

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
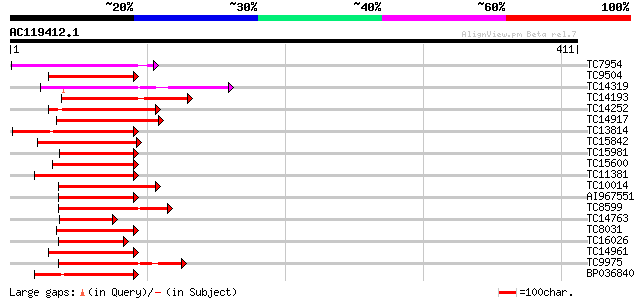
Query= AC119412.1 - phase: 0
(411 letters)
Database: LJGI
28,460 sequences; 14,692,800 total letters
Searching..................................................done
Score E
Sequences producing significant alignments: (bits) Value
TC7954 similar to UP|Q7XAU4 (Q7XAU4) Ethylene responsive protein... 81 3e-16
TC9504 similar to UP|Q9SXS8 (Q9SXS8) Ethylene responsive element... 81 4e-16
TC14319 similar to UP|AP23_ARATH (P42736) AP2 domain transcripti... 80 5e-16
TC14193 similar to UP|Q8S2S7 (Q8S2S7) Ethylene responsive elemen... 80 8e-16
TC14252 homologue to UP|Q9ZR85 (Q9ZR85) Ethylene-responsive elem... 80 8e-16
TC14917 similar to GB|AAP37839.1|30725634|BT008480 At5g64750 {Ar... 78 2e-15
TC13814 similar to UP|Q7XAU4 (Q7XAU4) Ethylene responsive protei... 78 3e-15
TC15842 homologue to GB|BAB62912.1|15207792|AB055883 ERF domain ... 77 5e-15
TC15981 similar to UP|ERF3_ARATH (O80339) Ethylene responsive el... 77 7e-15
TC15600 homologue to UP|BAD01556 (BAD01556) ERF-like protein, pa... 75 2e-14
TC11381 similar to UP|Q8GZE9 (Q8GZE9) Ethylene responsive elemen... 71 3e-13
TC10014 similar to UP|Q8LLR3 (Q8LLR3) Ethylene-responsive elemen... 67 4e-12
AI967551 67 5e-12
TC8599 homologue to UP|Q40478 (Q40478) EREBP-4, partial (26%) 67 5e-12
TC14763 homologue to GB|AAM16262.1|20334802|AY094001 AT3g20310/M... 67 7e-12
TC8031 similar to UP|Q9FR33 (Q9FR33) Ripening regulated protein ... 66 9e-12
TC16026 similar to UP|Q9SGZ3 (Q9SGZ3) F28K19.29, partial (32%) 66 9e-12
TC14961 similar to UP|Q9LEM7 (Q9LEM7) AP2-domain DNA-binding pro... 66 9e-12
TC9975 similar to UP|AAR15448 (AAR15448) AP2 transcription facto... 66 1e-11
BP036840 65 2e-11
>TC7954 similar to UP|Q7XAU4 (Q7XAU4) Ethylene responsive protein, partial
(75%)
Length = 1708
Score = 81.3 bits (199), Expect = 3e-16
Identities = 46/108 (42%), Positives = 63/108 (57%), Gaps = 1/108 (0%)
Frame = +2
Query: 2 ARKRKVSEAVEEGSMVWDDM-MKEAASLGGARRVRKRFVGVRQRPSGRWVAEIKDTIQKI 60
A K K S + +GS + E A +++ + ++ G+RQRP G+W AEI+D + +
Sbjct: 413 AAKTKRSNSFSKGSTAAKSVESNEQAEKYASKKRKNQYRGIRQRPWGKWAAEIRDPRKGV 592
Query: 61 RVWLGTFDTAEEAARAYDEAACLLRGANTRTNFWPCSQSSTSPALSSK 108
RVWLGTF+TAEEAARAYD A +RG + NF P LSSK
Sbjct: 593 RVWLGTFNTAEEAARAYDAEARRIRGNKAKVNF------PVEPQLSSK 718
>TC9504 similar to UP|Q9SXS8 (Q9SXS8) Ethylene responsive element binding
factor, partial (35%)
Length = 633
Score = 80.9 bits (198), Expect = 4e-16
