

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
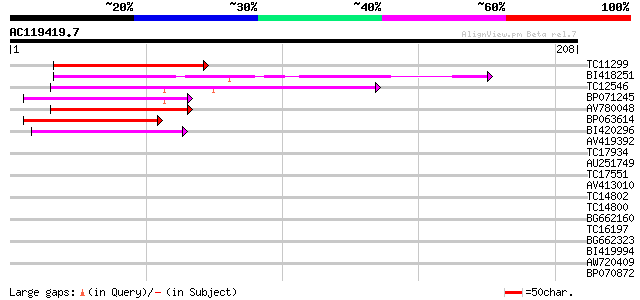
Query= AC119419.7 - phase: 0
(208 letters)
Database: LJGI
28,460 sequences; 14,692,800 total letters
Searching..................................................done
Score E
Sequences producing significant alignments: (bits) Value
TC11299 65 7e-12
BI418251 62 9e-11
TC12546 weakly similar to UP|Q9FJ30 (Q9FJ30) Similarity to heat ... 59 6e-10
BP071245 55 7e-09
AV780048 53 4e-08
BP063614 52 7e-08
BI420296 48 1e-06
AV419392 38 0.001
TC17934 similar to PIR|T06667|T06667 argininosuccinate synthase ... 37 0.002
AU251749 37 0.002
TC17551 weakly similar to UP|AAR24662 (AAR24662) At5g26960, part... 35 0.009
AV413010 34 0.016
TC14802 weakly similar to UP|CAD56661 (CAD56661) S locus F-box (... 31 0.14
TC14800 weakly similar to UP|Q9FHP3 (Q9FHP3) Genomic DNA, chromo... 31 0.14
BG662160 30 0.23
TC16197 similar to PIR|H96696|H96696 protein F1N21.16 [imported]... 30 0.30
BG662323 30 0.30
BI419994 30 0.40
AW720409 28 1.2
BP070872 28 1.5
>TC11299
Length = 498
Score = 65.5 bits (158), Expect = 7e-12
Identities = 28/57 (49%), Positives = 40/57 (70%)
Frame = +1
Query: 17 DRISSLPDDVLNHILSYLPIKTTVATGRLSRQWQHLWKHLHVLDFNEYDDYHRSDNR 73
DRIS+LPD+++ HILS+LP + VAT LS++W HLW+ + LDF+ Y D+R
Sbjct: 85 DRISALPDEIICHILSFLPTENAVATAVLSKRWTHLWRSVSALDFSSVRMYEPDDHR 255
>BI418251
Length = 546
Score = 61.6 bits (148), Expect = 9e-11
Identities = 48/167 (28%), Positives = 76/167 (44%), Gaps = 6/167 (3%)
Frame = +1
Query: 17 DRISSLPDDVLNHILSYLPIKTTVATGRLSRQWQHLWKHLHVLDFNEYDDYHRSDNRKEQ 76
DRIS LPD+VL ILS+LP + VAT L ++W LW + LDF DDY +K +
Sbjct: 37 DRISKLPDEVLCQILSFLPTEDAVATSVLCKRWSSLWLSVPTLDF---DDYRYLKGKKLK 207
Query: 77 LRR------FVVLVNNVLHRNRHGIRKMRLTCAHSLVDDDNFRSHTVDTWVRSVIGPQLE 130
L+ + ++++ LH+ I+ L+ V + V W+ + + Q+E
Sbjct: 208 LQSSFINFVYAIILSRALHQ---PIKNFTLS-----VISEECPYPDVKVWLNAAMQRQVE 363
Query: 131 ELELDLDLFSDDFDEPLDGPDFDEPLDGTDFKLPLSLFTCPNLVSLR 177
L++ L L + LP S+ +C LV L+
Sbjct: 364 NLDICLSLST----------------------LPCSILSCTTLVVLK 438
>TC12546 weakly similar to UP|Q9FJ30 (Q9FJ30) Similarity to heat shock
transcription factor HSF30, partial (9%)
Length = 755
Score = 58.9 bits (141), Expect = 6e-10
Identities = 43/126 (34%), Positives = 62/126 (49%), Gaps = 5/126 (3%)
Frame = +2
