

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC119417.2 + phase: 0
(898 letters)
Database: LJGI
28,460 sequences; 14,692,800 total letters
Searching..................................................done
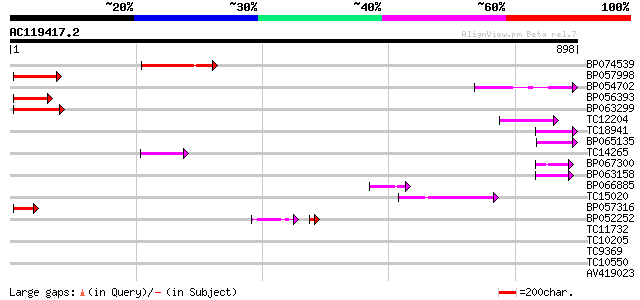
Score E
Sequences producing significant alignments: (bits) Value
BP074539 100 8e-22
BP057998 84 8e-17
BP054702 81 8e-16
BP056393 79 3e-15
BP063299 74 1e-13
TC12204 weakly similar to UP|Q9ZSN3 (Q9ZSN3) NBS-LRR-like protei... 52 5e-07
TC18941 weakly similar to UP|Q8GZF0 (Q8GZF0) Resistance protein ... 49 4e-06
BP065135 49 4e-06
TC14265 weakly similar to UP|AAN62760 (AAN62760) Disease resista... 48 6e-06
BP067300 45 7e-05
BP063158 44 9e-05
BP066885 43 2e-04
TC15020 homologue to UP|Q9QX99 (Q9QX99) Transcription factor (T-... 43 3e-04
BP057316 43 3e-04
BP052252 37 7e-04
TC11732 39 0.005
TC10205 weakly similar to UP|Q8RL77 (Q8RL77) HpaG, partial (10%) 38 0.006
TC9369 similar to UP|Q93X72 (Q93X72) Leucine-rich repeat resista... 38 0.006
TC10550 weakly similar to UP|Q40392 (Q40392) Virus resistance (N... 37 0.018
AV419023 37 0.018
>BP074539
Length = 357
Score = 100 bits (250), Expect = 8e-22
Identities = 59/120 (49%), Positives = 80/120 (66%)
Frame = +1
Query: 210 LVQLRYLNLSHTDIKCLPDTTCDLYYLQTLLLSGCWKLIELPIHVGKLINLRHLDISYTK 269
L L YL+LSHT+I+ LPD+TC LY LQ L L C L ELP+++ KL NLR+LD S T+
Sbjct: 4 LKHLCYLDLSHTNIEKLPDSTCLLYNLQILKLKNCRYLKELPLNLHKLTNLRYLDFSGTQ 183
Query: 270 IKKMPMQIVRLENLQTLTVFLVGKQKVGLSIRELGKFPNLRGKLCIKNLQNAIDVSEACD 329
++KMPM +L+NLQ L F VG Q +I++L + NL+G L I LQN I+ +A +
Sbjct: 184 VRKMPMHFGKLKNLQVLNSFFVG-QGSEYNIQQLDEL-NLQGTLSISELQNIINPLDALE 357
>BP057998
Length = 566
Score = 84.3 bits (207), Expect = 8e-17
Identities = 39/77 (50%), Positives = 52/77 (66%), Gaps = 2/77 (2%)
Frame = +2
Query: 7 ILNSDIWNIPNDN--IMPSLFLTYQHLPSHLKRCFAYCSIFPKGYPFNRKKLILLWMAEG 64
+LN DIW + I+PSL ++Y +LPS+LKRCFAY S++PK Y F + +ILLWMAE
Sbjct: 335 VLNCDIWELSESESKIIPSLRISYHYLPSYLKRCFAYFSLYPKDYEFEKNDVILLWMAED 514
Query: 65 FLEHSMVGKAVEEVGDD 81
L GK + EVGD+
Sbjct: 515 LLPPPKQGKTLNEVGDE 565
>BP054702
Length = 528
Score = 80.9 bits (198), Expect = 8e-16
Identities = 55/166 (33%), Positives = 79/166 (47%), Gaps = 3/166 (1%)
Frame = -3
Query: 736 FQSLCFLSDLHI---GGDNIVNTLLKKKLLPPLLVSLTITNLTEMMRLKGNRLQHISTLK 792
