

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC119419.5 - phase: 0
(735 letters)
Database: LJGI
28,460 sequences; 14,692,800 total letters
Searching..................................................done
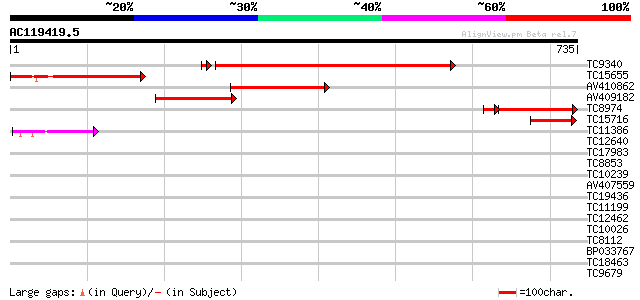
Score E
Sequences producing significant alignments: (bits) Value
TC9340 homologue to PIR|S58083|S58083 transketolase precursor -... 583 e-169
TC15655 similar to UP|Q8LE99 (Q8LE99) Transketolase-like protein... 242 2e-64
AV410862 237 4e-63
AV409182 207 5e-54
TC8974 similar to UP|TKTC_CRAPL (Q42676) Transketolase, chloropl... 180 5e-53
TC15716 similar to UP|O78327 (O78327) Transketolase 1 , partial ... 103 8e-23
TC11386 similar to UP|Q8LE99 (Q8LE99) Transketolase-like protein... 93 1e-19
TC12640 similar to UP|Q7X9B4 (Q7X9B4) Allene oxide synthase prec... 34 0.096
TC17983 similar to UP|Q9XEX5 (Q9XEX5) Cytidine deaminase , part... 33 0.21
TC8853 similar to PIR|T09217|T09217 protein sam2B - spinach {Spi... 33 0.21
TC10239 similar to UP|Q7NEZ8 (Q7NEZ8) Glr3729 protein, partial (8%) 32 0.28
AV407559 32 0.28
TC19436 similar to UP|Q93Y06 (Q93Y06) Receptor protein kinase-li... 32 0.36
TC11199 31 0.62
TC12462 weakly similar to UP|Q8TW78 (Q8TW78) Uncharacterized mem... 31 0.62
TC10026 31 0.62
TC8112 similar to UP|ODPA_PEA (P52902) Pyruvate dehydrogenase E1... 31 0.81
BP033767 31 0.81
TC18463 similar to AAR92278 (AAR92278) At4g22580, partial (34%) 31 0.81
TC9679 weakly similar to UP|Q8H1B5 (Q8H1B5) Hin1-like protein, p... 30 1.1
>TC9340 homologue to PIR|S58083|S58083 transketolase precursor - potato
(fragment) {Solanum tuberosum;} , partial (47%)
Length = 990
Score = 583 bits (1502), Expect(2) = e-169
Identities = 282/311 (90%), Positives = 296/311 (94%)
Frame = +1
Query: 268 AFTESVEKRFEGLGWHVIWVKNGNTGFDEIRAAIKEAKAVKDRPTLIKFTTTIGYGSPNK 327
AFTESV+ RFEGLGWHVIWVKNGN G+D+IRAAIKEAKAV D+PTLIK TTTIG+GSPNK
Sbjct: 58 AFTESVDSRFEGLGWHVIWVKNGNNGYDDIRAAIKEAKAVPDKPTLIKVTTTIGFGSPNK 237
Query: 328 SNSYSVHGSALGAKEVDATRNNLGWPYEPFHVPEDVKKHWSRHIREGAALESEWNAKFAD 387
+NSYSVHGSALGAKEV+ATR NL WP+EPFHVPEDVKKHWSRHI EGAALE+EWNAKFA+
Sbjct: 238 ANSYSVHGSALGAKEVEATRANLAWPHEPFHVPEDVKKHWSRHIPEGAALEAEWNAKFAE 417
Query: 388 YEKKYKEEAAVLKSIISGDLPAGWEKALPTYTPEIPADATRNLSQQNLNALANVLPGLIG 447
YEKKYKEEAA KSII+GDLPAGWEKALPTYTPE PADATRNLSQ NLNALA VLPGL+G
Sbjct: 418 YEKKYKEEAAEFKSIINGDLPAGWEKALPTYTPETPADATRNLSQTNLNALAKVLPGLLG 597
