

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
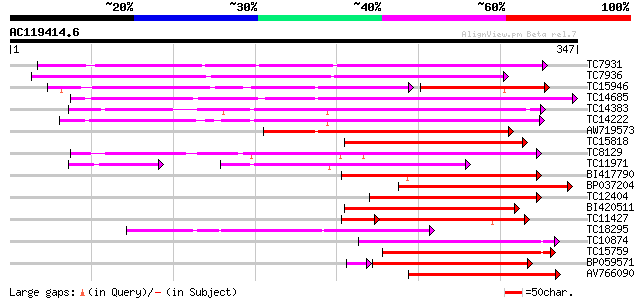
Query= AC119414.6 - phase: 0
(347 letters)
Database: LJGI
28,460 sequences; 14,692,800 total letters
Searching..................................................done
Score E
Sequences producing significant alignments: (bits) Value
TC7931 weakly similar to UP|Q43640 (Q43640) Dioxygenase, partial... 193 4e-50
TC7936 similar to UP|Q9SP53 (Q9SP53) Flavanone 3-hydroxylase, pa... 182 6e-47
TC15946 weakly similar to UP|Q43383 (Q43383) 2A6 protein, partia... 128 8e-45
TC14685 weakly similar to UP|Q43383 (Q43383) 2A6 protein, partia... 169 5e-43
TC14383 homologue to UP|Q8W3Y3 (Q8W3Y3) 1-aminocyclopropane-1-ca... 156 6e-39
TC14222 similar to UP|Q41681 (Q41681) 1-aminocyclopropane-1-carb... 152 6e-38
AW719573 128 2e-30
TC15818 similar to UP|Q9FFF6 (Q9FFF6) Leucoanthocyanidin dioxyge... 127 3e-30
TC8129 similar to UP|Q9XG83 (Q9XG83) GA 2-oxidase, partial (90%) 127 4e-30
TC11971 weakly similar to UP|Q43419 (Q43419) ACC oxidase, partia... 106 6e-29
BI417790 122 1e-28
BP037204 110 3e-25
TC12404 similar to UP|FLS_CITUN (Q9ZWQ9) Flavonol synthase (FLS... 107 2e-24
BI420511 107 2e-24
TC11427 weakly similar to UP|Q9LTH7 (Q9LTH7) Leucoanthocyanidin ... 89 5e-24
TC18295 similar to UP|Q39224 (Q39224) SRG1 mRNA (F6I1.30/F6I1.30... 104 2e-23
TC10874 similar to UP|Q8LF06 (Q8LF06) Leucoanthocyanidin dioxyge... 95 2e-20
TC15759 similar to UP|Q8LF06 (Q8LF06) Leucoanthocyanidin dioxyge... 91 3e-19
BP059571 86 4e-18
AV766090 82 1e-16
>TC7931 weakly similar to UP|Q43640 (Q43640) Dioxygenase, partial (29%)
Length = 1477
Score = 193 bits (490), Expect = 4e-50
Identities = 110/313 (35%), Positives = 172/313 (54%), Gaps = 1/313 (0%)
Frame = +1
Query: 18 NFILPEHKRPHLSEVKYLDSIPIIDLSYCDGNNPSSLEVIHKISKACEEFGFFQIVNHGV 77
+++ P RP V ++P++DL D E + +I KA EE+GFFQ++NHGV
Sbjct: 73 SYVQPPESRPGTVTVASGKAVPVVDLGGHD-----RAETLKQIFKASEEYGFFQVINHGV 237
Query: 78 PDQVCTKMMKAITNFFELAPEEREHLSSTDNTKNVRLFNYYLQVDGGEKVKLWSECFAHP 137
+++ + F + EE+ SS D + L+ +++ + ++ W + HP
Sbjct: 238 SNELIEDTLNIFKEFHAMPAEEKLSESSRDPNGSCMLYTSR-EINNKDCIQFWKDTLRHP 414
Query: 138 WYPIDDIIQLLPEKIGTQYREAFTEYAKEVGSLVRRLLSLISIGLGLEEDCLLKKLGEQP 197
P ++ ++ P K +YRE +Y KE+ L ++L ++ GLGL+ L E P
Sbjct: 415 CTPSEEFMEFWPLK-PARYREVVEKYTKELRELGSQILEMLCEGLGLDPRYCCAGLSENP 591
Query: 198 RQRAQSNFYPPCPDPELTMGLNEHTDLNALTVLLQ-SEVSGLQVNKDGKWISIPCIPNAF 256
SN YPPCP+P LT+GL++H D +T+LLQ ++V+ LQV KDG+WISI IP AF