F L LS L++ G +++NTLLK++LLP LV L + N + + L+GN LQH+S+L
Sbjct: 487 FPHLTSLSHLYVFGFGDQDLMNTLLKERLLPTSLVFLGLYNFSSLKFLEGNGLQHLSSLH 308
Query: 793 NLSFKCCSTLETCKDFFPSFLKSLVFINCPKLMSLPDMFPSSLETLEFDDCPRLGLLPRS 852
L CP L+SLP+
Sbjct: 307 QLHID----------------------KCPSLVSLPE----------------------D 260
Query: 853 GFPSSLKLLSISHCPLLKSRYATKRNDHLSKISHIPVVKINGIVTI 898
P SL +L++ CPLL++RY ++ H SK+S IPV+KIN V I
Sbjct: 259 QLPHSLAVLNLEKCPLLEARYRIRKGKHWSKVSRIPVIKINEEVII 122
>BP056393
Length = 421
Score = 79.0 bits (193), Expect = 3e-15
Identities = 36/63 (57%), Positives = 47/63 (74%), Gaps = 2/63 (3%)
Frame = +1
Query: 7 ILNSDIWNIPND--NIMPSLFLTYQHLPSHLKRCFAYCSIFPKGYPFNRKKLILLWMAEG 64
ILNS+IW +P + NI+ +L ++Y +LP HLKRC YCS+FPK Y F+ +LILLWMAE
Sbjct: 223 ILNSNIWELPENESNIILALRISYHYLPPHLKRCXVYCSLFPKDYKFDXDELILLWMAED 402
Query: 65 FLE 67
LE
Sbjct: 403 LLE 411
>BP063299
Length = 580
Score = 73.9 bits (180), Expect = 1e-13
Identities = 35/82 (42%), Positives = 54/82 (65%), Gaps = 1/82 (1%)
Frame = +3
Query: 7 ILNSDIWNIPNDN-IMPSLFLTYQHLPSHLKRCFAYCSIFPKGYPFNRKKLILLWMAEGF 65
+ S +WN+P++ I+P+L L+Y HL L++CFA+C+IFPK ++ LI LWMA GF
Sbjct: 309 VKESTLWNLPDERCILPALMLSYFHLTPTLRQCFAFCAIFPKDAKIIKEDLIYLWMANGF 488
Query: 66 LEHSMVGKAVEEVGDDYFNELL 87
+ S K VE+VG+ + E +
Sbjct: 489 IS-STKNKEVEDVGNSIWKECI 551
>TC12204 weakly similar to UP|Q9ZSN3 (Q9ZSN3) NBS-LRR-like protein cD7
(Fragment), partial (3%)
Length = 328
Score = 51.6 bits (122), Expect = 5e-07
Identities = 33/99 (33%), Positives = 46/99 (46%), Gaps = 5/99 (5%)
Frame = +1
Query: 776 EMMRLKGNRLQHISTLKNLSFKCCSTLETCKD--FFPSFLKSLVFINCPKLMSLPDMFPS 833
E + + G +LQ++ +L +L C E+ + + +L C KL S P
Sbjct: 10 ESLVVTGVQLQYLQSLNSLRICNCPNFESFPEGGLRAPNMTNLHLEKCKKLKSFPQQMNK 189
Query: 834 ---SLETLEFDDCPRLGLLPRSGFPSSLKLLSISHCPLL 869
SL TL +CP L +P GFP SL LL I HC L
Sbjct: 190 MLLSLMTLNIKECPELESIPEGGFPDSLNLLEIFHCAKL 306
Score = 32.0 bits (71), Expect = 0.45
Identities = 23/82 (28%), Positives = 36/82 (43%)
Frame = +1
Query: 633 LTSFQLNGFPVLQILSIEGCSSLKSIFISEKNSSLSLSTLQSLKVSNCKSLRSLPQRMDT 692
+T QL L L I C + +S L + +L + CK L+S PQ+M+
Sbjct: 22 VTGVQLQYLQSLNSLRICNCPNFESF----PEGGLRAPNMTNLHLEKCKKLKSFPQQMNK 189
Query: 693 LFVLKSLTLDKLSLCCEVACLP 714
+ L SL + C E+ +P
Sbjct: 190 M--LLSLMTLNIKECPELESIP 249