Query: 448 GSADLASSNMTLLKKFGDFQSDSPAERNVRFGVREHGMGAICNGIALHSRGLIPYCATFF 507
GSADLASSNMTLLK FG+FQ D+PAERNVRFGVREHGMGAICNGIALHS GLIPYCATFF
Sbjct: 598 GSADLASSNMTLLKMFGNFQKDTPAERNVRFGVREHGMGAICNGIALHSPGLIPYCATFF 777
Query: 508 VFTDYMRGAIRLSALSEAGVIYVMTHDSIGLGEDGPTHQPIEHLASFRAMPNILMLRPAD 567
VFTDYMRGAIRLSALSEAGVIYVMTHDSIGLGEDGPTHQPIEHLASFRAMPNILMLRPAD
Sbjct: 778 VFTDYMRGAIRLSALSEAGVIYVMTHDSIGLGEDGPTHQPIEHLASFRAMPNILMLRPAD 957
Query: 568 GNETAGAYKVA 578
GNETAGAYKVA
Sbjct: 958 GNETAGAYKVA 990
Score = 29.6 bits (65), Expect(2) = e-169
Identities = 12/13 (92%), Positives = 13/13 (99%)
Frame = +3
Query: 249 KLIAFYDDNHISI 261
KLIAFYDDNHIS+
Sbjct: 3 KLIAFYDDNHISM 41
>TC15655 similar to UP|Q8LE99 (Q8LE99) Transketolase-like protein, partial
(17%)
Length = 575
Score = 242 bits (617), Expect = 2e-64
Identities = 128/185 (69%), Positives = 146/185 (78%), Gaps = 10/185 (5%)
Frame = +2
Query: 2 SSSSSTSLYLTNPHLTRHTSSPSTTRLSSLPS--------TITLNRTPTPTTHLSQSHHL 53
S+S+S SL L+ LTR +S PS+TRLS LP+ T++L+ + TPTTH
Sbjct: 38 STSTSPSLSLSQALLTRPSSLPSSTRLS-LPAFSGLRQSTTLSLSLSHTPTTHRRLP--- 205
Query: 54 RHTAIHASVSAPPS--TTTDSSLVEKSVNTIRFLAVDSVEKANSGHPGLPMGCAPMGHVL 111
A+ ASV+A + TTD SLV+KSVNTIRFLAVD+VEKANSGHPGLPMGCAPMGHVL
Sbjct: 206 --AAVKASVAAETAKKVTTDESLVDKSVNTIRFLAVDAVEKANSGHPGLPMGCAPMGHVL 379
Query: 112 YDEVMRYNPKNPFWFNRDRFVLSAGHGCMLQYALLHLAGYDSVKVEDLKQFRQWESRTPG 171
YDE+MRYNPKNP WFNRDRFVLSAGHGCML YALLHLAG+DSVK EDLK+FRQW SRTPG
Sbjct: 380 YDEIMRYNPKNPKWFNRDRFVLSAGHGCMLHYALLHLAGFDSVKEEDLKEFRQWGSRTPG 559
Query: 172 HPENF 176
HPENF
Sbjct: 560 HPENF 574
>AV410862
Length = 387
Score = 237 bits (605), Expect = 4e-63
Identities = 111/128 (86%), Positives = 122/128 (94%)
Frame = +2
Query: 287 VKNGNTGFDEIRAAIKEAKAVKDRPTLIKFTTTIGYGSPNKSNSYSVHGSALGAKEVDAT 346
VKNGNTG+DEIRAAIKEAKAVKD+PTLIK TTTIG+GSPNK+NSYSVHGSALGAKEVDAT
Sbjct: 2 VKNGNTGYDEIRAAIKEAKAVKDKPTLIKVTTTIGFGSPNKANSYSVHGSALGAKEVDAT 181
Query: 347 RNNLGWPYEPFHVPEDVKKHWSRHIREGAALESEWNAKFADYEKKYKEEAAVLKSIISGD 406
R NLGWP+EPFHVPE+VKKHWSRH+ EGAALE+EWNAKFA+YEKKYKE+A LKSII+G
Sbjct: 182 RKNLGWPHEPFHVPEEVKKHWSRHVPEGAALEAEWNAKFAEYEKKYKEDAEELKSIITGV 361
Query: 407 LPAGWEKA 414
LPAGWEKA
Sbjct: 362 LPAGWEKA 385
>AV409182
Length = 312
Score = 207 bits (527), Expect = 5e-54
Identities = 95/104 (91%), Positives = 100/104 (95%)
Frame = +1
Query: 190 QGIANAVGLALAEKHLAARFNKPDNEIIDHYTYCILGDGCQMEGIANEACSLAGHWGLGK 249