Query: 16 DDRISSLPDDVLNHILSYLPIKTTVATGRLSRQWQHLWKH---LHVLDFNEYDDYHRSDN 72
+D IS+LPD +L HILS+L K VAT LSR+W LW+ LH D N + D +D+
Sbjct: 146 EDGISTLPDALLCHILSFLTSKEAVATSVLSRRWIPLWRSVPTLHFKDANYHTDIGHADH 325
Query: 73 R--KEQLRRFVVLVNNVLHRNRHGIRKMRLTCAHSLVDDDNFRSHTVDTWVRSVIGPQLE 130
K+ R V V V+ + + + V + V WV +V+ QLE
Sbjct: 326 DIVKDVRSRHVQSVYAVILSRDFQLPIKKFYLRLNDVCQPFYDPANVSVWVNAVVQRQLE 505
Query: 131 ELELDL 136
L++ L
Sbjct: 506 HLDISL 523
>BP071245
Length = 552
Score = 55.5 bits (132), Expect = 7e-09
Identities = 29/67 (43%), Positives = 40/67 (59%), Gaps = 5/67 (7%)
Frame = +1
Query: 6 SKRQKASEENDDRISSLPDDVLNHILSYLPIKTTVATGRLSRQWQHLWKH-----LHVLD 60
S+ + +EEN DR+S LPD VL HILS++ K V T LS +W+ LWK LH D
Sbjct: 349 SESESENEENKDRLSDLPDCVLLHILSFVNAKYAVQTCTLSTRWKDLWKRLPSLILHSSD 528
Query: 61 FNEYDDY 67
F+ + +
Sbjct: 529 FSTFKGF 549
>AV780048
Length = 501
Score = 52.8 bits (125), Expect = 4e-08
Identities = 25/52 (48%), Positives = 34/52 (65%)
Frame = +2
Query: 16 DDRISSLPDDVLNHILSYLPIKTTVATGRLSRQWQHLWKHLHVLDFNEYDDY 67
+DR SSLPD +L HILS L K V T LS++W LW+ + LDF++ + Y
Sbjct: 248 EDRFSSLPDPILCHILSLLTTKEAVTTSILSKRWIPLWRSVPTLDFDDSNWY 403
>BP063614
Length = 508
Score = 52.0 bits (123), Expect = 7e-08
Identities = 24/51 (47%), Positives = 33/51 (64%)
Frame = +2
Query: 6 SKRQKASEENDDRISSLPDDVLNHILSYLPIKTTVATGRLSRQWQHLWKHL 56
S + +E+N DR+S LPD +L HILS++ K V T LS +W+ LWK L
Sbjct: 134 SLNENENEDNKDRLSDLPDCILLHILSFVKAKAAVQTCILSTRWKDLWKRL 286
>BI420296
Length = 592
Score = 48.1 bits (113), Expect = 1e-06
Identities = 23/57 (40%), Positives = 33/57 (57%)
Frame = +2
Query: 9 QKASEENDDRISSLPDDVLNHILSYLPIKTTVATGRLSRQWQHLWKHLHVLDFNEYD 65
++ +E DR+S LPD VL H+L ++ K V T LS +W+ LWK + L N D
Sbjct: 170 ERDGKEERDRLSDLPDLVLLHVLKFMSTKQAVQTCVLSTRWKDLWKGVTTLALNSSD 340
>AV419392
Length = 318
Score = 37.7 bits (86), Expect = 0.001
Identities = 22/61 (36%), Positives = 31/61 (50%)
Frame = +3
Query: 29 HILSYLPIKTTVATGRLSRQWQHLWKHLHVLDFNEYDDYHRSDNRKEQLRRFVVLVNNVL 88
HILSY+ K + LS++W+ LW+ L VL+F+ H S F V +VL
Sbjct: 3 HILSYVETKDAMQCSVLSKRWRSLWRALPVLNFDSASFTHYSS--------FANFVLHVL 158
Query: 89 H 89
H
Sbjct: 159H 161
>TC17934 similar to PIR|T06667|T06667 argininosuccinate synthase -
Arabidopsis thaliana {Arabidopsis
thaliana;} , partial (11%)
Length = 564
Score = 37.4 bits (85), Expect = 0.002
