

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC119411.2 - phase: 0
(302 letters)
Database: LJGI
28,460 sequences; 14,692,800 total letters
Searching..................................................done
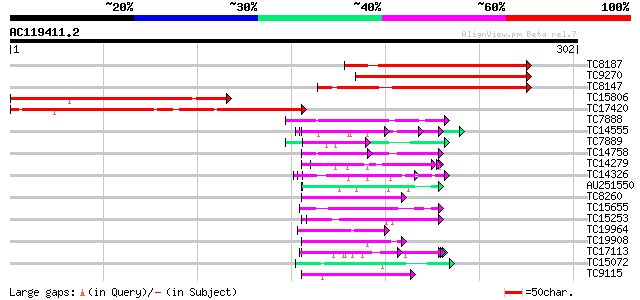
Score E
Sequences producing significant alignments: (bits) Value
TC8187 similar to UP|Q9FQ74 (Q9FQ74) Biotin carboxyl carrier pro... 155 6e-39
TC9270 similar to UP|Q9FQ74 (Q9FQ74) Biotin carboxyl carrier pro... 152 5e-38
TC8147 similar to UP|BCCP_SOYBN (Q42783) Biotin carboxyl carrier... 132 9e-32
TC15806 similar to UP|Q9FQ74 (Q9FQ74) Biotin carboxyl carrier pr... 130 4e-31
TC17420 similar to UP|Q9FQ74 (Q9FQ74) Biotin carboxyl carrier pr... 128 1e-30
TC7888 similar to UP|Q40336 (Q40336) Proline-rich cell wall prot... 49 8e-07
TC14555 similar to UP|Q43558 (Q43558) Proline rich protein precu... 47 3e-06
TC7889 similar to UP|Q40336 (Q40336) Proline-rich cell wall prot... 47 4e-06
TC14758 similar to UP|O24601 (O24601) Glycine-rich RNA binding p... 47 5e-06
TC14279 similar to UP|Q09085 (Q09085) Hydroxyproline-rich glycop... 46 7e-06
TC14326 weakly similar to UP|EXLP_TOBAC (Q03211) Pistil-specific... 46 9e-06
AU251550 45 2e-05
TC8260 similar to UP|Q94ES8 (Q94ES8) Nodule extensin (Fragment),... 45 2e-05
TC15655 similar to UP|Q8LE99 (Q8LE99) Transketolase-like protein... 45 2e-05
TC15253 weakly similar to UP|Q9ZT16 (Q9ZT16) Arabinogalactan-pro... 45 2e-05
TC19964 similar to UP|Q40363 (Q40363) NuM1 protein, partial (11%) 44 3e-05
TC19908 similar to GB|CAA60022.1|791150|VURNEXT26 extensin-like ... 44 4e-05
TC17113 weakly similar to UP|Q9LFU8 (Q9LFU8) Proline-rich protei... 44 4e-05
TC15072 43 6e-05
TC9115 similar to PIR|T00425|T00425 photolyase/blue-light recept... 43 6e-05
>TC8187 similar to UP|Q9FQ74 (Q9FQ74) Biotin carboxyl carrier protein
subunit , partial (33%)
Length = 602
Score = 155 bits (393), Expect = 6e-39
Identities = 77/100 (77%), Positives = 83/100 (83%)
Frame = +3
Query: 179 PPPPVVASTPPSAPPSNVVPALPPAKTNASSHPQLKCPMAGTFYRCPGPGEPPFVKVGDQ 238
PPP VAS P A PALPP K+++SS PQLKCPMAGTFYR P PGEPPFVKVGD+
Sbjct: 3 PPPLAVASAPTKA-----APALPPGKSSSSSLPQLKCPMAGTFYRSPAPGEPPFVKVGDK 167
Query: 239 VQKGQVVCIIEAMKLMNEIEADRSGTIVEVLVEDGKPVSV 278
VQKGQV+CIIEAMKLMNEIEAD+SGTI EVL EDGKPVSV