QGIAN VGLALAEKHLAARFNKPD+EI+DHYTY ILGDGCQMEGI+NEACSLAGHWGLGK
Sbjct: 1 QGIANGVGLALAEKHLAARFNKPDSEIVDHYTYVILGDGCQMEGISNEACSLAGHWGLGK 180
Query: 250 LIAFYDDNHISIDGDTEIAFTESVEKRFEGLGWHVIWVKNGNTG 293
LIA YDDNHISIDGDTEIAFTE+V+KRFE LGWHVIWVKNGNTG
Sbjct: 181 LIALYDDNHISIDGDTEIAFTENVDKRFEALGWHVIWVKNGNTG 312
>TC8974 similar to UP|TKTC_CRAPL (Q42676) Transketolase, chloroplast (TK)
(Fragment) , partial (23%)
Length = 706
Score = 180 bits (457), Expect(2) = 5e-53
Identities = 92/102 (90%), Positives = 97/102 (94%)
Frame = +2
Query: 634 EIAYKAGEDLRKEGKAVRVVSFVSWELFDEQSQAYKESVLPTAVTARVSIEAGSTFGWEK 693
+ A KA +DLRKEGKAVRVVSFVSWELFDEQS+AYKESVLP+AV+ARVSIEAGSTFGWEK
Sbjct: 59 KFAAKAADDLRKEGKAVRVVSFVSWELFDEQSEAYKESVLPSAVSARVSIEAGSTFGWEK 238
Query: 694 IVGSKGKAIGIDRFGASAPAGTIYKEFGITKEAVIAAAKELI 735
IVGSKGKAIGID FGASAPAG IYKEFGITKEAVIAAAKELI
Sbjct: 239 IVGSKGKAIGIDHFGASAPAGRIYKEFGITKEAVIAAAKELI 364
Score = 45.4 bits (106), Expect(2) = 5e-53
Identities = 21/21 (100%), Positives = 21/21 (100%)
Frame = +1
Query: 615 DNSTGNKPDVILIGTGSELEI 635
DNSTGNKPDVILIGTGSELEI
Sbjct: 1 DNSTGNKPDVILIGTGSELEI 63
>TC15716 similar to UP|O78327 (O78327) Transketolase 1 , partial (8%)
Length = 567
Score = 103 bits (258), Expect = 8e-23
Identities = 51/59 (86%), Positives = 56/59 (94%)
Frame = +2
Query: 676 AVTARVSIEAGSTFGWEKIVGSKGKAIGIDRFGASAPAGTIYKEFGITKEAVIAAAKEL 734
AVTARVSIEA STFGW K+VGS+GKAIG+DRFGASAPAG IYKEFGITKEAVIAAA+E+
Sbjct: 5 AVTARVSIEAASTFGWHKLVGSQGKAIGLDRFGASAPAGKIYKEFGITKEAVIAAAQEV 181
>TC11386 similar to UP|Q8LE99 (Q8LE99) Transketolase-like protein, partial
(11%)
Length = 461
Score = 93.2 bits (230), Expect = 1e-19
Identities = 57/126 (45%), Positives = 75/126 (59%), Gaps = 14/126 (11%)
Frame = +1
Query: 4 SSSTSLYLT-----NPHLTRHTSSPSTTRL---------SSLPSTITLNRTPTPTTHLSQ 49
+SS+SL+L+ N H ++SS S+ R+ S L S + TP+
Sbjct: 85 ASSSSLHLSQALAVNLHGGFNSSSSSSDRVALSNSFPAFSGLKSHSSCKHAATPSRR-RV 261
Query: 50 SHHLRHTAIHASVSAPPSTTTDSSLVEKSVNTIRFLAVDSVEKANSGHPGLPMGCAPMGH 109
T + A+ T + SLVEKSVNTIRFL++D+VEKANSGHPGLPMGCAPMGH
Sbjct: 262 GGTTTSTMVRATAVETLDKTAEVSLVEKSVNTIRFLSIDAVEKANSGHPGLPMGCAPMGH 441
Query: 110 VLYDEV 115
+LYDE+
Sbjct: 442 ILYDEI 459
>TC12640 similar to UP|Q7X9B4 (Q7X9B4) Allene oxide synthase precursor ,
partial (16%)
Length = 738
Score = 33.9 bits (76), Expect = 0.096
Identities = 19/55 (34%), Positives = 30/55 (54%), Gaps = 1/55 (1%)