Identities = 40/65 (61%), Positives = 48/65 (73%)
Frame = +2
Query: 29 GGARRVRKRFVGVRQRPSGRWVAEIKDTIQKIRVWLGTFDTAEEAARAYDEAACLLRGAN 88
G A R+ GVR+RP GR+ AEI+D ++K RVWLGTFD+AEEAARAYD AA LRG
Sbjct: 407 GSAVLKEPRYRGVRKRPWGRFAAEIRDPMKKARVWLGTFDSAEEAARAYDAAARALRGPK 586
Query: 89 TRTNF 93
+TNF
Sbjct: 587 AKTNF 601
>TC14319 similar to UP|AP23_ARATH (P42736) AP2 domain transcription factor
RAP2.3 (Related to AP2 protein 3) (Cadmium-induced
protein AS30), partial (37%)
Length = 1130
Score = 80.5 bits (197), Expect = 5e-16
Identities = 53/143 (37%), Positives = 71/143 (49%), Gaps = 3/143 (2%)
Frame = +3
Query: 23 KEAASLGGARRVRKR---FVGVRQRPSGRWVAEIKDTIQKIRVWLGTFDTAEEAARAYDE 79
K GGA + R + G+RQRP G+W AEI+D + +RVWLGTF TAEEAARAYDE
Sbjct: 348 KSVVGSGGAGKKNARKNVYRGIRQRPWGKWAAEIRDPQKGVRVWLGTFPTAEEAARAYDE 527
Query: 80 AACLLRGANTRTNFWPCSQSSTSPALSSKITNLLLQRLKERNMNNSCSSFSSSKYTSLLG 139
AA +RG + NF P + ++ P QR SS SS T+ G
Sbjct: 528 AAKRIRGDKAKLNF-PDTVAAPPPTKK--------QRCLSPEQPEYSSSGSSGPTTTTTG 680
Query: 140 SPEERREQQRANDHDRQEMQQQV 162
S + D+ ++ Q+
Sbjct: 681 SSDSSAPIITTGLEDQPNLKLQI 749
>TC14193 similar to UP|Q8S2S7 (Q8S2S7) Ethylene responsive element binding
factor 4-like protein, partial (50%)
Length = 1128
Score = 79.7 bits (195), Expect = 8e-16
Identities = 45/95 (47%), Positives = 59/95 (61%)
Frame = +3
Query: 38 FVGVRQRPSGRWVAEIKDTIQKIRVWLGTFDTAEEAARAYDEAACLLRGANTRTNFWPCS 97
F GVR+RP GR+ AEI+D +K RVWLGTFDTAEEAARAYD AA RG +TNF
Sbjct: 177 FRGVRKRPWGRYAAEIRDPGKKSRVWLGTFDTAEEAARAYDAAARQFRGPKAKTNF---- 344
Query: 98 QSSTSPALSSKITNLLLQRLKERNMNNSCSSFSSS 132
S SP ++ ++++ +K + S+ SS
Sbjct: 345 PQSPSPDNNTGPNDVVVANVKANQSPSQSSTVESS 449
>TC14252 homologue to UP|Q9ZR85 (Q9ZR85) Ethylene-responsive element binding
protein homolog, partial (47%)
Length = 1174
Score = 79.7 bits (195), Expect = 8e-16
Identities = 46/83 (55%), Positives = 56/83 (67%), Gaps = 2/83 (2%)
Frame = +1
Query: 29 GGARRVRKRFVGVRQRPSGRWVAEIKDTIQKIRVWLGTFDTAEEAARAYDEAACLLRGAN 88
GG + V F GVR+RP GR+ AEI+D +K RVWLGTFDTAEEAARAYD AA RG
Sbjct: 178 GGVKEVH--FRGVRKRPWGRYAAEIRDPGKKSRVWLGTFDTAEEAARAYDTAAREFRGPK 351
Query: 89 TRTNF-WPCSQ-SSTSPALSSKI 109
+TNF +P + SP+ SS +
Sbjct: 352 AKTNFPFPSDNVKNQSPSQSSTV 420
>TC14917 similar to GB|AAP37839.1|30725634|BT008480 At5g64750 {Arabidopsis
thaliana;}, partial (17%)
Length = 1051
Score = 78.2 bits (191), Expect = 2e-15
Identities = 36/77 (46%), Positives = 52/77 (66%)
Frame = +3
Query: 35 RKRFVGVRQRPSGRWVAEIKDTIQKIRVWLGTFDTAEEAARAYDEAACLLRGANTRTNFW 94
