

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC119412.2 - phase: 0
(184 letters)
Database: LJGI
28,460 sequences; 14,692,800 total letters
Searching..................................................done
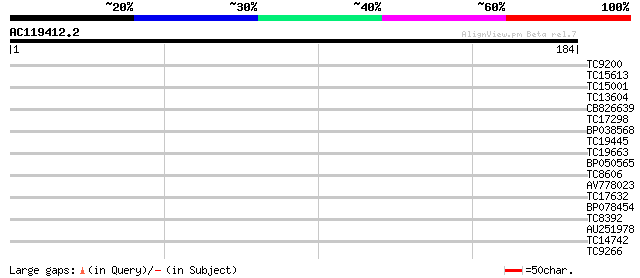
Score E
Sequences producing significant alignments: (bits) Value
TC9200 similar to UP|Q8LLE6 (Q8LLE6) RNA-binding protein AKIP1, ... 30 0.32
TC15613 similar to UP|Q8S9K2 (Q8S9K2) At1g75460/F1B16_22, partia... 30 0.32
TC15001 similar to UP|GLE1_YEAST (Q12315) RNA export factor GLE1... 30 0.32
TC13604 30 0.32
CB826639 28 0.72
TC17298 similar to UP|Q9AT29 (Q9AT29) bZIP transcription factor,... 28 0.94
BP038568 28 1.2
TC19445 UP|Q83ZH3 (Q83ZH3) IcaA (Fragment), partial (8%) 28 1.2
TC19663 similar to UP|Q8H6S8 (Q8H6S8) Translation initiation fac... 28 1.2
BP050565 28 1.2
TC8606 similar to UP|AAQ82688 (AAQ82688) Epa4p, partial (7%) 27 1.6
AV778023 27 2.1
TC17632 similar to GB|AAO44044.1|28466871|BT004778 At4g24310 {Ar... 27 2.7
BP078454 26 3.6
TC8392 similar to UP|Q8LK52 (Q8LK52) Cp10-like protein, partial ... 26 4.7
AU251978 25 6.1
TC14742 similar to UP|Q9XFC0 (Q9XFC0) Cytosolic ascorbate peroxi... 25 6.1
TC9266 similar to UP|Q9ASW5 (Q9ASW5) Epoxide hydrolase (Express... 25 6.1
>TC9200 similar to UP|Q8LLE6 (Q8LLE6) RNA-binding protein AKIP1, partial
(31%)
Length = 727
Score = 29.6 bits (65), Expect = 0.32
Identities = 28/90 (31%), Positives = 38/90 (42%)
Frame = +1
Query: 38 KQENNSNTFEEVGKIVEEGVDDTEEDIILEECSTMKMVEELEPLHPEEFPQERPYTEEAE 97
K E +EEV + VEE V++ EE EE +E EE E
Sbjct: 172 KLEAEVQEYEEVEEEVEEEVEEEEE---------------------EEEEEEEGEEEEEE 288
Query: 98 TVENEEVMEVTEKEDRILIKEESMEEKGKK 127
E EEV E+ED I ++ +E GK+
Sbjct: 289 EEEEEEVEAEAEEEDDEPI-QKLLEPMGKE 375
Score = 28.5 bits (62), Expect = 0.72
Identities = 21/76 (27%), Positives = 35/76 (45%)
Frame = +1
Query: 57 VDDTEEDIILEECSTMKMVEELEPLHPEEFPQERPYTEEAETVENEEVMEVTEKEDRILI 116
+ ++ + ++ E T K + E P + EAE E EEV E E+E
Sbjct: 64 LQESPNEAMVNESITKKRKLVPKSSQSAELPFKLQNKLEAEVQEYEEVEEEVEEEVEEEE 243
Query: 117 KEESMEEKGKKVNKVE 132
+EE EE+G++ + E
Sbjct: 244 EEEEEEEEGEEEEEEE 291
>TC15613 similar to UP|Q8S9K2 (Q8S9K2) At1g75460/F1B16_22, partial (30%)
Length = 491
Score = 29.6 bits (65), Expect = 0.32
Identities = 15/30 (50%), Positives = 18/30 (60%)
Frame = -1
Query: 56 GVDDTEEDIILEECSTMKMVEELEPLHPEE 85