Frame = +2
Query: 16 LTRHTSSPSTTRLSSLPSTITLNRTPTPTTHLSQSH-HLRHTAIHASVSAPPSTT 69
L + + P RL+SL S +N+ PT + Q H H+ +IH S+ PP++T
Sbjct: 77 LCTNITIPFHQRLNSLVSISQVNKRPTLLFNCQQLHTHIYTHSIHISIKTPPNST 241
>TC17983 similar to UP|Q9XEX5 (Q9XEX5) Cytidine deaminase , partial (42%)
Length = 519
Score = 32.7 bits (73), Expect = 0.21
Identities = 28/97 (28%), Positives = 38/97 (38%)
Frame = +2
Query: 2 SSSSSTSLYLTNPHLTRHTSSPSTTRLSSLPSTITLNRTPTPTTHLSQSHHLRHTAIHAS 61
SSS STS L +P T S+PS++ + PST TP+P
Sbjct: 200 SSSESTSNSLASPSTT--PSTPSSSSSPTSPSTTKQISTPSP------------------ 319
Query: 62 VSAPPSTTTDSSLVEKSVNTIRFLAVDSVEKANSGHP 98
S PP T +S + S RF + + HP
Sbjct: 320 -SPPPPVATAASSSKNSAEPRRFKSSSPQNPTRNSHP 427
>TC8853 similar to PIR|T09217|T09217 protein sam2B - spinach {Spinacia
oleracea;}, partial (43%)
Length = 1157
Score = 32.7 bits (73), Expect = 0.21
Identities = 27/90 (30%), Positives = 42/90 (46%), Gaps = 7/90 (7%)
Frame = -3
Query: 1 MSSSSSTSLYLTNPHLTRH-------TSSPSTTRLSSLPSTITLNRTPTPTTHLSQSHHL 53
+SS SS+SL + P+L H T PS+ LSS + P+ T ++ H
Sbjct: 429 LSSLSSSSLLQSPPYLLHHHHQPDSATPPPSSPALSSPQTHQLAAAFPSSTHPHPKTLHP 250
Query: 54 RHTAIHASVSAPPSTTTDSSLVEKSVNTIR 83
RH+ S+P ST++ + S T+R
Sbjct: 249 RHSR*SRRRSSPNSTSSHAPPPRGSATTLR 160
>TC10239 similar to UP|Q7NEZ8 (Q7NEZ8) Glr3729 protein, partial (8%)
Length = 629
Score = 32.3 bits (72), Expect = 0.28
Identities = 26/71 (36%), Positives = 37/71 (51%), Gaps = 4/71 (5%)
Frame = -3
Query: 3 SSSSTSLYLTNPHLTRHTSSPSTTRLSSLPSTITLNR----TPTPTTHLSQSHHLRHTAI 58
S SS+SL T+P+ T +P T SS P ++T R TP P+TH S +
Sbjct: 429 SKSSSSLSPTSPN---PTPAPPPTASSSKPPSLTPTRPFRLTPIPSTHGS---------L 286
Query: 59 HASVSAPPSTT 69
A ++PPST+
Sbjct: 285 PAPKTSPPSTS 253
>AV407559
Length = 429
Score = 32.3 bits (72), Expect = 0.28
Identities = 21/70 (30%), Positives = 34/70 (48%)
Frame = +1
Query: 15 HLTRHTSSPSTTRLSSLPSTITLNRTPTPTTHLSQSHHLRHTAIHASVSAPPSTTTDSSL 74
H + T ++++L S PST +R S S + H+A H ++S P T + SL
Sbjct: 28 HCHQPTPIATSSQLRSPPSTHRRHRGMNE*MTSSSSTNHSHSAHHHALSLNPQTLSSPSL 207
Query: 75 VEKSVNTIRF 84
+ S+ T F
Sbjct: 208LHSSIQTTPF 237
>TC19436 similar to UP|Q93Y06 (Q93Y06) Receptor protein kinase-like protein,
partial (19%)
Length = 570
Score = 32.0 bits (71), Expect = 0.36
Identities = 32/107 (29%), Positives = 49/107 (44%), Gaps = 4/107 (3%)
Frame = +2
Query: 4 SSSTSLYLTNPHLTRHTSSPSTTRLSSLPSTITLNRTPTPT--THLSQ-SHHLRHTAIHA 60