Identities = 16/35 (45%), Positives = 22/35 (62%)
Frame = -1
Query: 19 ISSLPDDVLNHILSYLPIKTTVATGRLSRQWQHLW 53
I+ LPD + ILS LPI T LSR+W+++W
Sbjct: 564 INQLPDGIPGAILSKLPINEAAKTSILSRKWRYMW 460
>AU251749
Length = 362
Score = 37.4 bits (85), Expect = 0.002
Identities = 19/31 (61%), Positives = 22/31 (70%)
Frame = +2
Query: 17 DRISSLPDDVLNHILSYLPIKTTVATGRLSR 47
D IS+LPD +LN ILS+LP K VAT SR
Sbjct: 137 DWISALPDTILNFILSFLPTKQVVATSGGSR 229
>TC17551 weakly similar to UP|AAR24662 (AAR24662) At5g26960, partial (22%)
Length = 505
Score = 35.0 bits (79), Expect = 0.009
Identities = 18/54 (33%), Positives = 30/54 (55%)
Frame = +2
Query: 10 KASEENDDRISSLPDDVLNHILSYLPIKTTVATGRLSRQWQHLWKHLHVLDFNE 63
K ++ ISSLPDD++ L+ +P + + + R+W HL LH DF++
Sbjct: 101 KPPQQPPTTISSLPDDIILDFLTRVPPSSLPSLSLVCRRWSHL---LHSPDFSD 253
>AV413010
Length = 416
Score = 34.3 bits (77), Expect = 0.016
Identities = 15/45 (33%), Positives = 26/45 (57%)
Frame = +3
Query: 5 NSKRQKASEENDDRISSLPDDVLNHILSYLPIKTTVATGRLSRQW 49
N +R+ +EEN S LPD+++ IL LP+ + + +S+ W
Sbjct: 45 NEQRRSCTEENPSLSSILPDELIIQILLQLPVTSLLRFKSVSKSW 179
>TC14802 weakly similar to UP|CAD56661 (CAD56661) S locus F-box (SLF)-S4
protein, partial (10%)
Length = 551
Score = 31.2 bits (69), Expect = 0.14
Identities = 20/57 (35%), Positives = 30/57 (52%), Gaps = 8/57 (14%)
Frame = +3
Query: 22 LPDDVLNHILSYLPIKTTVATGRLSRQW-------QHLWKH-LHVLDFNEYDDYHRS 70
LPD+++ ILS+LP+K+ + +S+ W Q + H LH L F D H S
Sbjct: 21 LPDELIIEILSWLPVKSLLQFRVVSKTWKSFISDPQFVKLHLLHRLSFRNADFEHTS 191
>TC14800 weakly similar to UP|Q9FHP3 (Q9FHP3) Genomic DNA, chromosome 5, TAC
clone:K14B20, partial (7%)
Length = 782
Score = 31.2 bits (69), Expect = 0.14
Identities = 20/57 (35%), Positives = 30/57 (52%), Gaps = 8/57 (14%)
Frame = +3
Query: 22 LPDDVLNHILSYLPIKTTVATGRLSRQW-------QHLWKH-LHVLDFNEYDDYHRS 70
LPD+++ ILS+LP+K+ + +S+ W Q + H LH L F D H S
Sbjct: 48 LPDELIIEILSWLPVKSLLQFRVVSKTWKSFISDPQFVKLHLLHRLSFRNADFEHTS 218
>BG662160
Length = 445
Score = 30.4 bits (67), Expect = 0.23
Identities = 17/50 (34%), Positives = 26/50 (52%), Gaps = 7/50 (14%)
Frame = +2
Query: 15 NDDRISSLPDDVLNHILSYLPIKTTVATGRLSRQW-------QHLWKHLH 57
N R LP+++ ILS+LP+K+ V +S+ W Q + HLH
Sbjct: 2 NSARAQFLPEELRLEILSWLPVKSLVRFRCVSKSWKSIISDSQFIKLHLH 151
>TC16197 similar to PIR|H96696|H96696 protein F1N21.16 [imported] -
