

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
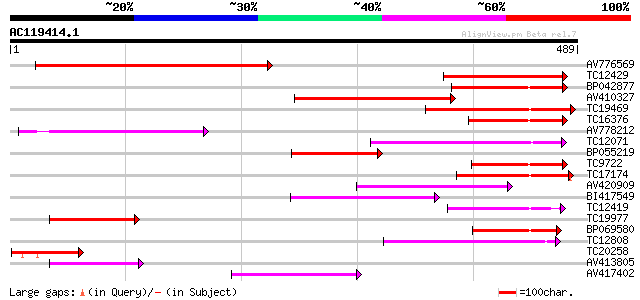
Query= AC119414.1 + phase: 0
(489 letters)
Database: LJGI
28,460 sequences; 14,692,800 total letters
Searching..................................................done
Score E
Sequences producing significant alignments: (bits) Value
AV776569 203 4e-53
TC12429 similar to UP|Q9FKQ1 (Q9FKQ1) Emb|CAB89401.1 (AT5g65380/... 173 5e-44
BP042877 127 4e-30
AV410327 124 3e-29
TC19469 weakly similar to UP|Q9LP32 (Q9LP32) T9L6.1 protein, par... 119 9e-28
TC16376 similar to UP|Q9LVD9 (Q9LVD9) Gb|AAC28507.1, partial (17%) 117 4e-27
AV778212 117 6e-27
TC12071 weakly similar to GB|AAP31968.1|30387605|BT006624 At2g04... 112 1e-25
BP055219 108 2e-24
TC9722 similar to UP|Q9LVD9 (Q9LVD9) Gb|AAC28507.1, partial (16%) 107 4e-24
TC17174 weakly similar to UP|Q940N9 (Q940N9) AT4g21910/T8O5_120,... 105 2e-23
AV420909 97 5e-21
BI417549 87 6e-18
TC12419 84 5e-17
TC19977 similar to UP|Q9LVD9 (Q9LVD9) Gb|AAC28507.1, partial (20%) 84 7e-17
BP069580 76 1e-14
TC12808 weakly similar to GB|CAA16564.1|2827556|ATT12H17 predict... 70 1e-12
TC20258 similar to UP|Q9FR32 (Q9FR32) Ripening regulated protein... 64 4e-11
AV413805 60 1e-09
AV417402 43 1e-04
>AV776569
Length = 633
Score = 203 bits (517), Expect = 4e-53
Identities = 98/205 (47%), Positives = 135/205 (65%), Gaps = 1/205 (0%)
Frame = +2
Query: 23 DEHENEV-LGRRVWNESKKLWHIAGPAIFNRVSNYSMLVITQVFAGHLGDMELAATSIAM 81
DE E + L RR ESKKLW IAGPAI + YS+ T F GH+G+++LAA S+
Sbjct: 17 DEDEKPLSLVRRFGIESKKLWKIAGPAILTMLCQYSLGAFTLTFVGHVGELDLAAVSVEN 196
Query: 82 NLILGLDLGIMLGMSSALETLCGQAFGAKQFNMLGIYMQRSWIVLFITGILLLPLFIFAT 141
+ I G G+MLGM SALETLCGQAFGA Q MLG+YMQRSW++LF T ++L+P ++++
Sbjct: 197 SCIAGFSFGVMLGMGSALETLCGQAFGAGQSRMLGVYMQRSWVILFTTALILVPAYVWSP 376
Query: 142 PILNFFGQPQEISELAGVISMWLIPTHVTYAFFFPLYFFLQSQLKNNIIGWVSLVALLVH 201
PIL GQ EISE AG ++W++P YAF FP+ FLQSQ K ++ W+S L++H
Sbjct: 377 PILRVIGQTTEISEAAGKFALWMLPQLFAYAFNFPMQKFLQSQRKVQVMLWISATVLVLH 556
Query: 202 VFLCWLVVVKFKLGVIALVASGNVA 226
+F WL+++K G+ + N +
Sbjct: 557 IFFSWLLILKLGWGLTGAAITLNAS 631
>TC12429 similar to UP|Q9FKQ1 (Q9FKQ1) Emb|CAB89401.1 (AT5g65380/MNA5_11),
partial (22%)
Length = 523
Score = 173 bits (439), Expect = 5e-44