Sbjct: 592 L--LLSNHYPPCPEPSLTLGLSKHRDPTLVTILLQDTDVNALQVFKDGEWISIEPIPYAF 765
Query: 257 VINLADQIEVLSNGRYKSVLHRAVTNNVQPRISMAMFYGPNPETIIGPIHELIDEEHPPK 316
V+N+ ++++SNGR V HR VTN+ R ++A F P E I+ P L+ P
Sbjct: 766 VVNIGLLLQIISNGRLVGVEHRVVTNSSISRTTVAYFIRPKKEEIVEPAKALVRTGAHPI 945
Query: 317 YRNYHFSKFLEEF 329
YR+ F +FL F
Sbjct: 946 YRSIAFEEFLRIF 984
>TC7936 similar to UP|Q9SP53 (Q9SP53) Flavanone 3-hydroxylase, partial
(94%)
Length = 1492
Score = 182 bits (463), Expect = 6e-47
Identities = 103/294 (35%), Positives = 164/294 (55%), Gaps = 2/294 (0%)
Frame = +3
Query: 14 SLTPNFILPEHKRPHLSEVKYLDSIPIIDLSYCDGNNPSSLEVIHKISKACEEFGFFQIV 73
+L +F+ E +RP ++ + + IP+I L+ D + E+ +KI +ACE +G FQ+V
Sbjct: 189 TLESSFVRDEDERPKVAYNNFSNEIPVISLAGIDEVDDRRSEICNKIVEACENWGIFQVV 368
Query: 74 NHGVPDQVCTKMMKAITNFFELAPEEREHLSSTDNTKNVRLFNYYLQVDGGEKVKLWSEC 133
+HGV ++ + M FF L PEE+ T K + + +LQ GE V+ W E
Sbjct: 369 DHGVDTELVSHMTTLAKEFFALPPEEKLRFDMTGGKKGGFIVSSHLQ---GESVQDWREI 539
Query: 134 FAHPWYPIDDIIQLLPEKIGTQYREAFTEYAKEVGSLVRRLLSLISIGLGLEEDCLLKKL 193
+ YPI + ++ EY++++ L +LL ++S +GLE++ L K
Sbjct: 540 VTYFSYPIRNRDYSRWPDTPAGWKAVTEEYSEKLMGLACKLLEVLSEAMGLEKEALTKAC 719
Query: 194 GEQPRQRAQSNFYPPCPDPELTMGLNEHTDLNALTVLLQSEVSGLQVNKD-GK-WISIPC 251
+ Q+ N+YP CP P+LT+GL HTD +T+LLQ +V GLQ +D GK WI++
Sbjct: 720 VDMD-QKVVVNYYPKCPQPDLTLGLKRHTDPGTITLLLQDQVGGLQATRDNGKTWITVQP 896
Query: 252 IPNAFVINLADQIEVLSNGRYKSVLHRAVTNNVQPRISMAMFYGPNPETIIGPI 305
+ AFV+NL D LSNGR+K+ H+AV N+ R+S+A F P P+ + P+
Sbjct: 897 VEGAFVVNLGDHGHYLSNGRFKNADHQAVVNSNSSRLSIATFQNPAPDATVYPL 1058
>TC15946 weakly similar to UP|Q43383 (Q43383) 2A6 protein, partial (56%)
Length = 1252
Score = 128 bits (321), Expect(2) = 8e-45
Identities = 80/227 (35%), Positives = 115/227 (50%), Gaps = 3/227 (1%)
Frame = +1
Query: 24 HKRPHLS---EVKYLDSIPIIDLSYCDGNNPSSLEVIHKISKACEEFGFFQIVNHGVPDQ 80
H HL E L S+P+IDL+ EVI KI AC E+GFFQ++NHG+P
Sbjct: 178 HANKHLDITPENSKLSSVPLIDLT------DEHSEVIGKIRSACHEWGFFQVINHGIPTS 339
Query: 81 VCTKMMKAITNFFELAPEEREHLSSTDNTKNVRLFNYYLQVDGGEKVKLWSECFAHPWYP 140
V +M+ I F E E R+ S D K V + G + W + F P
Sbjct: 340 VLDEMIDGIRRFHEQDSEVRKQFHSRDLKKKVMYYTNISLFSG--QAANWRDTFGFAVAP 513
Query: 141 IDDIIQLLPEKIGTQYREAFTEYAKEVGSLVRRLLSLISIGLGLEEDCLLKKLGEQPRQR 200
P++I R+ EY+K++ L +L L+S GLGL LK+L
Sbjct: 514 DPP----KPDEIPLVCRDIVMEYSKKIKELGFTILELLSEGLGLNPS-YLKELNCAEGLF 678
Query: 201 AQSNFYPPCPDPELTMGLNEHTDLNALTVLLQSEVSGLQVNKDGKWI 247
++YPPCP+P+LTMG +HTD N +T+LLQ ++ GLQ+ + +W+
Sbjct: 679 VLGHYYPPCPEPKLTMGTTKHTDGNFITLLLQDQLGGLQILHENQWV 819
Score = 68.9 bits (167), Expect(2) = 8e-45