GVDD E +I +EEC V ELE L E+
Sbjct: 431 GVDDAEAEIGVEECVHHDAVAELEDLERED 342
>TC15001 similar to UP|GLE1_YEAST (Q12315) RNA export factor GLE1, partial
(4%)
Length = 988
Score = 29.6 bits (65), Expect = 0.32
Identities = 25/87 (28%), Positives = 45/87 (50%), Gaps = 8/87 (9%)
Frame = +2
Query: 61 EEDIILEECSTMKMVEEL------EPLHPEEFPQE--RPYTEEAETVENEEVMEVTEKED 112
E ++ L E T K VEE E L+ EE E R E + + E +++ ++++
Sbjct: 368 EAELKLIEEETAKRVEEAIRKRVEESLNSEEVQVEIQRRLEEGRKRLIVEVAVQLEKEKE 547
Query: 113 RILIKEESMEEKGKKVNKVEIDRIIDE 139
LI+ EE+ +K K ++DR+++E
Sbjct: 548 AALIEAREKEEQARK-EKEDLDRMLEE 625
>TC13604
Length = 520
Score = 29.6 bits (65), Expect = 0.32
Identities = 17/66 (25%), Positives = 36/66 (53%), Gaps = 5/66 (7%)
Frame = +3
Query: 79 EPLHPEEFPQERPYTEEAETVENEEVMEVTEKEDRI-----LIKEESMEEKGKKVNKVEI 133
+P H + Q++ TEE + + NE++ TE++D I +K + EE+ + K++
Sbjct: 42 DPYHSYKNSQKKYDTEEEDHIRNEKMKFHTEEDDHIRNEKMKLKFYTEEEEDIRNEKMKF 221
Query: 134 DRIIDE 139
D + ++
Sbjct: 222 DTLFED 239
>CB826639
Length = 542
Score = 28.5 bits (62), Expect = 0.72
Identities = 19/62 (30%), Positives = 29/62 (46%)
Frame = +3
Query: 25 IMDIQANIQSTHGKQENNSNTFEEVGKIVEEGVDDTEEDIILEECSTMKMVEELEPLHPE 84
++++ A + T G EE GK VEE VD E D ++ + + L P PE
Sbjct: 291 VLELAAVLTDTRG---------EETGKWVEEAVDFEEADRLVYLKAALAETLRLYPSVPE 443
Query: 85 EF 86
+F
Sbjct: 444 DF 449
>TC17298 similar to UP|Q9AT29 (Q9AT29) bZIP transcription factor, partial
(39%)
Length = 484
Score = 28.1 bits (61), Expect = 0.94
Identities = 22/111 (19%), Positives = 46/111 (40%), Gaps = 10/111 (9%)
Frame = +1
Query: 15 SFQMEMMQEQIMDIQANIQSTHGKQENNSNTFEEVGKIVEEGVD-DTEEDIILEECSTMK 73
+FQ++ Q Q N+Q+ NS++ +E D ++ +I+ E +
Sbjct: 142 TFQLQKFSNQFFGYQNNLQAFQDFSPQNSSSCISSNSTSDEAEDQQQQQSLIINERKHRR 321
Query: 74 MVEELEPLHPEEFPQERPYTEEAETV-----ENEEVME----VTEKEDRIL 115
M+ E +++ E V EN +++E ++E DR++
Sbjct: 322 MISNRESARRSRMRKQKHLDELWSMVVCLRNENHQLLEKLNHLSESHDRVV 474
>BP038568
Length = 553
Score = 27.7 bits (60), Expect = 1.2
Identities = 15/56 (26%), Positives = 27/56 (47%)
Frame = +1
Query: 84 EEFPQERPYTEEAETVENEEVMEVTEKEDRILIKEESMEEKGKKVNKVEIDRIIDE 139
EE E+P TEE E + V E E++ E + K++ E +++++E
Sbjct: 25 EELGDEKPATEEKEAAYGNKESPVNESEEK--------EPEDKEMTLEEYEKVLEE 168
>TC19445 UP|Q83ZH3 (Q83ZH3) IcaA (Fragment), partial (8%)
Length = 553
Score = 27.7 bits (60), Expect = 1.2
