

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC119412.5 + phase: 0
(221 letters)
Database: LJGI
28,460 sequences; 14,692,800 total letters
Searching..................................................done
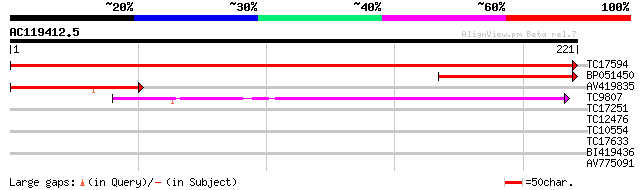
Score E
Sequences producing significant alignments: (bits) Value
TC17594 similar to UP|VT13_ARATH (Q9LVP9) Vesicle transport v-SN... 382 e-107
BP051450 82 6e-17
AV419835 75 1e-14
TC9807 weakly similar to UP|ME11_ARATH (Q9SJL6) Membrin 11 (AtME... 44 2e-05
TC17251 similar to UP|Q9M3V0 (Q9M3V0) Protein phosphatase 2C (PP... 29 0.73
TC12476 similar to GB|AAC72109.1|3850569|F15K9 ESTs gb|T21276, g... 27 2.8
TC10554 similar to PIR|T48847|T48847 syntaxin synt4 [imported] -... 27 3.6
TC17633 similar to UP|BAD08108 (BAD08108) Uridine kinase-like pr... 26 6.2
BI419436 25 8.1
AV775091 25 8.1
>TC17594 similar to UP|VT13_ARATH (Q9LVP9) Vesicle transport v-SNARE 13
(AtVTI13) (Vesicle transport v-SNARE protein VTI13)
(Vesicle soluble NSF attachment protein receptor 13),
complete
Length = 1097
Score = 382 bits (981), Expect = e-107
Identities = 194/221 (87%), Positives = 211/221 (94%)
Frame = +3
Query: 1 MSNVFEGYERQYCELSANLSKKCTAATALNGEQKKQKVSEVKSGIDEAEALIRKMDLEAR 60
MSNVFEGYERQYCELSANLSKKCTAA +LNGEQKKQKVSE+K+GIDEAEALIRKMDLEAR
Sbjct: 120 MSNVFEGYERQYCELSANLSKKCTAAGSLNGEQKKQKVSEIKAGIDEAEALIRKMDLEAR 299
Query: 61 SMQPNIKGVLLAKLREYKSDLNNLKSEVKKIVSGNLNPSARDELLESGMADTMTASADQR 120
S+QPN++ VLLAKLREYKSDLNNLKSEVKKIVSGNLNPSARDELLESGMADT+TAS+DQR
Sbjct: 300 SLQPNVRAVLLAKLREYKSDLNNLKSEVKKIVSGNLNPSARDELLESGMADTLTASSDQR 479
Query: 121 TRLMTSTERLNKTGERVKDSRRTMLETEELGVSILQDLHSQRQSLLHAHNTLHGVDDNIG 180
+RLM ST+RLNKTG+RV+ SR+TMLETEELGVSILQDLHSQRQSLLHAH TLHGVDDNIG
Sbjct: 480 SRLMMSTDRLNKTGDRVQQSRKTMLETEELGVSILQDLHSQRQSLLHAHTTLHGVDDNIG 659
Query: 181 KSKKIMTNMSRRMNKNKWIIGCIVLVLVVAIIAILYFKLVK 221
KS+KI+TNM+RRMNKNKWII IVLVLV+AI ILYFKL K
Sbjct: 660 KSQKILTNMTRRMNKNKWIICGIVLVLVIAIALILYFKLSK 782
>BP051450
Length = 460
Score = 82.4 bits (202), Expect = 6e-17
Identities = 40/54 (74%), Positives = 45/54 (83%)
Frame = -2
Query: 168 AHNTLHGVDDNIGKSKKIMTNMSRRMNKNKWIIGCIVLVLVVAIIAILYFKLVK 221
A +TLHGVD+NIGKSKKI++NMSRRMNKNKWII I +LV II ILYFKL K
Sbjct: 459 AGDTLHGVDNNIGKSKKILSNMSRRMNKNKWIISIITAILVFVIILILYFKLSK 298
>AV419835
Length = 186
Score = 74.7 bits (182), Expect = 1e-14
Identities = 34/53 (64%), Positives = 48/53 (90%), Gaps = 1/53 (1%)
Frame = +2
Query: 1 MSNVFEGYERQYCELSANLSKKCTAATALNG-EQKKQKVSEVKSGIDEAEALI 52
MS VFEGYERQYC+LSANLS+KC++A+ ++ EQK+QK+SE+K+G+D+A+ LI
Sbjct: 26 MSEVFEGYERQYCDLSANLSRKCSSASLVSDLEQKQQKLSEIKAGLDDADVLI 184
>TC9807 weakly similar to UP|ME11_ARATH (Q9SJL6) Membrin 11 (AtMEMB11)
(Golgi SNAP receptor complex member 2-1) (27 kDa Golgi
SNARE protein), partial (68%)
Length = 1062
Score = 44.3 bits (103), Expect = 2e-05
Identities = 39/180 (21%), Positives = 82/180 (44%), Gaps = 2/180 (1%)
Frame = +3
Query: 41 VKSGIDEAEALIRKMDLEARSM--QPNIKGVLLAKLREYKSDLNNLKSEVKKIVSGNLNP 98
VK I + ++L +MD S+ +P + + K+ + + +LK K NL
Sbjct: 225 VKKDIAQVQSLCVEMDRLWGSVGGKPQ-RDLWRRKVEQIAEEAESLKESWDKF---NLRN 392
Query: 99 SARDELLESGMADTMTASADQRTRLMTSTERLNKTGERVKDSRRTMLETEELGVSILQDL 158
R + E+ + A+ + +M + + + V+ S R + +G +IL +
Sbjct: 393 QKR--ISEAKERAELLGRANGDSHVMRIFDEEAQAMQSVRTSARELENANAIGEAILSSI 566