Identities = 78/107 (72%), Positives = 95/107 (87%)
Frame = +1
Query: 375 VIEEVNKLSPLLGFTILLNSVQPVLSGVAIGSGWQKYVAYIDLGCYYLIGMPLGFLMGFV 434
V++EVN LS LL FTILLNS+QPVLSGVA+GSGWQ YVAYI+LGCYYL+G+PLGF+MG+V
Sbjct: 1 VLDEVNNLSLLLAFTILLNSIQPVLSGVAVGSGWQSYVAYINLGCYYLVGVPLGFIMGWV 180
Query: 435 FQFGVEGLWAGLVCGGPAIQTLILAWVTIRCDWNKEAERAKLHLSKW 481
F GV G+WAG++ GG A+QTLIL +TIRCDW+ EAE+AKLHL+KW
Sbjct: 181 FNQGVMGIWAGMIFGGTALQTLILGVITIRCDWDGEAEKAKLHLTKW 321
>BP042877
Length = 489
Score = 127 bits (319), Expect = 4e-30
Identities = 54/100 (54%), Positives = 76/100 (76%)
Frame = -1
Query: 382 LSPLLGFTILLNSVQPVLSGVAIGSGWQKYVAYIDLGCYYLIGMPLGFLMGFVFQFGVEG 441
L PLL +ILL+ +QPVLSGVA+G GW +VAY+++GCYY +G+PLG ++GF FQFG +G
Sbjct: 483 LCPLLALSILLHGIQPVLSGVAVGCGWPAFVAYVNVGCYYGVGIPLGSVLGFYFQFGAQG 304
Query: 442 LWAGLVCGGPAIQTLILAWVTIRCDWNKEAERAKLHLSKW 481
+W G++ GG + T+IL WVT R DW++E E A L++W
Sbjct: 303 IWLGML-GGTTMPTIILMWVTFRTDWDQEVEEAAQRLNQW 187
>AV410327
Length = 424
Score = 124 bits (312), Expect = 3e-29
Identities = 65/139 (46%), Positives = 91/139 (64%)
Frame = +2
Query: 246 TWTGFSMEAFFDLWEFAKLSAASGVMLCLEVWYDKVLMLMTGNLHNAKKFVEALTICLTL 305
TW GFS +AF +L EF KLSAAS VMLCLE WY ++L+L+ G L + + +++L+IC T+
Sbjct: 8 TWQGFSWQAFSELPEFFKLSAASAVMLCLETWYFQILVLLAGLLPHPELALDSLSICTTI 187
Query: 306 NIWELMFPLGFLAATGVRVANELGAGNGQAAKFASAVAVVTSIIISVFFWLLIMIFRRQI 365
+ W M +GF AA VRV+NELGA N ++A F+ V + S I+SV L+++ R I
Sbjct: 188 SGWVFMISVGFNAAASVRVSNELGARNPKSASFSVVVVTLISFIMSVIAALVVLALRDII 367
Query: 366 GYLFTSSELVIEEVNKLSP 384
Y+FT E V V+ L P
Sbjct: 368 SYVFTGGEEVAAAVSDLCP 424
>TC19469 weakly similar to UP|Q9LP32 (Q9LP32) T9L6.1 protein, partial (26%)
Length = 501
Score = 119 bits (299), Expect = 9e-28
Identities = 53/130 (40%), Positives = 85/130 (64%)
Frame = +1
Query: 359 MIFRRQIGYLFTSSELVIEEVNKLSPLLGFTILLNSVQPVLSGVAIGSGWQKYVAYIDLG 418
+ R ++ Y+FT V E V LSPLL +ILLNSV PVLSGV++G+GW VAY+++G
Sbjct: 1 LFLRERLAYVFTLDSEVAEAVGDLSPLLSISILLNSVPPVLSGVSVGAGWPSVVAYVNIG 180
Query: 419 CYYLIGMPLGFLMGFVFQFGVEGLWAGLVCGGPAIQTLILAWVTIRCDWNKEAERAKLHL 478
CYYLIG+P+G ++G + +G+W G++ G +QT++L +T++ DW + A+ +
Sbjct: 181 CYYLIGIPVGVVLGNLIHLQEKGVWIGMLF-GTFVQTVMLITITLKTDWENQVVIARNRV 357
Query: 479 SKWGAPNHAQ 488
+KW + +
Sbjct: 358 NKWAVKENTE 387
>TC16376 similar to UP|Q9LVD9 (Q9LVD9) Gb|AAC28507.1, partial (17%)