++++ GVRQRP G+W AEI+D + RVWLGTF+TAEEAARAYD+A+ RG + NF
Sbjct: 201 KRKYRGVRQRPWGKWAAEIRDPFKAARVWLGTFNTAEEAARAYDQASLRFRGNKAKLNFP 380
Query: 95 PCSQSSTSPALSSKITN 111
+ P+++ T+
Sbjct: 381 ENVRLLQQPSINQPTTH 431
>TC13814 similar to UP|Q7XAU4 (Q7XAU4) Ethylene responsive protein, partial
(20%)
Length = 535
Score = 77.8 bits (190), Expect = 3e-15
Identities = 44/92 (47%), Positives = 60/92 (64%), Gaps = 1/92 (1%)
Frame = +2
Query: 3 RKRKVSEAVEEGSMVWDDMMKEAASLGGARRVRKR-FVGVRQRPSGRWVAEIKDTIQKIR 61
+KR + E E+GS +E G +RVRK + G+RQRP G+W AEI+D + +R
Sbjct: 236 KKRVMIEVEEKGSDEKSGESEEGIKKG--KRVRKNLYRGIRQRPWGKWAAEIRDPRKGVR 409
Query: 62 VWLGTFDTAEEAARAYDEAACLLRGANTRTNF 93
V LGTF+TAEEAA+AYD+AA +RG + NF
Sbjct: 410 VGLGTFNTAEEAAQAYDDAAKRIRGDKAKLNF 505
>TC15842 homologue to GB|BAB62912.1|15207792|AB055883 ERF domain protein12
{Arabidopsis thaliana;} , partial (41%)
Length = 746
Score = 77.0 bits (188), Expect = 5e-15
Identities = 40/75 (53%), Positives = 49/75 (65%)
Frame = +2
Query: 21 MMKEAASLGGARRVRKRFVGVRQRPSGRWVAEIKDTIQKIRVWLGTFDTAEEAARAYDEA 80
++ AS + + GVR+RP GR+ AEI+D +K RVWLGTFDT EEAA AYD A
Sbjct: 134 LLPSMASSSSSASREGHYRGVRKRPWGRYAAEIRDPWKKTRVWLGTFDTPEEAALAYDGA 313
Query: 81 ACLLRGANTRTNFWP 95
A LRGA +TNF P
Sbjct: 314 ARSLRGAKAKTNFPP 358
>TC15981 similar to UP|ERF3_ARATH (O80339) Ethylene responsive element
binding factor 3 (AtERF3), partial (45%)
Length = 671
Score = 76.6 bits (187), Expect = 7e-15
Identities = 37/57 (64%), Positives = 44/57 (76%)
Frame = +1
Query: 37 RFVGVRQRPSGRWVAEIKDTIQKIRVWLGTFDTAEEAARAYDEAACLLRGANTRTNF 93
RF GVR+RP GR+ AEI+D +K RVWLGTFD+AE+AARAYD AA RG +TNF
Sbjct: 238 RFRGVRKRPWGRFAAEIRDPWKKARVWLGTFDSAEDAARAYDAAARSFRGPKAKTNF 408
>TC15600 homologue to UP|BAD01556 (BAD01556) ERF-like protein, partial (36%)
Length = 570
Score = 75.5 bits (184), Expect = 2e-14
Identities = 34/62 (54%), Positives = 45/62 (71%)
Frame = +3
Query: 32 RRVRKRFVGVRQRPSGRWVAEIKDTIQKIRVWLGTFDTAEEAARAYDEAACLLRGANTRT 91
R+ + + G+RQRP G+W AEI+D + +RVWLGTF+TAEEAARAYD A +RG +
Sbjct: 378 RQRKNLYRGIRQRPWGKWAAEIRDPRKGVRVWLGTFNTAEEAARAYDREARKIRGKKAKV 557
Query: 92 NF 93
NF
Sbjct: 558 NF 563
>TC11381 similar to UP|Q8GZE9 (Q8GZE9) Ethylene responsive element binding
protein, partial (36%)
Length = 497
Score = 71.2 bits (173), Expect = 3e-13
Identities = 35/76 (46%), Positives = 52/76 (68%), Gaps = 1/76 (1%)
Frame = +3
Query: 19 DDMMKEAASLGGARRVRKRFVGVRQRPSGRWVAEIKDTIQK-IRVWLGTFDTAEEAARAY 77
+++ E G + R K + GVR+RP G++ AEI+D+ + +RVWLGTFD+AE AA AY