Identities = 32/85 (37%), Positives = 53/85 (61%), Gaps = 6/85 (7%)
Frame = +2
Query: 252 IPNAFVINLADQIEVLSNGRYKSVLHRAVTNNVQPRISMAMFYGPNPETI------IGPI 305
+ A V+N+ D +++++N R+ SV HR ++ N+ PRIS+A F+ +P+ I GPI
Sbjct: 833 VHGALVVNIGDLLQLITNDRFVSVYHRVLSQNIGPRISVASFFVNSPDPIEGTLKVYGPI 1012
Query: 306 HELIDEEHPPKYRNYHFSKFLEEFF 330
EL+ EE+PP YR+ FL ++
Sbjct: 1013KELLTEENPPIYRDVTIKDFLAHYY 1087
>TC14685 weakly similar to UP|Q43383 (Q43383) 2A6 protein, partial (38%)
Length = 1323
Score = 169 bits (429), Expect = 5e-43
Identities = 110/313 (35%), Positives = 161/313 (51%), Gaps = 3/313 (0%)
Frame = +1
Query: 38 IPIIDLSYCDGNNPSSLEVIHKISKACEEFGFFQIVNHGVPDQVCTKMMKAITNFFELAP 97
IP IDL+ N S V+ +I +A GFFQ+VNHGVP + + + AI F E
Sbjct: 298 IPTIDLAAV---NESRAAVVDQIRRAASSVGFFQVVNHGVPLDLLRRTVAAIKAFHEQPE 468
Query: 98 EEREHLSSTDNTKNVRLFNYYLQVDGGE-KVKLWSECFAHPWYPIDDIIQLLPEKIGTQY 156
EER + + + V +Y VD + K W + P ++PE
Sbjct: 469 EERARVYTREMGTGV---SYISNVDLFQSKAASWRDTLQLRMGPTAVDQSMIPEVC---- 627
Query: 157 REAFTEYAKEVGSLVRRLLSLISIGLGLEEDCLLKKLGEQPRQRAQSNFYPPCPDPELTM 216
R+ E+ KEV + L L+S GLGL+ L ++G + ++YP CP P+LT+
Sbjct: 628 RQEVMEWDKEVVRVGEVLYGLLSEGLGLDAG-RLSEMGLVQGRVMVGHYYPFCPQPDLTV 804
Query: 217 GLNEHTDLNALTVLLQSEVSGLQVNKDGKWISIPCIPNAFVINLADQIEVLSNGRYKSVL 276
GLN H D ALTV+LQ + GLQV W+ + + A VIN+ D ++++SN YKS
Sbjct: 805 GLNSHADPGALTVVLQDHIGGLQVRTTEGWVDVKPLDGALVINIGDLLQIISNEEYKSAD 984
Query: 277 HRAVTNNV-QPRISMAMFYGP-NPETIIGPIHELIDEEHPPKYRNYHFSKFLEEFFNQEG 334
HR + N+ +PR+S+A+F P N E + GP+ EL E P YRN+ +FL FF +E
Sbjct: 985 HRVLANSANEPRVSVAVFLNPGNREKLFGPLPELTSAEKPALYRNFTLDEFLTRFFKKEL 1164
Query: 335 TRRIVKEVFELPC 347
+ + F C
Sbjct: 1165DGKSLTNYFRQ*C 1203
>TC14383 homologue to UP|Q8W3Y3 (Q8W3Y3) 1-aminocyclopropane-1-carboxylic
acid oxidase, complete
Length = 1249
Score = 156 bits (394), Expect = 6e-39
Identities = 96/301 (31%), Positives = 151/301 (49%), Gaps = 9/301 (2%)
Frame = +3
Query: 37 SIPIIDLSYCDGNNPSSLEVIHKISKACEEFGFFQIVNHGVPDQVCTKMMKAITNFFELA 96
+ P+I L +G +++ +I ACE +GFF++VNHG+P V + + + +
Sbjct: 45 NFPVISLEKLNGEERKDIKL--QIKDACENWGFFELVNHGIPHDVMDTVERLTKEHYRIC 218
Query: 97 PEEREHLSSTDNTKNVRLFNYYLQVDGGEKVKL------WSECFAHPWYPIDDIIQLLPE 150
E+R F + G E V+ W F P +I ++
Sbjct: 219 MEQR--------------FKELMASKGLEAVQTEVKDMDWESTFHLRHLPESNISEI--P 350
Query: 151 KIGTQYREAFTEYAKEVGSLVRRLLSLISIGLGLEEDCLLKKL--GEQPRQRAQSNFYPP 208
+ +YR+ ++A + L LL L+ LGLE+ L K P + YPP
Sbjct: 351 DLTDEYRKVMKDFALRIEKLSEDLLDLLCENLGLEKGYLKKAFHGSRGPTFGTKVANYPP 530