Identities = 16/71 (22%), Positives = 35/71 (48%), Gaps = 1/71 (1%)
Frame = +1
Query: 12 KNLSFQMEMMQEQIMDIQANIQSTHGKQENNSNTFEEVGKIVEE-GVDDTEEDIILEECS 70
K L+ + + ++ + +++ NIQ T GK+E V +I ++ G + + + +
Sbjct: 19 KKLAGKFQALKGKFKEMKGNIQKTAGKEEPQDEPAGSVDEIKKKYGFSSSSNETSAAKLA 198
Query: 71 TMKMVEELEPL 81
K+ E L+ L
Sbjct: 199ESKLQENLQKL 231
>TC19663 similar to UP|Q8H6S8 (Q8H6S8) Translation initiation factor,
partial (10%)
Length = 485
Score = 27.7 bits (60), Expect = 1.2
Identities = 21/88 (23%), Positives = 39/88 (43%)
Frame = +3
Query: 70 STMKMVEELEPLHPEEFPQERPYTEEAETVENEEVMEVTEKEDRILIKEESMEEKGKKVN 129
S K ++ P H E + +EAE + E E KE+ ++E +E + ++
Sbjct: 162 SKTKAADKKLPKHVREMQEALARRKEAEERKKREEEEKLRKEEEERRRQEELERQAEEAR 341
Query: 130 KVEIDRIIDEICALFNKPKLGRIWTPHQ 157
+ + +R E L K + G++ T Q
Sbjct: 342 RRKKER---EKEKLLKKKQEGKLLTGKQ 416
>BP050565
Length = 542
Score = 27.7 bits (60), Expect = 1.2
Identities = 19/55 (34%), Positives = 28/55 (50%)
Frame = -1
Query: 31 NIQSTHGKQENNSNTFEEVGKIVEEGVDDTEEDIILEECSTMKMVEELEPLHPEE 85
N + T K +S +F +VGK VEE + T E ++ E + EE LH E+
Sbjct: 443 NTKGTPVKNVCSSGSFADVGKFVEEILQKTSELLLSE------IKEETIGLHGEQ 297
>TC8606 similar to UP|AAQ82688 (AAQ82688) Epa4p, partial (7%)
Length = 1008
Score = 27.3 bits (59), Expect = 1.6
Identities = 22/77 (28%), Positives = 36/77 (46%)
Frame = -2
Query: 37 GKQENNSNTFEEVGKIVEEGVDDTEEDIILEECSTMKMVEELEPLHPEEFPQERPYTEEA 96
G +EN + +E G+ E+ DD E+D +E EE EE +E +E
Sbjct: 449 GPEENGEDEEDEDGEDQEDDDDDDEDD---DEEDDGGEDEE------EEGVEEEDNEDEE 297
Query: 97 ETVENEEVMEVTEKEDR 113
E E+EE ++ +K +
Sbjct: 296 EDEEDEEALQPPKKRKK 246
>AV778023
Length = 583
Score = 26.9 bits (58), Expect = 2.1
Identities = 14/36 (38%), Positives = 24/36 (65%), Gaps = 1/36 (2%)
Frame = +2
Query: 44 NTFEEVG-KIVEEGVDDTEEDIILEECSTMKMVEEL 78
N+ E++ K VEE E+D+I++E T+ ++EEL
Sbjct: 293 NSIEDLDIKKVEEQAAKLEKDLIVKELETLDVLEEL 400
>TC17632 similar to GB|AAO44044.1|28466871|BT004778 At4g24310 {Arabidopsis
thaliana;}, partial (31%)
Length = 396
Score = 26.6 bits (57), Expect = 2.7
Identities = 11/22 (50%), Positives = 15/22 (68%)
Frame = +1
Query: 31 NIQSTHGKQENNSNTFEEVGKI 52
N +S K++NNSNT EE G +
Sbjct: 91 NNESDEAKKKNNSNTEEEAGAV 156
>BP078454
Length = 385
Score = 26.2 bits (56), Expect = 3.6
Identities = 14/48 (29%), Positives = 22/48 (45%)
Frame = -3