>TC18941 weakly similar to UP|Q8GZF0 (Q8GZF0) Resistance protein KR4,
partial (3%)
Length = 569
Score = 48.9 bits (115), Expect = 4e-06
Identities = 28/66 (42%), Positives = 37/66 (55%)
Frame = +2
Query: 833 SSLETLEFDDCPRLGLLPRSGFPSSLKLLSISHCPLLKSRYATKRNDHLSKISHIPVVKI 892
+SLE LE P+L LP G PSSL +L I CPLL++ + KI+HIP + I
Sbjct: 71 TSLEKLEICYSPQLESLPEEGLPSSLSVLIIXPCPLLEASCHSHGGKECPKIAHIPCIII 250
Query: 893 NGIVTI 898
+ V I
Sbjct: 251 HRPVII 268
Score = 37.4 bits (85), Expect = 0.011
Identities = 26/86 (30%), Positives = 43/86 (49%), Gaps = 1/86 (1%)
Frame = +2
Query: 766 LVSLTITNLTEMMRLKGNRLQHISTLKNLSFKCCSTLETC-KDFFPSFLKSLVFINCPKL 824
LVSL I NL ++ L G LQH+++L+ L LE+ ++ PS L L+ CP
Sbjct: 2 LVSLVICNLHDVKCLGGIWLQHLTSLEKLEICYSPQLESLPEEGLPSSLSVLIIXPCP-- 175
Query: 825 MSLPDMFPSSLETLEFDDCPRLGLLP 850
+ +S + +CP++ +P
Sbjct: 176 -----LLEASCHSHGGKECPKIAHIP 238
>BP065135
Length = 492
Score = 48.9 bits (115), Expect = 4e-06
Identities = 27/65 (41%), Positives = 36/65 (54%), Gaps = 1/65 (1%)
Frame = -3
Query: 835 LETLEFDDCPRLGLLPRSGFP-SSLKLLSISHCPLLKSRYATKRNDHLSKISHIPVVKIN 893
L L D CP L LP ++L + S+ CP L++RY + H SK+S IPV+KIN
Sbjct: 244 LHQLHIDQCPSLVALPEDATGHTALAV*SVEKCP*LEARYQIGKGKHWSKVSRIPVIKIN 65
Query: 894 GIVTI 898
V I
Sbjct: 64 EEVII 50
Score = 28.1 bits (61), Expect = 6.5
Identities = 17/32 (53%), Positives = 23/32 (71%), Gaps = 4/32 (12%)
Frame = -1
Query: 736 FQSLCFLSDLH---IGGD-NIVNTLLKKKLLP 763
F+ L LS L+ GGD ++VNTLLK++LLP
Sbjct: 423 FRHLTSLSHLYAFGFGGDQDLVNTLLKERLLP 328
>TC14265 weakly similar to UP|AAN62760 (AAN62760) Disease resistance
protein-like protein MsR1, partial (5%)
Length = 766
Score = 48.1 bits (113), Expect = 6e-06
Identities = 32/78 (41%), Positives = 43/78 (55%), Gaps = 1/78 (1%)
Frame = +1
Query: 207 INKLVQLRYLNLSH-TDIKCLPDTTCDLYYLQTLLLSGCWKLIELPIHVGKLINLRHLDI 265
I KL L L LS TD+K LPD+ L L+ L +S C L LP +G L NL+ L +
Sbjct: 7 IGKLENLELLRLSSCTDLKGLPDSIGRLSKLRLLDISNCISLPSLPEEIGNLCNLKSLYM 186
Query: 266 SYTKIKKMPMQIVRLENL 283
+ ++P IV L+NL
Sbjct: 187 TSCAGCELPSSIVNLQNL 240
Score = 39.7 bits (91), Expect = 0.002
Identities = 24/64 (37%), Positives = 36/64 (55%)
Frame = +1
Query: 644 LQILSIEGCSSLKSIFISEKNSSLSLSTLQSLKVSNCKSLRSLPQRMDTLFVLKSLTLDK 703
L++L + C+ LK + +S LS L+ L +SNC SL SLP+ + L LKSL +
Sbjct: 25 LELLRLSSCTDLKGL----PDSIGRLSKLRLLDISNCISLPSLPEEIGNLCNLKSLYMTS 192