Query: 209 CPDPELTMGLNEHTDLNALTVLLQSE-VSGLQVNKDGKWISIPCIPNAFVINLADQIEVL 267
CP P L GL HTD + +L Q + VSGLQ+ KDG+W+ +P + ++ V+NL DQ+EV+
Sbjct: 531 CPKPNLVKGLRAHTDAGGIILLFQDDKVSGLQLLKDGEWVDVPPMRHSIVVNLGDQLEVI 710
Query: 268 SNGRYKSVLHRAVTNNVQPRISMAMFYGPNPETIIGPIHELIDEEHPPKYRNYHFSKFLE 327
+NG+YKSV HR + R+S+A FY P + +I P EL+++E K N + KF+
Sbjct: 711 TNGKYKSVEHRVIAQTDGTRMSIASFYNPGSDAVIYPAPELLEKEAEEK--NQLYPKFVF 884
Query: 328 E 328
E
Sbjct: 885 E 887
>TC14222 similar to UP|Q41681 (Q41681) 1-aminocyclopropane-1-carboxylate
oxidase homolog (ACC oxidase) , partial (97%)
Length = 1597
Score = 152 bits (385), Expect = 6e-38
Identities = 91/302 (30%), Positives = 157/302 (51%), Gaps = 5/302 (1%)
Frame = +1
Query: 31 EVKYLDSIPIIDLSYCDGNNPSSLEVIHKISKACEEFGFFQIVNHGVPDQVCTKMMKAIT 90
E K + + P++D+S N + + I ACE +GFF++VNH + + + +
Sbjct: 34 EEKKMANFPVVDMSKL--NTEERVATMELIKDACENWGFFELVNHEISTEFMDTVERLTK 207
Query: 91 NFFELAPEER-EHLSSTDNTKNVRLFNYYLQVDGGEKVKLWSECFAHPWYPIDDIIQLLP 149
++ E+R + + +T + V+ ++D + W F P +I ++
Sbjct: 208 EHYKRFMEQRFKEMVATKGLETVQS-----EIDDLD----WESTFFLRHLPSSNISEV-- 354
Query: 150 EKIGTQYREAFTEYAKEVGSLVRRLLSLISIGLGLEEDCLLKKL--GEQPRQRAQSNFYP 207
+ YR+ E+A ++ L LL L+ LGLE+ L K + P + + YP
Sbjct: 355 PDLDEDYRKVMKEFAVKLEKLAENLLDLLCENLGLEKGYLKKVFYGSKGPNFGTKVSNYP 534
Query: 208 PCPDPELTMGLNEHTDLNALTVLLQSE-VSGLQVNKDGKWISIPCIPNAFVINLADQIEV 266
PCP P+L GL HTD + +L Q + VSGLQ+ KD +WI +P + ++ V+NL DQ+EV
Sbjct: 535 PCPKPDLIKGLRAHTDAGGIILLFQDDKVSGLQLLKDDQWIDVPPMRHSIVVNLGDQLEV 714
Query: 267 LSNGRYKSVLHRAVTNNVQPRISMAMFYGPNPETIIGPIHELIDEEHPPK-YRNYHFSKF 325
++NG+YKSV+HR + R+S+A FY P + +I P L+ E+ + Y + F +
Sbjct: 715 ITNGKYKSVMHRVIAQTDGARMSLASFYNPGDDAVISPAPSLVKEDETSQVYPKFVFEDY 894
Query: 326 LE 327
++
Sbjct: 895 MK 900
>AW719573
Length = 521
Score = 128 bits (321), Expect = 2e-30
Identities = 60/154 (38%), Positives = 96/154 (61%), Gaps = 1/154 (0%)
Frame = +2
Query: 156 YREAFTEYAKEVGSLVRRLLSLISIGLGLEEDCLLKKLGEQPRQRAQSNFYPPCPDPELT 215
+RE Y E+ +L +++ L++ L ++ + ++ + Q + N+YPPCP PE
Sbjct: 62 FRENLEIYCAELNNLSIKIMELMANALKIDPK-EITEIYNEGSQTIRMNYYPPCPQPERV 238
Query: 216 MGLNEHTDLNALTVLLQS-EVSGLQVNKDGKWISIPCIPNAFVINLADQIEVLSNGRYKS 274
+GL HTD LT+LLQ+ ++ GLQ+ KDG W+ + +PNAF+IN+ D +E+ +NG Y S
Sbjct: 239 IGLKSHTDGAGLTILLQANDIEGLQIRKDGHWVPVQPLPNAFIINIGDMLEITTNGIYAS 418
Query: 275 VLHRAVTNNVQPRISMAMFYGPNPETIIGPIHEL 308
+ HRA N+ + RIS+A FYGPN + + P L
Sbjct: 419 IEHRATVNSEKERISLATFYGPNMQATLAPAPSL 520
>TC15818 similar to UP|Q9FFF6 (Q9FFF6) Leucoanthocyanidin dioxygenase-like