+SS S T+P +PS SS+ S +P P+ +HLSQ S+H TA +
Sbjct: 248 ASSASSSKTSPSPAPSPRTPSRASTSSVSSRSATTPSPDPSLISHLSQTSNHSSSTATTS 427
Query: 61 -SVSAPPSTTTDSSLVEKSVNTIRFLAVDSVEKANSGHPGLPMGCAP 106
++S S+ + +S + +S T V S + S LP G P
Sbjct: 428 PALSRRQSSPSTASSLSRSATTTS--PVSSRFSSTSXDXSLPSGWTP 562
Score = 27.7 bits (60), Expect = 6.9
Identities = 15/35 (42%), Positives = 21/35 (59%)
Frame = +2
Query: 2 SSSSSTSLYLTNPHLTRHTSSPSTTRLSSLPSTIT 36
+S+ S+S T+P L+R SSPST S +T T
Sbjct: 392 TSNHSSSTATTSPALSRRQSSPSTASSLSRSATTT 496
>TC11199
Length = 743
Score = 31.2 bits (69), Expect = 0.62
Identities = 26/70 (37%), Positives = 40/70 (57%), Gaps = 3/70 (4%)
Frame = +1
Query: 2 SSSSSTSLYLTNPHLTRHTSSPSTTRLSSLPSTITLNRTPTPTTHLSQSHHLRHTAIHA- 60
+++S+TSL +P R S+ TTR SS PSTI + +PT++ S L T +
Sbjct: 205 TTTSATSLAPKSPRSKR--SARRTTRTSSSPSTI--SDPSSPTSNPSSPPSLTPTPSSSP 372
Query: 61 --SVSAPPST 68
++S+PPST
Sbjct: 373 SLALSSPPST 402
>TC12462 weakly similar to UP|Q8TW78 (Q8TW78) Uncharacterized membrane
protein specific for M.kandleri, MK-9 family, partial
(6%)
Length = 812
Score = 31.2 bits (69), Expect = 0.62
Identities = 24/64 (37%), Positives = 29/64 (44%), Gaps = 6/64 (9%)
Frame = +3
Query: 17 TRHTSSPS-TTRLSSLPSTITLNRTPTPTTHLSQ-----SHHLRHTAIHASVSAPPSTTT 70
T+HTSSPS R S S + TP PT L HH R A H + + PP+
Sbjct: 123 TKHTSSPSRLPRFSLSLSRCHSSTTPPPTLSLKHKLQPAQHHHRRPATHCT-TPPPTQPL 299
Query: 71 DSSL 74
SL
Sbjct: 300 SLSL 311
>TC10026
Length = 599
Score = 31.2 bits (69), Expect = 0.62
Identities = 26/73 (35%), Positives = 35/73 (47%), Gaps = 2/73 (2%)
Frame = +3
Query: 4 SSSTSLYL--TNPHLTRHTSSPSTTRLSSLPSTITLNRTPTPTTHLSQSHHLRHTAIHAS 61
+S TS+Y T PH T S T S LPS+ +L T + H S T+ +
Sbjct: 57 TSETSIYPSPTTPHSPELTYSTLPTPESPLPSSPSLYLTRSKPPHSRAS-----TSPTPT 221
Query: 62 VSAPPSTTTDSSL 74
+S+PPS TT L
Sbjct: 222 LSSPPSMTTTRPL 260
>TC8112 similar to UP|ODPA_PEA (P52902) Pyruvate dehydrogenase E1 component
alpha subunit, mitochondrial precursor (PDHE1-A) ,
partial (90%)
Length = 1816
Score = 30.8 bits (68), Expect = 0.81
Identities = 34/161 (21%), Positives = 69/161 (42%), Gaps = 4/161 (2%)
Frame = +3
Query: 186 GPLGQGIANAVGLALAEKHLAARFNKPDNEIIDHYTYCILGDGCQMEGIANEACSLAGHW 245
G +G + G+A A+K+ D + T+ + GDG +G EA ++A W
Sbjct: 639 GIVGAQVPLGCGVAFAQKY------SKDEAV----TFALYGDGAANQGQLFEALNIAALW 788
Query: 246 GLGKLIAFYDDNHISIDGDTEIAFTESVEKRFEGLGWHVIWVKNGNTGFDEIRAAIKEAK 305