Query: 159 HSQRQSLLHAHNTLHGVDDNIGKSKKIMTNMSRRMNKNKWIIGCIVLVLVVAIIAILYFK 218
H QR+ L AH + + +G S ++ + RR ++WI +L+ V+ + A ++++
Sbjct: 567 HGQRERLKSAHRKALDILNTVGISNSVLRLIERRNRVDQWIKYAGMLLTVIFLFAFVFWR 746
>TC17251 similar to UP|Q9M3V0 (Q9M3V0) Protein phosphatase 2C (PP2C) ,
partial (30%)
Length = 715
Score = 28.9 bits (63), Expect = 0.73
Identities = 17/60 (28%), Positives = 31/60 (51%)
Frame = -1
Query: 161 QRQSLLHAHNTLHGVDDNIGKSKKIMTNMSRRMNKNKWIIGCIVLVLVVAIIAILYFKLV 220
+R S LHA+NT HG+ N ++ +++ W G ++V I++L+F+ V
Sbjct: 706 KRTS*LHANNTDHGLH*NSNHTQPRQGYQNKKGKW*TW*KGAKATFIMVIRISLLFFQFV 527
>TC12476 similar to GB|AAC72109.1|3850569|F15K9 ESTs gb|T21276, gb|T45403,
and gb|AA586113 come from this gene. {Arabidopsis
thaliana;}, partial (7%)
Length = 791
Score = 26.9 bits (58), Expect = 2.8
Identities = 13/40 (32%), Positives = 25/40 (62%)
Frame = +2
Query: 29 LNGEQKKQKVSEVKSGIDEAEALIRKMDLEARSMQPNIKG 68
L+ + + ++VSEVK+ D +E+ + D+ S QP++ G
Sbjct: 197 LDIKNEVKEVSEVKADHDNSESSCKDSDISIVSTQPSMPG 316
>TC10554 similar to PIR|T48847|T48847 syntaxin synt4 [imported] -
Arabidopsis thaliana {Arabidopsis thaliana;}, partial
(11%)
Length = 545
Score = 26.6 bits (57), Expect = 3.6
Identities = 9/22 (40%), Positives = 15/22 (67%)
Frame = +1
Query: 194 NKNKWIIGCIVLVLVVAIIAIL 215
N KW CI+L+LV+ ++ +L
Sbjct: 10 NTRKWTCFCILLLLVIILVVVL 75
>TC17633 similar to UP|BAD08108 (BAD08108) Uridine kinase-like protein,
partial (3%)
Length = 559
Score = 25.8 bits (55), Expect = 6.2
Identities = 19/66 (28%), Positives = 32/66 (47%), Gaps = 1/66 (1%)
Frame = +1
Query: 154 ILQDLHSQRQSLLHAHNTL-HGVDDNIGKSKKIMTNMSRRMNKNKWIIGCIVLVLVVAII 212
+L QRQ L+H +TL H + + G+ + RRM + + I +VL L + +
Sbjct: 43 VLLVARGQRQ-LIHQLDTLSHLLHEYFGERSRQDRTEQRRMGEIESIAIPVVLTLAIGAV 219
Query: 213 AILYFK 218
+ FK
Sbjct: 220 GVFLFK 237
>BI419436
Length = 601
Score = 25.4 bits (54), Expect = 8.1
Identities = 9/31 (29%), Positives = 19/31 (61%)
Frame = +2
Query: 188 NMSRRMNKNKWIIGCIVLVLVVAIIAILYFK 218
+++RR+ + WI G + + L ++ LYF+
Sbjct: 125 SLARRITAHLWIFGVLAVCLQSYLLGSLYFR 217
>AV775091
Length = 398
Score = 25.4 bits (54), Expect = 8.1
Identities = 14/41 (34%), Positives = 25/41 (60%), Gaps = 1/41 (2%)
Frame = +3
Query: 126 STERLNKTG-ERVKDSRRTMLETEELGVSILQDLHSQRQSL 165
+ E+ N+ G +++KD + L E + +S LQ+ H + QSL
Sbjct: 27 TNEKHNEEGRQKIKDYTFSQLFREGIDISALQNSHRKIQSL 149
Database: LJGI
Posted date: Jul 30, 2004 11:16 AM
Number of letters in database: 14,692,800
Number of sequences in database: 28,460
Lambda K H
0.314 0.129 0.342
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 2,639,890
Number of Sequences: 28460
Number of extensions: 24848
Number of successful extensions: 142
Number of sequences better than 10.0: 20
Number of HSP's better than 10.0 without gapping: 141
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 141
length of query: 221
length of database: 4,897,600
effective HSP length: 87
effective length of query: 134
effective length of database: 2,421,580
effective search space: 324491720
effective search space used: 324491720
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 16 ( 7.2 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 42 (22.0 bits)
S2: 53 (25.0 bits)
Medicago: description of AC119412.5