protein (AT5g05600/MOP10_14), partial (37%)
Length = 618
Score = 127 bits (319), Expect = 3e-30
Identities = 60/113 (53%), Positives = 79/113 (69%), Gaps = 1/113 (0%)
Frame = +2
Query: 206 YPPCPDPELTMGLNEHTDLNALTVLLQSE-VSGLQVNKDGKWISIPCIPNAFVINLADQI 264
YP CP P+LT+GL+ H+D LT+LL + VSGLQV K WI++ +PNAF+IN+ DQI
Sbjct: 2 YPKCPQPDLTLGLSPHSDPGGLTILLPDDFVSGLQVRKGNYWITVKPVPNAFIINIGDQI 181
Query: 265 EVLSNGRYKSVLHRAVTNNVQPRISMAMFYGPNPETIIGPIHELIDEEHPPKY 317
+VLSN YKSV HR + N+ Q R+S+AMFY P + +I P EL+ EE P Y
Sbjct: 182 QVLSNTIYKSVEHRVIVNSNQDRVSLAMFYNPKSDLLIEPAKELVTEEKPALY 340
>TC8129 similar to UP|Q9XG83 (Q9XG83) GA 2-oxidase, partial (90%)
Length = 1278
Score = 127 bits (318), Expect = 4e-30
Identities = 87/302 (28%), Positives = 145/302 (47%), Gaps = 14/302 (4%)
Frame = +2
Query: 38 IPIIDLSYCDGNNPSSLEVIHKISKACEEFGFFQIVNHGVPDQVCTKMMKAITNFFELAP 97
+P +DLS+ + I AC+EFGFF+I+NH +P ++ + + +FF
Sbjct: 197 VPEVDLSHPEAKT--------LIVNACKEFGFFKIINHDIPMELISNLENEALSFFMQPQ 352
Query: 98 EEREHLSSTDNTKNVRLFNYYLQVDGGEKVKLWSECFAHPWYPIDDIIQ----LLPEKIG 153
E+E S D F Y + G W E P D+I LL E+
Sbjct: 353 SEKEKASLDDP------FGYGSKRIGKNGDVGWVEYLLFNTNP--DVISPNPLLLLEQNQ 508
Query: 154 TQYREAFTEYAKEVGSLVRRLLSLISIGLGLEEDCLLKKLGEQPRQRA--QSNFYPPCPD 211
+ A +Y V +L +L L++ GLG+E+ + +L + + N YP C +
Sbjct: 509 NVFSSAVNDYTLAVKNLSCEILELMADGLGIEQRNVFSRLIRDEKSDCIFRLNHYPACGE 688
Query: 212 PELT-------MGLNEHTDLNALTVLLQSEVSGLQVN-KDGKWISIPCIPNAFVINLADQ 263
EL +G EHTD ++VL + SGLQ+ +DG W+SIP +F IN+ D
Sbjct: 689 LELQAIGGRNLIGFGEHTDPQMISVLRSNNTSGLQICLRDGTWVSIPPDQTSFFINVGDS 868
Query: 264 IEVLSNGRYKSVLHRAVTNNVQPRISMAMFYGPNPETIIGPIHELIDEEHPPKYRNYHFS 323
++V++NG ++SV HR +++ R+SM F G + P+ L+ +E Y+ + +
Sbjct: 869 LQVMTNGSFRSVKHRVLSDTAMSRVSMIYFGGAPLTEKLAPLPSLVSKEEDSLYKEFTWG 1048
Query: 324 KF 325
++
Sbjct: 1049EY 1054
>TC11971 weakly similar to UP|Q43419 (Q43419) ACC oxidase, partial (69%)
Length = 730
Score = 106 bits (265), Expect(2) = 6e-29
Identities = 63/157 (40%), Positives = 87/157 (55%), Gaps = 4/157 (2%)
Frame = +3
Query: 130 WSECFAHPWYPIDDIIQLLPEKIGTQYREAFTEYAKEVGSLVRRLLSLISIGLGLEEDCL 189
W F P +I Q+ + + R+ +Y +++ L L L+S LGLE+D +
Sbjct: 252 WESAFFIWHRPKSNIKQI--SNLSEELRQTMDQYIEKLIQLAEDLSGLMSENLGLEKDYI 425
Query: 190 LKKLG--EQPRQRAQSNFYPPCPDPELTMGLNEHTDLNALTVLLQSE-VSGLQVNKDGKW 246
K + P + YP CP+PEL GL EHTD + +LLQ + V GL+ +KDGKW
Sbjct: 426 KKAFSGSDGPAMGTKVAKYPQCPNPELVRGLREHTDAGGIILLLQDDQVPGLEFSKDGKW 605
Query: 247 ISIPCIPN-AFVINLADQIEVLSNGRYKSVLHRAVTN 282
+ IP N A IN DQIEVLSNG YKSV+HR + +
Sbjct: 606 VEIPPSKNNAIFINTGDQIEVLSNGLYKSVVHRVMAD 716