Sbjct: 168 VQKGQVICIIEAMKLMNEIEADKSGTIAEVLAEDGKPVSV 287
>TC9270 similar to UP|Q9FQ74 (Q9FQ74) Biotin carboxyl carrier protein
subunit , partial (36%)
Length = 625
Score = 152 bits (385), Expect = 5e-38
Identities = 73/95 (76%), Positives = 80/95 (83%), Gaps = 1/95 (1%)
Frame = +2
Query: 185 ASTPPSAPPSNVVPALP-PAKTNASSHPQLKCPMAGTFYRCPGPGEPPFVKVGDQVQKGQ 243
AS P +PPS PALP P K + SSHP LKCPMAGTFYR P PGEPPFVKVGD+VQKGQ
Sbjct: 2 ASAPVXSPPSKAAPALPSPGKESKSSHPPLKCPMAGTFYRSPAPGEPPFVKVGDKVQKGQ 181
Query: 244 VVCIIEAMKLMNEIEADRSGTIVEVLVEDGKPVSV 278
V+CIIEAMKLMNEIEAD+SGTI E++ EDGKPVSV
Sbjct: 182 VICIIEAMKLMNEIEADQSGTITEIVAEDGKPVSV 286
>TC8147 similar to UP|BCCP_SOYBN (Q42783) Biotin carboxyl carrier protein
of acetyl-CoA carboxylase, chloroplast precursor (BCCP)
, partial (42%)
Length = 731
Score = 132 bits (331), Expect = 9e-32
Identities = 70/114 (61%), Positives = 81/114 (70%)
Frame = +3
Query: 165 PYPAYPPSYQAPLPPPPPVVASTPPSAPPSNVVPALPPAKTNASSHPQLKCPMAGTFYRC 224
P P PP A P P VA++ PS S + K+ SSHP LK PMAGTFYR
Sbjct: 3 PSPTMPPV--ATASNPAPAVAASTPSPSTSLAI------KSAKSSHPPLKSPMAGTFYRS 158
Query: 225 PGPGEPPFVKVGDQVQKGQVVCIIEAMKLMNEIEADRSGTIVEVLVEDGKPVSV 278
P PGEP FVKVGD+V+KGQV+CIIEAMKLMNEIEAD+SGT+V+V+ EDGK VSV
Sbjct: 159 PAPGEPAFVKVGDKVKKGQVLCIIEAMKLMNEIEADQSGTVVDVIAEDGKSVSV 320
>TC15806 similar to UP|Q9FQ74 (Q9FQ74) Biotin carboxyl carrier protein
subunit , partial (28%)
Length = 619
Score = 130 bits (326), Expect = 4e-31
Identities = 75/124 (60%), Positives = 88/124 (70%), Gaps = 6/124 (4%)
Frame = +3
Query: 1 MASFTTVPCPKCLTFHHLGLKSQTSS-RNVQ----LGFMKSQSFGSLSCDLVS-NGIQCL 54
MAS TVPCPKCL+F GL S+T +N+Q LG +S SFGSLS D + NGIQ L
Sbjct: 252 MASSFTVPCPKCLSFTGSGLNSRTPPPQNMQIRSGLGLKESGSFGSLSHDSAAPNGIQAL 431
Query: 55 ERKQFSVWRSQALPNEVATIENSSNSVPVLINEPNGALPKEKDNHNGKPPGPSASTDASS 114
RKQ+SVW+ QALP+E AT+ NSSNS PVL+ EP A +EKD NGKP GPS STDASS
Sbjct: 432 NRKQYSVWKFQALPSEAATVGNSSNSAPVLVKEPKVASLEEKD--NGKPSGPSTSTDASS 605
Query: 115 VSTF 118
+S F
Sbjct: 606 ISAF 617
>TC17420 similar to UP|Q9FQ74 (Q9FQ74) Biotin carboxyl carrier protein
subunit , partial (18%)
Length = 600
Score = 128 bits (322), Expect = 1e-30
Identities = 85/160 (53%), Positives = 105/160 (65%), Gaps = 2/160 (1%)
Frame = -3
