

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
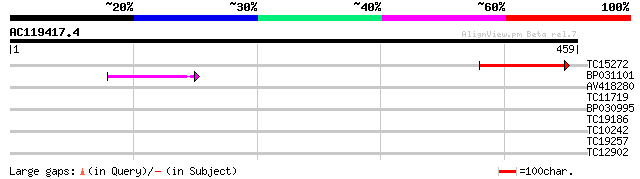
Query= AC119417.4 - phase: 0 /pseudo
(459 letters)
Database: LJGI
28,460 sequences; 14,692,800 total letters
Searching..................................................done
Score E
Sequences producing significant alignments: (bits) Value
TC15272 weakly similar to UP|Q9FUJ0 (Q9FUJ0) Cytokinin oxidase, ... 76 1e-14
BP031101 42 3e-04
AV418280 30 0.64
TC11719 29 1.4
BP030995 29 1.4
TC19186 homologue to emb|Z18289.1|PPC16SRNA Palmaria palmata Chl... 27 5.4
TC10242 weakly similar to UP|O00561 (O00561) Calcium/calmodulin-... 27 7.1
TC19257 UP|Q9BDE4 (Q9BDE4) Selenoprotein P-like protein (Fragmen... 27 7.1
TC12902 27 9.2
>TC15272 weakly similar to UP|Q9FUJ0 (Q9FUJ0) Cytokinin oxidase, partial
(14%)
Length = 524
Score = 75.9 bits (185), Expect = 1e-14
Identities = 31/73 (42%), Positives = 52/73 (70%)
Frame = +3
Query: 381 SSSGPDELEHILSQNKRILEYCERAHLGVKQYLPHYTTQEEWQTHYGHKWEIFKQRKSIY 440
+++G DE++ +QN+ IL++C+ A + +K+YL T EEW H+G KW++++ RK+ +
Sbjct: 117 AANGGDEVQKYQAQNQEILQFCKGAGIKIKEYLTGNKTHEEWVEHFGPKWKLWEDRKAEF 296
Query: 441 DPLAILAPGQGIF 453
DP IL+PG GIF
Sbjct: 297 DPKRILSPGPGIF 335
>BP031101
Length = 472
Score = 41.6 bits (96), Expect = 3e-04
Identities = 29/74 (39%), Positives = 39/74 (52%)
Frame = +3
Query: 80 ENFLMWMCQEVNYG*RFCMRH*SMVWHQDLGQIICI*QLVVLYPMLVLVVRLLGMVHRSV 139
E+ M M E N+G C+ SM H LG I CI* L+PML V R MV +S
Sbjct: 207 ESSTMLMLVESNFGLMCCVPPWSMDLHLFLGLITCI*LWEGLFPMLA*VARHSNMVLKSQ 386
Query: 140 MFRRWRLLQSEEQN 153
+F +W + E++N
Sbjct: 387 VFLKW-MSSPEKEN 425
>AV418280
Length = 398
Score = 30.4 bits (67), Expect = 0.64
Identities = 23/80 (28%), Positives = 36/80 (44%), Gaps = 5/80 (6%)
Frame = -2
Query: 309 SKGLWDVPHPWLNLFI--PKSKIHNFAEVVFGNIVKETSNGPVLIY-PVHKSKWDKRTSV 365
S G+W H WL + + P S + NFA+ K G + I+ S W +R S+
Sbjct: 328 SMGVWQALHRWLGISVALPASTLANFAQFSITARNKNQRLGELAIWIATVWSLWIQRNSI 149
Query: 366 VIPDE--DIFYLVGFLASSS 383
+ + D YL+ + S S
Sbjct: 148 IFRNNALDHSYLLDLIQSRS 89
>TC11719
Length = 376
Score = 29.3 bits (64), Expect = 1.4
Identities = 11/51 (21%), Positives = 27/51 (52%)
Frame = -2
Query: 388 LEHILSQNKRILEYCERAHLGVKQYLPHYTTQEEWQTHYGHKWEIFKQRKS 438
++H+L++N R+ C+ H+ +K+ ++W+ +++KQ S
Sbjct: 303 VKHVLTENARV*NLCQPEHIYLKKKQKQTKQMQDWKKITMQNLKVWKQNAS 151
>BP030995
Length = 458
Score = 29.3 bits (64), Expect = 1.4
Identities = 11/51 (21%), Positives = 27/51 (52%)
Frame = +3
Query: 388 LEHILSQNKRILEYCERAHLGVKQYLPHYTTQEEWQTHYGHKWEIFKQRKS 438