Score = 37.4 bits (85), Expect(2) = 6e-29
Identities = 19/58 (32%), Positives = 32/58 (54%)
Frame = +2
Query: 37 SIPIIDLSYCDGNNPSSLEVIHKISKACEEFGFFQIVNHGVPDQVCTKMMKAITNFFE 94
+IPIID S +G+ E + + +AC+ +G F I NHG+ + K+ + I +E
Sbjct: 5 AIPIIDFSTLNGDKRG--ETMALLHEACKNWGCFLIENHGIDKSLMEKVKQLINTHYE 172
>BI417790
Length = 610
Score = 122 bits (305), Expect = 1e-28
Identities = 56/124 (45%), Positives = 82/124 (65%), Gaps = 2/124 (1%)
Frame = +3
Query: 204 NFYPPCPDPELTMGLNEHTDLNALTVLLQSEVSGLQVNK--DGKWISIPCIPNAFVINLA 261
N+YPPCP+P++ +G H D ALT+L Q EVSGLQV + DG+W+ + +P+A++IN+
Sbjct: 15 NYYPPCPNPDIVLGCGRHKDSGALTILAQDEVSGLQVRRKCDGQWVRLNPLPDAYIINVG 194
Query: 262 DQIEVLSNGRYKSVLHRAVTNNVQPRISMAMFYGPNPETIIGPIHELIDEEHPPKYRNYH 321
D I+V N Y+SV HR N + R+S F P+ T + P+ EL +EE+PPKY Y+
Sbjct: 195 DVIQVWCNDAYESVEHRVKLNPEKARLSYPFFLYPSHYTNVEPLEELTNEENPPKYTPYN 374
Query: 322 FSKF 325
+ KF
Sbjct: 375 WGKF 386
>BP037204
Length = 527
Score = 110 bits (276), Expect = 3e-25
Identities = 46/106 (43%), Positives = 76/106 (71%)
Frame = -1
Query: 239 QVNKDGKWISIPCIPNAFVINLADQIEVLSNGRYKSVLHRAVTNNVQPRISMAMFYGPNP 298
QV KD KW+++ +PN F++N+ DQI+V+SN +YKS LHRA+ NN + R+S+ FY P+P
Sbjct: 527 QVLKDEKWVTVNPVPNTFIVNIGDQIQVISNDKYKSALHRALVNNEKERMSIPTFYCPSP 348
Query: 299 ETIIGPIHELIDEEHPPKYRNYHFSKFLEEFFNQEGTRRIVKEVFE 344
+ +IGP +LID +HP +Y NY +S++ F+N+ ++ ++F+
Sbjct: 347 DALIGPAPQLIDIDHPAQYTNYAYSEYYHNFWNRGLSKTTCVDMFK 210
>TC12404 similar to UP|FLS_CITUN (Q9ZWQ9) Flavonol synthase (FLS) (CitFLS)
, partial (31%)
Length = 526
Score = 107 bits (268), Expect = 2e-24
Identities = 48/105 (45%), Positives = 73/105 (68%)
Frame = +2
Query: 221 HTDLNALTVLLQSEVSGLQVNKDGKWISIPCIPNAFVINLADQIEVLSNGRYKSVLHRAV 280
HTDL++LT+LL +EV GLQV KD +WI IP+A ++++ DQI++ SNGRYKSVLHR
Sbjct: 5 HTDLSSLTILLPNEVPGLQVLKDERWIDAKYIPHALIVHIGDQIQIASNGRYKSVLHRTT 184
Query: 281 TNNVQPRISMAMFYGPNPETIIGPIHELIDEEHPPKYRNYHFSKF 325
+ + R+S +F P E +GP+ +L+ +++PPKY+ F +
Sbjct: 185 VDKERTRMSWPVFLEPPAECEVGPLPQLVTQDNPPKYKVKKFKDY 319
>BI420511
Length = 384
Score = 107 bits (268), Expect = 2e-24
Identities = 50/108 (46%), Positives = 72/108 (66%), Gaps = 1/108 (0%)
Frame = +3
Query: 206 YPPCPDPELTMGLNEHTDLNALTVLLQSE-VSGLQVNKDGKWISIPCIPNAFVINLADQI 264
YP CP PEL GL HTD + +LLQ + VSGLQ+ KDG+W+ +P + ++ VINL DQ+
Sbjct: 36 YPACPKPELVKGLRAHTDAGGIILLLQDDKVSGLQLLKDGQWVDVPPMRHSIVINLGDQL 215
Query: 265 EVLSNGRYKSVLHRAVTNNVQPRISMAMFYGPNPETIIGPIHELIDEE 312
EV++NG+YKSV HR + R+S+A FY P + +I P L++ +
Sbjct: 216 EVITNGKYKSVEHRVIAKTDGTRMSIASFYNPANDAVIYPAPALLESK 359