Arabidopsis thaliana {Arabidopsis thaliana;}, partial
(35%)
Length = 582
Score = 30.0 bits (66), Expect = 0.30
Identities = 14/39 (35%), Positives = 20/39 (50%)
Frame = +3
Query: 1 MVESNSKRQKASEENDDRISSLPDDVLNHILSYLPIKTT 39
+ K+ A+ DD SLPDD++ ILS L +T
Sbjct: 108 LTRKRQKKSPAATSGDDHFESLPDDLVISILSKLSSSST 224
>BG662323
Length = 375
Score = 30.0 bits (66), Expect = 0.30
Identities = 12/31 (38%), Positives = 19/31 (60%)
Frame = +1
Query: 22 LPDDVLNHILSYLPIKTTVATGRLSRQWQHL 52
LP D+L HIL YLPI + + ++W+ +
Sbjct: 76 LPGDLLEHILMYLPIASIFRACCVCKRWREI 168
>BI419994
Length = 549
Score = 29.6 bits (65), Expect = 0.40
Identities = 13/46 (28%), Positives = 26/46 (56%)
Frame = +2
Query: 7 KRQKASEENDDRISSLPDDVLNHILSYLPIKTTVATGRLSRQWQHL 52
K + S E+ S LPD+++ IL LP+++ + + + W++L
Sbjct: 2 KHPQTSTESLSLFSHLPDELILQILLRLPVRSLLRFKTVCKSWRYL 139
>AW720409
Length = 575
Score = 28.1 bits (61), Expect = 1.2
Identities = 15/43 (34%), Positives = 25/43 (57%), Gaps = 6/43 (13%)
Frame = +3
Query: 22 LPDDVLNHILSYLPIKTTVATGRLSRQWQHL------WKHLHV 58
LP ++ ILS+LP+KT + +S+ W+ L +K LH+
Sbjct: 93 LPFELWIEILSWLPVKTLMRFSCVSKSWKSLIYQDRDFKKLHL 221
>BP070872
Length = 472
Score = 27.7 bits (60), Expect = 1.5
Identities = 12/39 (30%), Positives = 23/39 (58%)
Frame = +1
Query: 168 FTCPNLVSLRHVLLFFIASFSFLNVKCHCGKIESYTCTS 206
F P+ +SLR +++FF+ ++F N + ++ TC S
Sbjct: 88 FYAPHCISLRSLIMFFLRHYTFYN--NYSFSLQQVTCLS 198
Database: LJGI
Posted date: Jul 30, 2004 11:16 AM
Number of letters in database: 14,692,800
Number of sequences in database: 28,460
Lambda K H
0.323 0.139 0.431
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 4,224,240
Number of Sequences: 28460
Number of extensions: 64807
Number of successful extensions: 437
Number of sequences better than 10.0: 54
Number of HSP's better than 10.0 without gapping: 436
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 436
length of query: 208
length of database: 4,897,600
effective HSP length: 86
effective length of query: 122
effective length of database: 2,450,040
effective search space: 298904880
effective search space used: 298904880
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 16 ( 7.5 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.9 bits)
S2: 53 (25.0 bits)
Medicago: description of AC119419.7