++H+L++N R+ C+ H+ +K+ ++W+ +++KQ S
Sbjct: 72 VKHVLTENARV*NLCQPEHIYLKKKQKQTKQMQDWKKITMQNLKVWKQNAS 224
>TC19186 homologue to emb|Z18289.1|PPC16SRNA Palmaria palmata Chloroplast
for 16SrRNA gene, partial (10%)
Length = 563
Score = 27.3 bits (59), Expect = 5.4
Identities = 12/26 (46%), Positives = 16/26 (61%)
Frame = +2
Query: 372 IFYLVGFLASSSGPDELEHILSQNKR 397
I+YL+ S P EL+H+ SQ KR
Sbjct: 107 IYYLLNL*TRKSKPGELKHLSSQRKR 184
>TC10242 weakly similar to UP|O00561 (O00561) Calcium/calmodulin-dependent
protein kinase isoform A (Fragment), partial (6%)
Length = 522
Score = 26.9 bits (58), Expect = 7.1
Identities = 22/79 (27%), Positives = 37/79 (45%)
Frame = +2
Query: 259 YFNMEETLEVNQDIQKHLSHLNFIPSTLFQTEVTYVDFLDRVHISEVKLRSKGLWDVPHP 318
Y+ EET ++ I + S + L T+ ++DF+ + K R + HP
Sbjct: 14 YYANEETDQLEYIIPEESS----LEQHLQVTDAMFIDFVKYLLNINPKRRPTTKQALKHP 181
Query: 319 WLNLFIPKSKIHNFAEVVF 337
WL+ + KSK H F+ +F
Sbjct: 182 WLS-HVYKSK*HCFSGGMF 235
>TC19257 UP|Q9BDE4 (Q9BDE4) Selenoprotein P-like protein (Fragment), partial
(9%)
Length = 525
Score = 26.9 bits (58), Expect = 7.1
Identities = 17/66 (25%), Positives = 32/66 (47%), Gaps = 2/66 (3%)
Frame = -2
Query: 104 VWHQDLGQIIC-I*QLVVLYPMLVLVVRLLGMVHRSVMFRRWRLLQ-SEEQNGELFYSVL 161
+WH LG ++C + L P+ +L++ +L S+ +W L Q + + +S +
Sbjct: 404 IWHLQLGLLLCFLLDTTTLIPIPILILLML-----SL*LTQWHLQQCTNNKETNFVFSFI 240
Query: 162 GGLGQF 167
L QF
Sbjct: 239 ACLDQF 222
>TC12902
Length = 714
Score = 26.6 bits (57), Expect = 9.2
Identities = 8/12 (66%), Positives = 9/12 (74%)
Frame = -1
Query: 414 PHYTTQEEWQTH 425
PHYT Q +W TH
Sbjct: 123 PHYTPQHQWSTH 88
Database: LJGI
Posted date: Jul 30, 2004 11:16 AM
Number of letters in database: 14,692,800
Number of sequences in database: 28,460
Lambda K H
0.336 0.148 0.478
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 8,600,052
Number of Sequences: 28460
Number of extensions: 125840
Number of successful extensions: 898
Number of sequences better than 10.0: 18
Number of HSP's better than 10.0 without gapping: 886
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 898
length of query: 459
length of database: 4,897,600
effective HSP length: 93
effective length of query: 366
effective length of database: 2,250,820
effective search space: 823800120
effective search space used: 823800120
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 15 ( 7.3 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 39 (21.7 bits)
S2: 57 (26.6 bits)
Medicago: description of AC119417.4