Length = 550
Score = 117 bits (293), Expect = 4e-27
Identities = 50/86 (58%), Positives = 65/86 (75%)
Frame = +2
Query: 396 QPVLSGVAIGSGWQKYVAYIDLGCYYLIGMPLGFLMGFVFQFGVEGLWAGLVCGGPAIQT 455
QPVLSGVA+G GWQ +VAY+++GCYY IG+P G ++GF F+FG G+W G++ G +QT
Sbjct: 2 QPVLSGVAVGCGWQTFVAYVNVGCYYGIGIPFGAVLGFYFKFGAMGIWLGML-AGTVLQT 178
Query: 456 LILAWVTIRCDWNKEAERAKLHLSKW 481
+IL WVT R DWNKE E A L+KW
Sbjct: 179 IILVWVTFRTDWNKEIEEATKRLNKW 256
>AV778212
Length = 557
Score = 117 bits (292), Expect = 6e-27
Identities = 65/164 (39%), Positives = 96/164 (57%)
Frame = +1
Query: 8 EALVEAKVPLLKNNIDEHENEVLGRRVWNESKKLWHIAGPAIFNRVSNYSMLVITQVFAG 67
+ L + ++P LK ++ W E L+ +AGPAI + N M T+ FAG
Sbjct: 91 QVLSDTELPWLKRHLSA---------TWIELNLLFPLAGPAILVYLINNFMSTATRAFAG 243
Query: 68 HLGDMELAATSIAMNLILGLDLGIMLGMSSALETLCGQAFGAKQFNMLGIYMQRSWIVLF 127
HLG++ LAA ++ + I G+MLGM SA+ETLCGQA+GA +++MLGIYMQR++IVL
Sbjct: 244 HLGNLALAAANLGNDGIQLFAYGLMLGMGSAVETLCGQAYGAHKYDMLGIYMQRAFIVLT 423
Query: 128 ITGILLLPLFIFATPILNFFGQPQEISELAGVISMWLIPTHVTY 171
ITGI L +++F IL G+ +++ + LIP Y
Sbjct: 424 ITGIPLTVVYVFCKQILLVLGESDDVASAGAIFVYGLIPQIYAY 555
>TC12071 weakly similar to GB|AAP31968.1|30387605|BT006624 At2g04100
{Arabidopsis thaliana;}, partial (32%)
Length = 764
Score = 112 bits (281), Expect = 1e-25
Identities = 61/169 (36%), Positives = 96/169 (56%)
Frame = +3
Query: 312 FPLGFLAATGVRVANELGAGNGQAAKFASAVAVVTSIIISVFFWLLIMIFRRQIGYLFTS 371
+ G AA RV+NELGAG + A+ A ++ + + ++ + R +G+ F++
Sbjct: 18 YSYGVGAAVSTRVSNELGAGKPKEARDAIFAVIILATLDAIILSSALFCCRHVLGFAFSN 197
Query: 372 SELVIEEVNKLSPLLGFTILLNSVQPVLSGVAIGSGWQKYVAYIDLGCYYLIGMPLGFLM 431
V+ V K+ PLL ++ ++S VLSGVA G+GWQK A +L YY IG+P+ L+
Sbjct: 198 DLEVVHSVAKIVPLLCLSVCVDSFLGVLSGVARGAGWQKIGAVTNLLAYYAIGIPVALLL 377
Query: 432 GFVFQFGVEGLWAGLVCGGPAIQTLILAWVTIRCDWNKEAERAKLHLSK 480
GFV + +GLW G++ G QTL+LA +T W K+A A + S+
Sbjct: 378 GFVLKLNAKGLWIGILTGS-TTQTLVLALLTAFTKWEKQAPLATVRTSE 521
>BP055219
Length = 527
Score = 108 bits (270), Expect = 2e-24
Identities = 51/78 (65%), Positives = 63/78 (80%)
Frame = -1
Query: 244 PLTWTGFSMEAFFDLWEFAKLSAASGVMLCLEVWYDKVLMLMTGNLHNAKKFVEALTICL 303
P TW+G SMEAF LWEF KLSAASGVMLC+E WY +VL++MTGNL NA+ V+AL+IC+
Sbjct: 527 PHTWSGLSMEAFSGLWEFVKLSAASGVMLCVENWYCRVLIVMTGNLQNAEIAVDALSICM 348