Query: 1 MASFTTVPCPKCLTFHHLGLKS--QTSSRNVQLGFMKSQSFGSLSCDLVSNGIQCLERKQ 58
MASF+ VPC K LGL+S + +NV QSFGSL S+ I CL RKQ
Sbjct: 469 MASFS-VPCSKSPAPAPLGLRSSHKVEFQNVLCLKPSQQSFGSLCARPASSEIGCLNRKQ 293
Query: 59 FSVWRSQALPNEVATIENSSNSVPVLINEPNGALPKEKDNHNGKPPGPSASTDASSVSTF 118
SV + QA NE AT+ NS NS P+ +N A KEK+++NG + + DASS+STF
Sbjct: 292 LSVQKLQAQRNEAATVGNS-NSAPLPVNV---ASSKEKEDYNGSVSCGTVA-DASSISTF 128
Query: 119 MDQVSELVKLVDSRDIMELELKQAGYELMIRKKEALQPPP 158
M QVS+LVKLVDSRDI+EL+LKQ+ ELMIRK+EAL PPP
Sbjct: 127 MAQVSDLVKLVDSRDIVELQLKQSDCELMIRKREALHPPP 8
>TC7888 similar to UP|Q40336 (Q40336) Proline-rich cell wall protein,
partial (60%)
Length = 913
Score = 49.3 bits (116), Expect = 8e-07
Identities = 34/87 (39%), Positives = 39/87 (44%)
Frame = +3
Query: 148 IRKKEALQPPPVSQQPFPYPAYPPSYQAPLPPPPPVVASTPPSAPPSNVVPALPPAKTNA 207
I K + PPV + P PY PP P P PPVV TPP P VVP PP
Sbjct: 483 IVKPPYVPKPPVVKPP-PYVPKPPVVSPPYVPKPPVVPVTPPYVPKPPVVPVTPP---YV 650
Query: 208 SSHPQLKCPMAGTFYRCPGPGEPPFVK 234
P +K P+ Y P +PP VK
Sbjct: 651 PKPPIVKPPIV---YPPPYVPKPPIVK 722
Score = 37.4 bits (85), Expect = 0.003
Identities = 33/95 (34%), Positives = 38/95 (39%), Gaps = 1/95 (1%)
Frame = +3
Query: 152 EALQPPPVSQQPFPYPAYPPSYQAPLPPPPPVVASTPPSAP-PSNVVPALPPAKTNASSH 210
E PP S +P YP PP + P P PPVV PP P P V P P +
Sbjct: 309 EPCPPPKPSPKPPHYPK-PPVVKPPHVPKPPVV--KPPHVPKPPIVKPPYVPKPPHVPKP 479
Query: 211 PQLKCPMAGTFYRCPGPGEPPFVKVGDQVQKGQVV 245
P +K P +PP VK V K VV
Sbjct: 480 PIVKPPYV---------PKPPVVKPPPYVPKPPVV 557
Score = 35.8 bits (81), Expect = 0.009
Identities = 26/81 (32%), Positives = 33/81 (40%), Gaps = 9/81 (11%)
Frame = +3
Query: 163 PFPYPAYPPSYQAPLPP------PPPVV---ASTPPSAPPSNVVPALPPAKTNASSHPQL 213
P+P P+ PP + PP PP+V PP PP P P K S P+
Sbjct: 135 PYPAPSKPPKHPIVKPPVHKPPQHPPIVKPPVHKPPQHPPIVKPPVHKPPKYPPSHGPE- 311
Query: 214 KCPMAGTFYRCPGPGEPPFVK 234
CP + P +PP VK
Sbjct: 312 PCPPPKPSPKPPHYPKPPVVK 374
Score = 35.8 bits (81), Expect = 0.009
Identities = 23/74 (31%), Positives = 29/74 (39%), Gaps = 19/74 (25%)
Frame = +3
Query: 148 IRKKEALQPPPVSQQPFPYPAY---------PPSYQAPLPPPPPVV----------ASTP 188
+ K ++PP V + P P Y PP + P P PPVV +P
Sbjct: 384 VPKPPVVKPPHVPKPPIVKPPYVPKPPHVPKPPIVKPPYVPKPPVVKPPPYVPKPPVVSP 563
Query: 189 PSAPPSNVVPALPP 202
P P VVP PP
Sbjct: 564 PYVPKPPVVPVTPP 605