Sbjct: 240 EEVSSEENGCGSSPRKEKSYRGVRRRPWGKFAAEIRDSTRNGVRVWLGTFDSAEAAALAY 419
Query: 78 DEAACLLRGANTRTNF 93
D+AA +RG++ NF
Sbjct: 420 DQAAFAMRGSSAILNF 467
>TC10014 similar to UP|Q8LLR3 (Q8LLR3) Ethylene-responsive element binding
protein 1, partial (33%)
Length = 580
Score = 67.4 bits (163), Expect = 4e-12
Identities = 36/75 (48%), Positives = 47/75 (62%), Gaps = 1/75 (1%)
Frame = +3
Query: 36 KRFVGVRQRPSGRWVAEIKDTIQK-IRVWLGTFDTAEEAARAYDEAACLLRGANTRTNFW 94
KRF GVR+RP G++ AEI+D + RVWLGT++T EEA AYD AA +RG + NF
Sbjct: 291 KRFRGVRRRPWGKFAAEIRDPKKNGARVWLGTYETEEEAGLAYDRAAFKIRGRKAKLNFP 470
Query: 95 PCSQSSTSPALSSKI 109
S T P +S +
Sbjct: 471 HLIGSDTQPETASVV 515
>AI967551
Length = 466
Score = 67.0 bits (162), Expect = 5e-12
Identities = 33/59 (55%), Positives = 45/59 (75%), Gaps = 1/59 (1%)
Frame = +2
Query: 36 KRFVGVRQRPSGRWVAEIKDTIQK-IRVWLGTFDTAEEAARAYDEAACLLRGANTRTNF 93
K + GVR+RP G++ AEI+D+ + IRVWLGTFD+AE AA AYD+AA +RG++ NF
Sbjct: 278 KAYRGVRRRPWGKFAAEIRDSTRHGIRVWLGTFDSAEAAALAYDQAAFAMRGSSATLNF 454
>TC8599 homologue to UP|Q40478 (Q40478) EREBP-4, partial (26%)
Length = 632
Score = 67.0 bits (162), Expect = 5e-12
Identities = 38/84 (45%), Positives = 53/84 (62%), Gaps = 1/84 (1%)
Frame = +3
Query: 36 KRFVGVRQRPSGRWVAEIKDTIQK-IRVWLGTFDTAEEAARAYDEAACLLRGANTRTNFW 94
K + GVR+RP G++ AEI+D +K RVWLGTFDTA EAA+AYD AA +RG+ NF
Sbjct: 384 KHYRGVRRRPWGKFAAEIRDPNRKGSRVWLGTFDTAIEAAKAYDRAAFKMRGSRAILNF- 560
Query: 95 PCSQSSTSPALSSKITNLLLQRLK 118
P + S + N+ +R++
Sbjct: 561 PLEVGNCHEEEESSVVNVGEKRIR 632
>TC14763 homologue to GB|AAM16262.1|20334802|AY094001 AT3g20310/MQC12_6
{Arabidopsis thaliana;}, partial (18%)
Length = 618
Score = 66.6 bits (161), Expect = 7e-12
Identities = 30/42 (71%), Positives = 37/42 (87%)
Frame = +1
Query: 37 RFVGVRQRPSGRWVAEIKDTIQKIRVWLGTFDTAEEAARAYD 78
R+ GVR+RP GR+ AEI+D ++K RVWLGTFDTAE+AARAYD
Sbjct: 490 RYRGVRKRPWGRFAAEIRDPMKKARVWLGTFDTAEDAARAYD 615
>TC8031 similar to UP|Q9FR33 (Q9FR33) Ripening regulated protein DDTFR10/A
(Fragment), partial (44%)
Length = 1109
Score = 66.2 bits (160), Expect = 9e-12
Identities = 34/60 (56%), Positives = 44/60 (72%), Gaps = 1/60 (1%)
Frame = +2
Query: 35 RKRFVGVRQRPSGRWVAEIKDTIQK-IRVWLGTFDTAEEAARAYDEAACLLRGANTRTNF 93
++ + GVRQRP G++ AEI+D ++ RVWLGTFDTA EAA+AYD AA LRG+ NF
Sbjct: 542 KQHYRGVRQRPWGKFAAEIRDPNKRGSRVWLGTFDTAIEAAKAYDRAAFKLRGSKAILNF 721
>TC16026 similar to UP|Q9SGZ3 (Q9SGZ3) F28K19.29, partial (32%)
Length = 1183
Score = 66.2 bits (160), Expect = 9e-12
Identities = 33/51 (64%), Positives = 39/51 (75%)