Query: 304 TLNIWELMFPLGFLAATG 321
++N E+M PL F AATG
Sbjct: 347 SINGLEMMIPLSFFAATG 294
>TC9722 similar to UP|Q9LVD9 (Q9LVD9) Gb|AAC28507.1, partial (16%)
Length = 571
Score = 107 bits (267), Expect = 4e-24
Identities = 46/83 (55%), Positives = 63/83 (75%)
Frame = +3
Query: 399 LSGVAIGSGWQKYVAYIDLGCYYLIGMPLGFLMGFVFQFGVEGLWAGLVCGGPAIQTLIL 458
LSGVA+G G Q +VAY+++GCYY +G+PLG ++GF F+FG +G+W G++ GG +QT+IL
Sbjct: 3 LSGVAVGCGCQTFVAYVNVGCYYGVGIPLGAVLGFYFKFGAKGIWLGML-GGTVMQTIIL 179
Query: 459 AWVTIRCDWNKEAERAKLHLSKW 481
WVT R DW KE E A L+KW
Sbjct: 180 MWVTFRTDWIKEVEEASKRLNKW 248
>TC17174 weakly similar to UP|Q940N9 (Q940N9) AT4g21910/T8O5_120, partial
(18%)
Length = 723
Score = 105 bits (262), Expect = 2e-23
Identities = 47/103 (45%), Positives = 74/103 (71%), Gaps = 2/103 (1%)
Frame = +1
Query: 386 LGFTILLNSVQPVLSGVAIGSGWQKYVAYIDLGCYYLIGMPLGFLMGFVFQFGVEGLWAG 445
L +ILL SV PVLSGVA+G+GW+ VAY++LGCYY+IG+P+G ++G VF + V+G+W G
Sbjct: 1 LAVSILLYSVPPVLSGVALGAGWRSTVAYVNLGCYYIIGLPVGIVLGNVFHWQVKGIWIG 180
Query: 446 LVCGGPAIQTLILAWVTIRCDWNKEAERAKLHLSKWG--APNH 486
++ G +QT++L +T R DW ++ A+ +++W P+H
Sbjct: 181 MLF-GTLVQTIVLVIITYRTDWEEQVTIARNRVNRWSKVEPDH 306
>AV420909
Length = 421
Score = 97.4 bits (241), Expect = 5e-21
Identities = 49/134 (36%), Positives = 80/134 (59%)
Frame = +3
Query: 300 TICLTLNIWELMFPLGFLAATGVRVANELGAGNGQAAKFASAVAVVTSIIISVFFWLLIM 359
++CL AA RV+NELGAGN + AK A V V+ I+ ++ +
Sbjct: 3 SVCLNTTTLHYFVTYAVGAAASTRVSNELGAGNPKTAKGAVRVVVIVGIVEAIIVSAFFI 182
Query: 360 IFRRQIGYLFTSSELVIEEVNKLSPLLGFTILLNSVQPVLSGVAIGSGWQKYVAYIDLGC 419
FR +GY +++ + V++ V ++PLL +++ +S+ LSGVA G G+Q+ AY +LG
Sbjct: 183 CFRNVLGYAYSNDKEVVDYVADMAPLLCGSVIGDSLIGALSGVARGGGFQQIGAYANLGA 362
Query: 420 YYLIGMPLGFLMGF 433
YY++G+P+G L+GF
Sbjct: 363 YYVVGIPIGLLLGF 404
>BI417549
Length = 404
Score = 87.0 bits (214), Expect = 6e-18
Identities = 47/128 (36%), Positives = 70/128 (53%)
Frame = +1
Query: 243 CPLTWTGFSMEAFFDLWEFAKLSAASGVMLCLEVWYDKVLMLMTGNLHNAKKFVEALTIC 302
C T T FS A L EF +LS S M+C++ W ++L+L+ G N + L+IC
Sbjct: 19 CEKTRTPFSKRALSGLGEFFRLSVPSAAMVCVKWWASELLVLLAGLFSNPELETSILSIC 198
Query: 303 LTLNIWELMFPLGFLAATGVRVANELGAGNGQAAKFASAVAVVTSIIISVFFWLLIMIFR 362
LT++ P GF A RVANELGAGN +AA+ ++A+ + I +V + FR
Sbjct: 199 LTISTLHFTIPYGFGVAASTRVANELGAGNPKAARVVVSIAMSLAFIEAVIVSTALFGFR 378