>TC11427 weakly similar to UP|Q9LTH7 (Q9LTH7) Leucoanthocyanidin
dioxygenase-like protein (AT5g59540/f2o15_200), partial
(30%)
Length = 690
Score = 89.4 bits (220), Expect(2) = 5e-24
Identities = 38/99 (38%), Positives = 65/99 (65%), Gaps = 4/99 (4%)
Frame = +1
Query: 224 LNALTVLLQSEVSGLQVNKDGKWISIPCIPNAFVINLADQIEVLSNGRYKSVLHRAVTNN 283
L +T+LLQ ++ GLQ+ +W+ +P + + V+N+ D +++++NG++ SV HR ++ +
Sbjct: 91 LTLMTILLQDQIGGLQILHQNQWVDVPPVHGSLVVNIGDLLQLMTNGQFSSVYHRVLSRH 270
Query: 284 VQPRISMAMFY----GPNPETIIGPIHELIDEEHPPKYR 318
V PRIS+A F+ +IGPI EL+ EE+PP YR
Sbjct: 271 VGPRISIASFFVNISSQGKSKVIGPIKELLSEENPPIYR 387
Score = 38.1 bits (87), Expect(2) = 5e-24
Identities = 12/23 (52%), Positives = 21/23 (91%)
Frame = +3
Query: 204 NFYPPCPDPELTMGLNEHTDLNA 226
++YP CP+PELTMG ++H+D+++
Sbjct: 30 SYYPTCPEPELTMGTSKHSDIDS 98
>TC18295 similar to UP|Q39224 (Q39224) SRG1 mRNA (F6I1.30/F6I1.30)
(At1g17020/F6I1.30), partial (29%)
Length = 561
Score = 104 bits (259), Expect = 2e-23
Identities = 63/190 (33%), Positives = 100/190 (52%), Gaps = 1/190 (0%)
Frame = +3
Query: 72 IVNHGVPDQVCTKMMKAITNFFELAPEEREHLSSTDNTKNVRLFNYYLQVDGGEKVKLWS 131
++NHGV + M K + + F L EE++ L T F V +K++ W+
Sbjct: 3 LINHGVNPSLVENMKKGVQDLFNLPIEEKKKLWQTPEVGQG--FGQAFVVSEEQKLE-WA 173
Query: 132 ECFAHPWYPIDDIIQLLPEKIGTQYREAFTEYAKEVGSLVRRLLSLISIGLGLEEDCLLK 191
+ F +P++ L I +R+ Y E+ ++ ++ L+ L E + L++
Sbjct: 174 DLFYINTFPLNKRNTSLIPSIPQPFRDNLDAYCSELKNICVTIIGLMEKALKTETNELVE 353
Query: 192 KLGEQPRQRAQSNFYPPCPDPELTMGLNEHTDLNALTVLLQ-SEVSGLQVNKDGKWISIP 250
L E Q + N+YPPCP PE +GLN H+D A T+LLQ +E+ GLQ+ KDG W+ I
Sbjct: 354 -LFEDTSQGMRMNYYPPCPQPEKVIGLNPHSDSGAFTILLQVNEMEGLQIRKDGIWVPIK 530
Query: 251 CIPNAFVINL 260
+ NAFVIN+
Sbjct: 531 PLSNAFVINI 560
>TC10874 similar to UP|Q8LF06 (Q8LF06) Leucoanthocyanidin dioxygenase-like
protein, partial (38%)
Length = 723
Score = 95.1 bits (235), Expect = 2e-20
Identities = 49/124 (39%), Positives = 73/124 (58%), Gaps = 1/124 (0%)
Frame = +2
Query: 214 LTMGLNEHTDLNALTVLL-QSEVSGLQVNKDGKWISIPCIPNAFVINLADQIEVLSNGRY 272
LT+GL+ H+D +T+LL +V GLQV K WI++ +AF++N+ DQI+VLSN Y
Sbjct: 2 LTLGLSSHSDPGGMTMLLPDDQVRGLQVRKGDDWITVNPARHAFIVNIGDQIQVLSNAIY 181
Query: 273 KSVLHRAVTNNVQPRISMAMFYGPNPETIIGPIHELIDEEHPPKYRNYHFSKFLEEFFNQ 332
SV HR + N+ + R+S+A FY P + I P+ EL+ + P Y F ++ F
Sbjct: 182 TSVEHRVIVNSDKERVSLAFFYNPKSDIPIEPVKELVTPDKPALYPAMTFDEY-RLFIRM 358
Query: 333 EGTR 336
G R
Sbjct: 359 RGPR 370
>TC15759 similar to UP|Q8LF06 (Q8LF06) Leucoanthocyanidin dioxygenase-like
