) At1g19950/T20H2_25, partial (56%)
Length = 1343
Score = 30.0 bits (66), Expect = 2.2
Identities = 16/78 (20%), Positives = 39/78 (49%)
Frame = -2
Query: 149 MRAVTWALFASRKALNSITVKYRNGFVQAFHRDLKDNNIFYIYTDLSYNETNSFAAHEDT 208
+R+++ + FA+ L + Y FV +HR++K +N+ +Y + + +H
Sbjct: 478 LRSISGSCFATY-GLKKESYTYVVPFVFGYHRNMKKSNLASLYIGTHDIKVSPILSHTVK 302
Query: 209 NSNKSAIWYREQLDPVNG 226
+ K W++++ ++G
Sbjct: 301 TATKIQYWHQKRSCSISG 248
>TC9858 similar to UP|Q8L466 (Q8L466) At1g24050/T23E23_11, partial (58%)
Length = 865
Score = 29.3 bits (64), Expect = 3.8
Identities = 15/56 (26%), Positives = 30/56 (52%)
Frame = -3
Query: 428 LCIIVIGCVCILILTNGVSKEMNLRAELISHLEARRKAEASSNYKSQFLANMSHEL 483
L +I+IGC C+++ T +K L EL L + + ++SN++ + + + L
Sbjct: 557 LFLIIIGCKCLILQTTRTAKLQPLPLELEHFLHSVICSRSTSNHRIRVIRTANTHL 390
>TC9660 similar to UP|O49483 (O49483) Receptor protein kinase - like
protein (Receptor protein kinase-like protein), partial
(12%)
Length = 1022
Score = 29.3 bits (64), Expect = 3.8
Identities = 21/76 (27%), Positives = 33/76 (42%), Gaps = 8/76 (10%)
Frame = +1
Query: 662 WFEVDDTGCGIDPS--------KWDSVFESFEQADLSTTRLHGGTGLGLCIVRSLVNKMG 713
W V C DPS +W+ E D+ T ++ G G G+C ++ ++G
Sbjct: 154 WDGVVPPECHPDPSILRLSAALRWEPAHEPLH-TDIDTKKVCG-VGPGMCFANAVRRRVG 327
Query: 714 GEIKIVKKEGPGTLMR 729
GE +V GT M+
Sbjct: 328 GEFGLVPCAVGGTPMK 375
>TC12626 similar to GB|AAH41272.1|27371281|BC041272 mg:db03h09-prov protein
{Xenopus laevis;} , partial (30%)
Length = 540
Score = 29.3 bits (64), Expect = 3.8
Identities = 18/44 (40%), Positives = 23/44 (51%), Gaps = 4/44 (9%)
Frame = -3
Query: 609 IILRGWCENPNSYGDSENFTLEQKPFGCSRKT----KMKQHENH 648
I LRGW P S GD + L+Q+ + CS T + QHE H
Sbjct: 337 INLRGWDNVPISSGDHISPILKQEIYACSFHTNAFPQPYQHETH 206
>BP074964
Length = 418
Score = 28.9 bits (63), Expect = 5.0
Identities = 13/24 (54%), Positives = 16/24 (66%), Gaps = 1/24 (4%)
Frame = +2
Query: 662 WFEVD-DTGCGIDPSKWDSVFESF 684
W EVD D+G G+ P+KW VF F
Sbjct: 224 WSEVDADSGLGLAPAKWLHVFNLF 295
>TC7865 homologue to UP|SAHH_ARATH (O23255) Adenosylhomocysteinase
(S-adenosyl-L-homocysteine hydrolase) (AdoHcyase) ,
complete
Length = 1852
Score = 28.5 bits (62), Expect = 6.5
Identities = 24/79 (30%), Positives = 34/79 (42%), Gaps = 15/79 (18%)
Frame = +1
Query: 1026 LEGLRILLAEDTAVIQRVAT--------IMLEKM-----GATVVAVGDGQQAVD--ALNG 1070
+EGL++L ED + IM++ M A V +G +D L
Sbjct: 970 MEGLQVLTLEDVVSEADIFVTTTGNKDIIMVDHMRKMKNNAIVCNIGHFDNEIDMLGLES 1149
Query: 1071 MPGVERNTITSQTEILSFP 1089
PGV+R TI QT+ FP
Sbjct: 1150 YPGVKRITIKPQTDRFVFP 1206
>BP060559
Length = 386
Score = 28.1 bits (61), Expect = 8.5
Identities = 26/91 (28%), Positives = 41/91 (44%), Gaps = 7/91 (7%)
Frame = +3
Query: 853 LNFLHKYYARA--KFVWLQNHDSSNTVKT--ELRKKGHILTVNKPLYKAKMIHILEDVIK 908
LN L +Y + K W + T+K E +KKG ++ V K K++ +L K
Sbjct: 12 LNMLLQYCLKTL*KSGWRKLEPCFQTLKRVMEKQKKGRMMRVRKDQLKSESKVLLSKR*K 191
Query: 909 ERNVEVQKKNMKEDDL---HESLEIEYTHHC 936
VEVQ++ M+ L + LE+ C
Sbjct: 192 RTRVEVQQEKMQNLKLTLYQQLLELSQNFPC 284
Database: LJGI
Posted date: Jul 30, 2004 11:16 AM
Number of letters in database: 14,692,800
Number of sequences in database: 28,460
Lambda K H
0.318 0.133 0.384
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 18,700,752
Number of Sequences: 28460
Number of extensions: 241116
Number of successful extensions: 1246
Number of sequences better than 10.0: 28
Number of HSP's better than 10.0 without gapping: 1220
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 1237
length of query: 1172
length of database: 4,897,600
effective HSP length: 100
effective length of query: 1072
effective length of database: 2,051,600
effective search space: 2199315200
effective search space used: 2199315200
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 16 ( 7.3 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.7 bits)
S2: 60 (27.7 bits)
Medicago: description of AC121233.1