Query: 363 RQIGYLFT 370
+GY ++
Sbjct: 379 HILGYAYS 402
>TC12419
Length = 667
Score = 84.0 bits (206), Expect = 5e-17
Identities = 46/102 (45%), Positives = 60/102 (58%)
Frame = +3
Query: 378 EVNKLSPLLGFTILLNSVQPVLSGVAIGSGWQKYVAYIDLGCYYLIGMPLGFLMGFVFQF 437
E ++PLL +ILL+S+Q VLSGVA G GWQ AY++L +YLIG+P+ L+ F
Sbjct: 3 EFASVAPLLAISILLDSIQGVLSGVARGCGWQHLAAYVNLATFYLIGLPISCLLAFKTSL 182
Query: 438 GVEGLWAGLVCGGPAIQTLILAWVTIRCDWNKEAERAKLHLS 479
+GLW GL+C G Q L +T R W AKL LS
Sbjct: 183 QYKGLWIGLIC-GMVCQAGTLLLLTTRAKW------AKLELS 287
>TC19977 similar to UP|Q9LVD9 (Q9LVD9) Gb|AAC28507.1, partial (20%)
Length = 452
Score = 83.6 bits (205), Expect = 7e-17
Identities = 41/78 (52%), Positives = 56/78 (71%)
Frame = +3
Query: 35 WNESKKLWHIAGPAIFNRVSNYSMLVITQVFAGHLGDMELAATSIAMNLILGLDLGIMLG 94
W E K L+++A PA+ + NY M + TQ+F+GHLG++ELAA S+ I G+MLG
Sbjct: 216 WIELKLLFYLAVPAVIVYLINYVMSMSTQIFSGHLGNLELAAASLGNTGIQVFAYGLMLG 395
Query: 95 MSSALETLCGQAFGAKQF 112
M SA+ETLCGQA+GAK+F
Sbjct: 396 MGSAVETLCGQAYGAKKF 449
>BP069580
Length = 483
Score = 76.3 bits (186), Expect = 1e-14
Identities = 35/77 (45%), Positives = 49/77 (63%)
Frame = -1
Query: 400 SGVAIGSGWQKYVAYIDLGCYYLIGMPLGFLMGFVFQFGVEGLWAGLVCGGPAIQTLILA 459
SG A G G QK A+I+LG YYL+G+PL ++ FV G +GLW G++C +Q L
Sbjct: 264 SGTARGCGLQKIGAFINLGSYYLVGIPLAMVLAFVLHIGGKGLWLGIIC-ALIVQVFSLM 88
Query: 460 WVTIRCDWNKEAERAKL 476
+T+R DW KEA R ++
Sbjct: 87 IITLRIDWEKEARRPRI 37
>TC12808 weakly similar to GB|CAA16564.1|2827556|ATT12H17 predicted protein
{Arabidopsis thaliana;}, partial (32%)
Length = 624
Score = 69.7 bits (169), Expect = 1e-12
Identities = 43/153 (28%), Positives = 76/153 (49%)
Frame = +1
Query: 323 RVANELGAGNGQAAKFASAVAVVTSIIISVFFWLLIMIFRRQIGYLFTSSELVIEEVNKL 382
RV+NELGA A ++ V++ +I+ L+++ R G LF+ +I V K
Sbjct: 10 RVSNELGANQAGRAYRSACVSLALGLILGCIGSLVMVAARGIWGPLFSHDRGIINGVKKT 189
Query: 383 SPLLGFTILLNSVQPVLSGVAIGSGWQKYVAYIDLGCYYLIGMPLGFLMGFVFQFGVEGL 442
L+ + N V G+ G+ Y +LG +Y + +PLG + F G+ GL
Sbjct: 190 MLLMALVEVFNFPLAVCGGIVRGTARPWLGMYANLGGFYFLALPLGVVFAFKLNLGLFGL 369
Query: 443 WAGLVCGGPAIQTLILAWVTIRCDWNKEAERAK 475
+ GL+ G +L++ ++ R +W +EA +A+
Sbjct: 370 FIGLITGIVTCLSLLVIFIA-RINWEEEASKAQ 465
>TC20258 similar to UP|Q9FR32 (Q9FR32) Ripening regulated protein DDTFR18,