Query: 704 LSLC 707
+ C
Sbjct: 193 CAGC 204
>BP067300
Length = 523
Score = 44.7 bits (104), Expect = 7e-05
Identities = 23/61 (37%), Positives = 37/61 (59%)
Frame = -2
Query: 833 SSLETLEFDDCPRLGLLPRSGFPSSLKLLSISHCPLLKSRYATKRNDHLSKISHIPVVKI 892
+SLETL CP+L +P + P S+ L I P L+ R ++++ KI+HIP+++I
Sbjct: 489 TSLETLGIACCPKLQCMP-AKLPCSISTLHIVRSPRLEERCRGRKSEDWPKIAHIPMIRI 313
Query: 893 N 893
N
Sbjct: 312 N 310
>BP063158
Length = 431
Score = 44.3 bits (103), Expect = 9e-05
Identities = 25/61 (40%), Positives = 30/61 (48%), Gaps = 1/61 (1%)
Frame = -1
Query: 834 SLETLEFDDCPRLGLLPRSGFPSSLKLLSISH-CPLLKSRYATKRNDHLSKISHIPVVKI 892
SL L DC L LP G P S+ LSI CPLLK R + KISH ++ I
Sbjct: 329 SLTELRLSDCSGLPCLPEEGLPKSISTLSIRRMCPLLKQRCQKPHGEDWGKISHSKLIFI 150
Query: 893 N 893
+
Sbjct: 149 D 147
>BP066885
Length = 469
Score = 43.1 bits (100), Expect = 2e-04
Identities = 26/65 (40%), Positives = 38/65 (58%)
Frame = -1
Query: 571 TCLQHLDLIYISSLTAFPANGLPTSLQSLRIDECQNLAFLRPETWSNYTSLVTLELKNCC 630
TCLQ L + SS +FP + LP SL+SL I + + L F + + + L +L++ N C
Sbjct: 388 TCLQGLMIWSCSSALSFPGDCLPASLKSLYIRDFRELEFPK-QNQQXHELLESLDIDNSC 212
Query: 631 DSLTS 635
DSL S
Sbjct: 211 DSLIS 197
>TC15020 homologue to UP|Q9QX99 (Q9QX99) Transcription factor (T-cell
leukemia, homeobox 1), partial (5%)
Length = 1304
Score = 42.7 bits (99), Expect = 3e-04
Identities = 42/159 (26%), Positives = 72/159 (44%), Gaps = 1/159 (0%)
Frame = +1
Query: 616 SNYTSLVTLEL-KNCCDSLTSFQLNGFPVLQILSIEGCSSLKSIFISEKNSSLSLSTLQS 674
S T LVTL+L +N L+ L+ L++ S SI S +L +L S
Sbjct: 292 SQLTQLVTLDLAENSFSGPIPSSLSSLSNLKTLTLRSNSFSGSI----PPSITTLKSLYS 459
Query: 675 LKVSNCKSLRSLPQRMDTLFVLKSLTLDKLSLCCEVACLPPKLQFMHIESLGLATPVTEW 734
L S+ SLP M++L L+ + L L + LPP L + +++ L+ P+ +
Sbjct: 460 LDFSHNSLSGSLPNSMNSLINLRRIDLSFNKLTGSIPKLPPNLLELAVKANSLSGPLQKA 639
Query: 735 GFQSLCFLSDLHIGGDNIVNTLLKKKLLPPLLVSLTITN 773
F+ L L + + + + T+ L P L + ++N
Sbjct: 640 TFEGLNQLEVVELSENTLAGTVEPWFFLLPSLQQVDLSN 756
>BP057316
Length = 494
Score = 42.7 bits (99), Expect = 3e-04
Identities = 17/41 (41%), Positives = 28/41 (67%), Gaps = 2/41 (4%)
Frame = +2
Query: 7 ILNSDIWNI--PNDNIMPSLFLTYQHLPSHLKRCFAYCSIF 45
+L D W + D+I+P L L+YQ+L +++CFAYCS++
Sbjct: 371 VLQGDFWKLCEDEDSIIPVLKLSYQNLSPQIRQCFAYCSLY 493
>BP052252
Length = 472