protein, partial (34%)
Length = 929
Score = 90.9 bits (224), Expect = 3e-19
Identities = 43/106 (40%), Positives = 65/106 (60%)
Frame = +2
Query: 229 VLLQSEVSGLQVNKDGKWISIPCIPNAFVINLADQIEVLSNGRYKSVLHRAVTNNVQPRI 288
+L +V+GLQV K WI++ +P+AF++N+ DQI+VLSN YKSV HR + N+ + R+
Sbjct: 2 LLPDDQVTGLQVRKGDNWITVKPVPHAFIVNIGDQIQVLSNAIYKSVEHRVIVNSNKERV 181
Query: 289 SMAMFYGPNPETIIGPIHELIDEEHPPKYRNYHFSKFLEEFFNQEG 334
S+A FY P + I P EL+ E+ P Y F ++ F +G
Sbjct: 182 SLAFFYNPRGDIPIEPAKELVKEDKPALYTAMTFDEY-RRFIRMKG 316
>BP059571
Length = 386
Score = 85.5 bits (210), Expect(2) = 4e-18
Identities = 45/98 (45%), Positives = 61/98 (61%)
Frame = +2
Query: 223 DLNALTVLLQSEVSGLQVNKDGKWISIPCIPNAFVINLADQIEVLSNGRYKSVLHRAVTN 282
D +LT+L Q +V GLQV D +W SI NAFV+N+ D LSNGRYKS LHRAV N
Sbjct: 77 DPTSLTILHQDQVGGLQVFVDNEWHSISPDLNAFVVNIGDTFMALSNGRYKSCLHRAVVN 256
Query: 283 NVQPRISMAMFYGPNPETIIGPIHELIDEEHPPKYRNY 320
+ + R S+A F P + ++ P EL+D +P Y ++
Sbjct: 257 SEKTRKSLAFFLCPRNDKVVTPPCELVDSCNPRIYPDF 370
Score = 21.9 bits (45), Expect(2) = 4e-18
Identities = 7/15 (46%), Positives = 9/15 (59%)
Frame = +3
Query: 207 PPCPDPELTMGLNEH 221
P C P+LT+G H
Sbjct: 30 PSCQKPDLTLGTGPH 74
>AV766090
Length = 532
Score = 82.0 bits (201), Expect = 1e-16
Identities = 38/93 (40%), Positives = 59/93 (62%)
Frame = -1
Query: 245 KWISIPCIPNAFVINLADQIEVLSNGRYKSVLHRAVTNNVQPRISMAMFYGPNPETIIGP 304
KW+++ IPNAFV+N+ D +E+ SNG+YKSVLHR V N V+ R+S+A + +
Sbjct: 526 KWVTVQPIPNAFVVNIGDHLEIYSNGKYKSVLHRVVVNEVKSRVSVASLHSLPFTCTVRV 347
Query: 305 IHELIDEEHPPKYRNYHFSKFLEEFFNQEGTRR 337
+LIDEE+P +Y + F FL ++E ++
Sbjct: 346 SPKLIDEENPKRYMDTDFETFLAYVSSREPKKK 248
Database: LJGI
Posted date: Jul 30, 2004 11:16 AM
Number of letters in database: 14,692,800
Number of sequences in database: 28,460
Lambda K H
0.319 0.138 0.417
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 6,573,509
Number of Sequences: 28460
Number of extensions: 96844
Number of successful extensions: 611
Number of sequences better than 10.0: 128
Number of HSP's better than 10.0 without gapping: 570
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 573
length of query: 347
length of database: 4,897,600
effective HSP length: 91
effective length of query: 256
effective length of database: 2,307,740
effective search space: 590781440
effective search space used: 590781440
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 16 ( 7.4 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.7 bits)
S2: 56 (26.2 bits)
Medicago: description of AC119414.6