Frame = +2
Query: 36 KRFVGVRQRPSGRWVAEIKDTIQKIRVWLGTFDTAEEAARAYDEAACLLRG 86
K + GVRQR G+WVAEI+ + R+WLGTFDTAEEAA AYD+AA LRG
Sbjct: 1025 KLYRGVRQRHWGKWVAEIRLPKNRTRLWLGTFDTAEEAALAYDKAAYKLRG 1177
>TC14961 similar to UP|Q9LEM7 (Q9LEM7) AP2-domain DNA-binding protein,
partial (27%)
Length = 1135
Score = 66.2 bits (160), Expect = 9e-12
Identities = 34/65 (52%), Positives = 43/65 (65%)
Frame = +1
Query: 29 GGARRVRKRFVGVRQRPSGRWVAEIKDTIQKIRVWLGTFDTAEEAARAYDEAACLLRGAN 88
GG R + GVRQR G+WVAEI++ + R+WLGTF TA AA AYDEAA + G++
Sbjct: 361 GGPENSRCNYRGVRQRTWGKWVAEIREPNRGKRLWLGTFATAIGAALAYDEAARAMYGSS 540
Query: 89 TRTNF 93
R NF
Sbjct: 541 ARLNF 555
>TC9975 similar to UP|AAR15448 (AAR15448) AP2 transcription factor, partial
(41%)
Length = 865
Score = 65.9 bits (159), Expect = 1e-11
Identities = 41/94 (43%), Positives = 57/94 (60%), Gaps = 1/94 (1%)
Frame = +1
Query: 36 KRFVGVRQRPSGRWVAEIKDTIQK-IRVWLGTFDTAEEAARAYDEAACLLRGANTRTNFW 94
+R+ GVR+RP G++ AEI+D +K RVWLGTFDT +AA+AYD AA +RG NF
Sbjct: 178 RRYRGVRRRPWGKFAAEIRDPTRKGTRVWLGTFDTEIDAAKAYDYAAFRMRGRKAILNF- 354
Query: 95 PCSQSSTSPALSSKITNLLLQRLKERNMNNSCSS 128
P SP K +N +R +E+ + + SS
Sbjct: 355 PLEAGKASP----KPSNSGRKRGREQRESGTSSS 444
>BP036840
Length = 584
Score = 65.5 bits (158), Expect = 2e-11
Identities = 35/75 (46%), Positives = 47/75 (62%)
Frame = +3
Query: 19 DDMMKEAASLGGARRVRKRFVGVRQRPSGRWVAEIKDTIQKIRVWLGTFDTAEEAARAYD 78
D++ AA GG R R GVR+R GRWV+EI++ +K R+WLG+F E AA+AYD
Sbjct: 36 DNVTASAAVRGGTRHPVYR--GVRKRRWGRWVSEIREPRKKSRIWLGSFPVPEMAAKAYD 209
Query: 79 EAACLLRGANTRTNF 93
AA L+G + NF
Sbjct: 210 VAAYCLKGRKAQLNF 254
Database: LJGI
Posted date: Jul 30, 2004 11:16 AM
Number of letters in database: 14,692,800
Number of sequences in database: 28,460
Lambda K H
0.311 0.127 0.358
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 6,486,060
Number of Sequences: 28460
Number of extensions: 81319
Number of successful extensions: 486
Number of sequences better than 10.0: 72
Number of HSP's better than 10.0 without gapping: 471
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 476
length of query: 411
length of database: 4,897,600
effective HSP length: 92
effective length of query: 319
effective length of database: 2,279,280
effective search space: 727090320
effective search space used: 727090320
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 16 ( 7.2 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 42 (21.8 bits)
S2: 56 (26.2 bits)
Medicago: description of AC119412.1