Score = 37.4 bits (85), Expect(2) = 7e-04
Identities = 30/75 (40%), Positives = 37/75 (49%)
Frame = -2
Query: 383 SWLGDCSFSNMVYLSIKSCEYCITLPPLGQVPFLKELKIDGMSRVETIGPEFYGMTGGST 442
SWL +N+V +S+ C T+PPL +PFLK L I +E I Y G T
Sbjct: 453 SWL-----ANIVEISLIKCNTXRTIPPLEHLPFLKSLFICYHGELEFI----YFQDGRRT 301
Query: 443 NSPFQPFPSLEKLEF 457
FPSLEKL F
Sbjct: 300 F-----FPSLEKLCF 271
Score = 23.1 bits (48), Expect(2) = 7e-04
Identities = 9/15 (60%), Positives = 11/15 (73%)
Frame = -3
Query: 476 FPRLKTLMLRDCTEL 490
FPRL L +R+C EL
Sbjct: 179 FPRLSHLSIRNCEEL 135
>TC11732
Length = 530
Score = 38.5 bits (88), Expect = 0.005
Identities = 28/99 (28%), Positives = 58/99 (58%), Gaps = 2/99 (2%)
Frame = +2
Query: 192 MLSLSNYNITVLPNSINKLVQ-LRYLNLSHTDIKCLPDTTCDLYYLQTLLLSGCWKLIE- 249
+++L + + P+ I L + +R L+L+H I +P L +Q L+L+ LIE
Sbjct: 188 IVALRDSKLKTFPDEILDLDRSVRTLDLTHNRIVDIPLEISKLVNVQRLVLAD--NLIER 361
Query: 250 LPIHVGKLINLRHLDISYTKIKKMPMQIVRLENLQTLTV 288
LP+++GKL +L+ +++ +I +P ++ +L L+ L++
Sbjct: 362 LPVNLGKLQSLKLVNLDGNRITSLPDELGQLVRLERLSI 478
>TC10205 weakly similar to UP|Q8RL77 (Q8RL77) HpaG, partial (10%)
Length = 586
Score = 38.1 bits (87), Expect = 0.006
Identities = 22/63 (34%), Positives = 36/63 (56%), Gaps = 1/63 (1%)
Frame = +3
Query: 203 LPNSINKLVQLRYLNLSHT-DIKCLPDTTCDLYYLQTLLLSGCWKLIELPIHVGKLINLR 261
LP SI L Q+++L++S + LP+ +L L+ L + GC +L ELP + L NL
Sbjct: 9 LPKSITNLEQVQFLDISDCISLSHLPEDMGELKKLEKLNVRGCSRLTELPTSIMDLENLE 188
Query: 262 HLD 264
++
Sbjct: 189 EVE 197
>TC9369 similar to UP|Q93X72 (Q93X72) Leucine-rich repeat resistance
protein-like protein, partial (70%)
Length = 859
Score = 38.1 bits (87), Expect = 0.006
Identities = 31/100 (31%), Positives = 49/100 (49%), Gaps = 2/100 (2%)
Frame = +1
Query: 184 IPSIKRLRMLSLS-NYNITVLPNSINKLVQLRYLNLSHTDIKC-LPDTTCDLYYLQTLLL 241
I +KRL++L+L N +P I +L L +L LS K +P +L L+ L L
Sbjct: 67 IGRLKRLKILNLRWNKLQDAIPPEIGELKSLTHLYLSFNSFKGEIPKELANLPDLRYLYL 246
Query: 242 SGCWKLIELPIHVGKLINLRHLDISYTKIKKMPMQIVRLE 281
+ +P +G L NLRHLD + +++R+E
Sbjct: 247 HENRLIGRIPPELGTLQNLRHLDAGNNHLVGTIRELIRIE 366
>TC10550 weakly similar to UP|Q40392 (Q40392) Virus resistance (N), partial
(5%)
Length = 670
Score = 36.6 bits (83), Expect = 0.018
Identities = 25/71 (35%), Positives = 37/71 (51%), Gaps = 1/71 (1%)
Frame = +3