partial (8%)
Length = 303
Score = 64.3 bits (155), Expect = 4e-11
Identities = 38/67 (56%), Positives = 48/67 (70%), Gaps = 5/67 (7%)
Frame = +2
Query: 2 SKVEASEA--LVEAKVPLLKNNI--DEHENEV-LGRRVWNESKKLWHIAGPAIFNRVSNY 56
SK+ + E L E K PLL+ + D E E L R++W ESKKLWHIAGPAIF+R+++Y
Sbjct: 98 SKMPSKEVSPLGEIKAPLLEVHPLPDAAEQEQDLKRKIWVESKKLWHIAGPAIFSRIASY 277
Query: 57 SMLVITQ 63
SMLVITQ
Sbjct: 278 SMLVITQ 298
>AV413805
Length = 418
Score = 59.7 bits (143), Expect = 1e-09
Identities = 30/82 (36%), Positives = 48/82 (57%), Gaps = 1/82 (1%)
Frame = +3
Query: 35 WNESKKLWHIAGPAIFNRVSNYSMLVITQVFAGHLGDM-ELAATSIAMNLILGLDLGIML 93
+ E KK+ +A P + VS Y + V++ + GH ++ + +IA++ ++L
Sbjct: 171 YQELKKVSFMAAPMVAVTVSQYLLQVVSLMMVGHFSELVSFSGVAIAISFAEVTGFSVLL 350
Query: 94 GMSSALETLCGQAFGAKQFNML 115
GMS ALETLCGQ FGA++F L
Sbjct: 351 GMSGALETLCGQTFGAEEFGKL 416
>AV417402
Length = 371
Score = 43.1 bits (100), Expect = 1e-04
Identities = 27/113 (23%), Positives = 54/113 (46%), Gaps = 1/113 (0%)
Frame = +3
Query: 192 WVSLVALLVHVFLCWLVVVKFKLGVIALVASGNVAWIVLVFGFFGYAVLCGC-PLTWTGF 250
+ +++++L+H+ + +L+V LG+ + LV Y + G TWTG
Sbjct: 9 YCAVLSILLHIPINYLLVSVLHLGIRGIALGAVWTNFNLVISLIIYIWVSGIHKKTWTGI 188
Query: 251 SMEAFFDLWEFAKLSAASGVMLCLEVWYDKVLMLMTGNLHNAKKFVEALTICL 303
S F L+ S + +CLE W+ ++++L+ G L N V ++ + +
Sbjct: 189 SSACFKGWKALLNLAIPSCISVCLEWWWYEIMILLCGLLINPHATVASMGVLI 347
Database: LJGI
Posted date: Jul 30, 2004 11:16 AM
Number of letters in database: 14,692,800
Number of sequences in database: 28,460
Lambda K H
0.328 0.142 0.451
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 9,831,713
Number of Sequences: 28460
Number of extensions: 156644
Number of successful extensions: 1199
Number of sequences better than 10.0: 61
Number of HSP's better than 10.0 without gapping: 1183
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 1191
length of query: 489
length of database: 4,897,600
effective HSP length: 94
effective length of query: 395
effective length of database: 2,222,360
effective search space: 877832200
effective search space used: 877832200
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 15 ( 7.1 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 40 (21.8 bits)
S2: 57 (26.6 bits)
Medicago: description of AC119414.1