Query: 83 PEEFPQERPYTEEAETVENEEVMEVTEKEDRILIKEESMEEKGKKVNK 130
PEE +E V+ + + KE++ +KEE E + KV K
Sbjct: 308 PEEDGDSSSGSETGSPVDKKAARKEARKENKKKVKEEKKEARKNKVPK 165
>TC8392 similar to UP|Q8LK52 (Q8LK52) Cp10-like protein, partial (81%)
Length = 1014
Score = 25.8 bits (55), Expect = 4.7
Identities = 11/26 (42%), Positives = 17/26 (65%)
Frame = +1
Query: 99 VENEEVMEVTEKEDRILIKEESMEEK 124
+E +EV ++ DR+LIK + EEK
Sbjct: 565 LETDEVQDLKPLNDRVLIKVAAAEEK 642
>AU251978
Length = 381
Score = 25.4 bits (54), Expect = 6.1
Identities = 15/39 (38%), Positives = 20/39 (50%), Gaps = 1/39 (2%)
Frame = -3
Query: 87 PQERPYTEEAETVENEEVMEVTEKEDRILIKE-ESMEEK 124
P E PY + +TE +D ILIK+ S+EEK
Sbjct: 157 PPEIPYPHPLPMLSCTYTSTITENKDPILIKK*NSVEEK 41
>TC14742 similar to UP|Q9XFC0 (Q9XFC0) Cytosolic ascorbate peroxidase,
complete
Length = 1256
Score = 25.4 bits (54), Expect = 6.1
Identities = 15/58 (25%), Positives = 28/58 (47%)
Frame = +3
Query: 60 TEEDIILEECSTMKMVEELEPLHPEEFPQERPYTEEAETVENEEVMEVTEKEDRILIK 117
T++DI+ + L HPE + P+TE+ +N +E+ ++E L+K
Sbjct: 525 TDKDIV-----ALSGAHTLGRAHPERSGFDGPWTEDPLKFDNSYFVELLKEESAGLLK 683
>TC9266 similar to UP|Q9ASW5 (Q9ASW5) Epoxide hydrolase (Expressed
protein) (At2g18360/T30D6.13) , partial (34%)
Length = 506
Score = 25.4 bits (54), Expect = 6.1
Identities = 9/27 (33%), Positives = 19/27 (70%)
Frame = +3
Query: 151 RIWTPHQLYFKFMEFLPTRRIEKDDVL 177
++W P++L+ F+E + T R E+ ++L
Sbjct: 21 KLWFPNRLHKDFLEVMFTNRKERGELL 101
Database: LJGI
Posted date: Jul 30, 2004 11:16 AM
Number of letters in database: 14,692,800
Number of sequences in database: 28,460
Lambda K H
0.313 0.132 0.367
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 2,498,488
Number of Sequences: 28460
Number of extensions: 25885
Number of successful extensions: 155
Number of sequences better than 10.0: 36
Number of HSP's better than 10.0 without gapping: 152
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 155
length of query: 184
length of database: 4,897,600
effective HSP length: 85
effective length of query: 99
effective length of database: 2,478,500
effective search space: 245371500
effective search space used: 245371500
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 16 ( 7.2 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 42 (21.9 bits)
S2: 52 (24.6 bits)
Medicago: description of AC119412.2