Query: 178 KVIDDLIPSIKRLRMLSLSNYNITVLPNSINKLVQLRYLNLSHTD-IKCLPDTTCDLYYL 236
K + S ++L +L L I LP+ I L +LRYLNLS + ++ LP+ +
Sbjct: 51 KALPSSFGSQRKLEILDLEQGRIESLPSCIVNLTRLRYLNLSDCEKLQTLPELPPS---V 221
Query: 237 QTLLLSGCWKL 247
+TLL GC L
Sbjct: 222 ETLLTRGCSSL 254
Score = 30.4 bits (67), Expect = 1.3
Identities = 29/96 (30%), Positives = 42/96 (43%), Gaps = 4/96 (4%)
Frame = +3
Query: 592 LPTSLQSLRIDECQNLAFLR----PETWSNYTSLVTLELKNCCDSLTSFQLNGFPVLQIL 647
LP+S S R E +L R P N T L L L +C T +L P ++ L
Sbjct: 57 LPSSFGSQRKLEILDLEQGRIESLPSCIVNLTRLRYLNLSDCEKLQTLPELP--PSVETL 230
Query: 648 SIEGCSSLKSIFISEKNSSLSLSTLQSLKVSNCKSL 683
GCSSLK++ + S + ++ C+SL
Sbjct: 231 LTRGCSSLKTVLFPSTATEQSQENQKRVEFFQCESL 338
>AV419023
Length = 410
Score = 36.6 bits (83), Expect = 0.018
Identities = 25/60 (41%), Positives = 32/60 (52%)
Frame = +3
Query: 193 LSLSNYNITVLPNSINKLVQLRYLNLSHTDIKCLPDTTCDLYYLQTLLLSGCWKLIELPI 252
L LS +I +PN+I +L L LNL + LP T L L L LS C KL +LP+
Sbjct: 12 LDLSFCSILEVPNAIGRLWCLERLNLQGNNFVELPSTISRLRRLAYLNLSHCIKLGDLPL 191
Score = 35.4 bits (80), Expect = 0.041
Identities = 26/76 (34%), Positives = 37/76 (48%), Gaps = 7/76 (9%)
Frame = +3
Query: 170 LQESYLSCKVIDDLIPSIKRLRMLSLSNYNITVLPNSINKLVQLRYLNLSHT----DIKC 225
L S+ S + + I + L L+L N LP++I++L +L YLNLSH D+
Sbjct: 12 LDLSFCSILEVPNAIGRLWCLERLNLQGNNFVELPSTISRLRRLAYLNLSHCIKLGDLPL 191
Query: 226 LPDTTCDL---YYLQT 238
LP L YY T
Sbjct: 192 LPSECAPLGGRYYETT 239
Database: LJGI
Posted date: Jul 30, 2004 11:16 AM
Number of letters in database: 14,692,800
Number of sequences in database: 28,460
Lambda K H
0.323 0.139 0.423
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 18,674,093
Number of Sequences: 28460
Number of extensions: 308325
Number of successful extensions: 1727
Number of sequences better than 10.0: 66
Number of HSP's better than 10.0 without gapping: 1666
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 1717
length of query: 898
length of database: 4,897,600
effective HSP length: 98
effective length of query: 800
effective length of database: 2,108,520
effective search space: 1686816000
effective search space used: 1686816000
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 16 ( 7.4 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.9 bits)
S2: 59 (27.3 bits)
Medicago: description of AC119417.2