L ++ ++++ + A + + KR G +V +K ++ A K AK
Sbjct: 789 DLPAILVCENNHYGMGTAEWRAAKSPAYYKR----GDYVPGLKVDGMDVLAVKQACKFAK 956
Query: 306 --AVKDRPTLIKFTT--TIGYGSPNKSNSYSVHGSALGAKE 342
A+K+ P +++ T G+ + ++Y G ++
Sbjct: 957 EHALKNGPIILEMDTYRYHGHSMSDPGSTYRTRDEISGVRQ 1079
>BP033767
Length = 541
Score = 30.8 bits (68), Expect = 0.81
Identities = 27/82 (32%), Positives = 37/82 (44%), Gaps = 8/82 (9%)
Frame = +2
Query: 2 SSSSSTSLYLTN-----PHLTRHTSSPSTTRLSSLPSTITLNRTPTPTTHLSQSHHLRH- 55
S +SS +L T+ P T HT PST +TIT++ T P + H H
Sbjct: 92 SCTSSATLIRTHHQQEDPPPTLHTRLPSTLSTPKPTTTITISLTSKPKIPIFLHPHNHHH 271
Query: 56 -TAIHASV-SAPPSTTTDSSLV 75
+H + P STTT SL+
Sbjct: 272 PKCLHTFLPHIPISTTTSFSLL 337
>TC18463 similar to AAR92278 (AAR92278) At4g22580, partial (34%)
Length = 555
Score = 30.8 bits (68), Expect = 0.81
Identities = 19/64 (29%), Positives = 31/64 (47%)
Frame = +3
Query: 20 TSSPSTTRLSSLPSTITLNRTPTPTTHLSQSHHLRHTAIHASVSAPPSTTTDSSLVEKSV 79
++S ST SS+P++ + R TP+T L+ + A P TT +S V ++
Sbjct: 276 SNSSSTVACSSIPASPPIQRRQTPSTSLTTPASILSGISTARSITPAPTTASTSSVSFAM 455
Query: 80 NTIR 83
T R
Sbjct: 456 MTTR 467
>TC9679 weakly similar to UP|Q8H1B5 (Q8H1B5) Hin1-like protein, partial
(54%)
Length = 710
Score = 30.4 bits (67), Expect = 1.1
Identities = 16/53 (30%), Positives = 21/53 (39%)
Frame = +1
Query: 14 PHLTRHTSSPSTTRLSSLPSTITLNRTPTPTTHLSQSHHLRHTAIHASVSAPP 66
P ++ P R L + L+ P THL + HH RH H PP
Sbjct: 100 PSAQKNLPPPKPRRRRMLRVLLRLHLQPHLQTHLHRPHHHRHRRPHLLAHRPP 258
Database: LJGI
Posted date: Jul 30, 2004 11:16 AM
Number of letters in database: 14,692,800
Number of sequences in database: 28,460
Lambda K H
0.315 0.133 0.395
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 12,134,473
Number of Sequences: 28460
Number of extensions: 171324
Number of successful extensions: 1315
Number of sequences better than 10.0: 87
Number of HSP's better than 10.0 without gapping: 1249
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 1301
length of query: 735
length of database: 4,897,600
effective HSP length: 97
effective length of query: 638
effective length of database: 2,136,980
effective search space: 1363393240
effective search space used: 1363393240
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 16 ( 7.3 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.6 bits)
S2: 59 (27.3 bits)
Medicago: description of AC119419.5