Score = 35.8 bits (81), Expect = 0.009
Identities = 25/83 (30%), Positives = 31/83 (37%)
Frame = +3
Query: 151 KEALQPPPVSQQPFPYPAYPPSYQAPLPPPPPVVASTPPSAPPSNVVPALPPAKTNASSH 210
K + PP+ P PY PP + P+ PPP V + P PP P P S
Sbjct: 654 KPPIVKPPIVYPP-PYVPKPPIVKPPIVYPPPYVPTPPIVKPPIVYPPPYVPLPPVVPSP 830
Query: 211 PQLKCPMAGTFYRCPGPGEPPFV 233
P + P P PP V
Sbjct: 831 PIVTPPTPTPPIVTPPTPTPPVV 899
Score = 35.4 bits (80), Expect = 0.012
Identities = 25/60 (41%), Positives = 28/60 (46%), Gaps = 6/60 (10%)
Frame = +3
Query: 148 IRKKEALQPPPVSQQP---FPYPAYPPSYQAPLPP---PPPVVASTPPSAPPSNVVPALP 201
I K + PPP P P YPP Y PLPP PP+V TPP+ P V P P
Sbjct: 714 IVKPPIVYPPPYVPTPPIVKPPIVYPPPY-VPLPPVVPSPPIV--TPPTPTPPIVTPPTP 884
Score = 32.0 bits (71), Expect = 0.13
Identities = 17/46 (36%), Positives = 19/46 (40%)
Frame = +3
Query: 157 PPVSQQPFPYPAYPPSYQAPLPPPPPVVASTPPSAPPSNVVPALPP 202
PPV PY PP + P+ PPP V P PP P P
Sbjct: 618 PPVVPVTPPYVPKPPIVKPPIVYPPPYVPKPPIVKPPIVYPPPYVP 755
Score = 30.8 bits (68), Expect = 0.29
Identities = 20/51 (39%), Positives = 23/51 (44%)
Frame = +3
Query: 148 IRKKEALQPPPVSQQPFPYPAYPPSYQAPLPPPPPVVASTPPSAPPSNVVP 198
I K + PPP P P+ PP P P PP V TPP+ P V P
Sbjct: 765 IVKPPIVYPPPYVPLPPVVPS-PPIVTPPTPTPPIV---TPPTPTPPVVTP 905
Score = 30.4 bits (67), Expect = 0.38
Identities = 24/80 (30%), Positives = 30/80 (37%)
Frame = +3
Query: 155 QPPPVSQQPFPYPAYPPSYQAPLPPPPPVVASTPPSAPPSNVVPALPPAKTNASSHPQLK 214
+PP VS P P P +P PP V + P P V P + P +K
Sbjct: 543 KPPVVSPPYVPKPPVVPVTPPYVPKPPVVPVTPPYVPKPPIVKPPIVYPPPYVPKPPIVK 722
Query: 215 CPMAGTFYRCPGPGEPPFVK 234
P+ Y P PP VK
Sbjct: 723 PPIV---YPPPYVPTPPIVK 773
Score = 29.3 bits (64), Expect = 0.86
Identities = 24/81 (29%), Positives = 31/81 (37%), Gaps = 1/81 (1%)
Frame = +3
Query: 155 QPPPVSQQPFPYPA-YPPSYQAPLPPPPPVVASTPPSAPPSNVVPALPPAKTNASSHPQL 213
Q PP+ + P P +PP + P+ PP PPS P P P K P +
Sbjct: 201 QHPPIVKPPVHKPPQHPPIVKPPVHKPP----KYPPSHGPEPCPPPKPSPKPPHYPKPPV 368
Query: 214 KCPMAGTFYRCPGPGEPPFVK 234
P P +PP VK
Sbjct: 369 VKP--------PHVPKPPVVK 407
Score = 26.9 bits (58), Expect = 4.2
Identities = 22/77 (28%), Positives = 28/77 (35%)
Frame = +3
Query: 157 PPVSQQPFPYPAYPPSYQAPLPPPPPVVASTPPSAPPSNVVPALPPAKTNASSHPQLKCP 216
PP +P P PP P PP+V P P V P + + P +K P
Sbjct: 600 PPYVPKPPVVPVTPPYVPKPPIVKPPIVYPPPYVPKPPIVKPPIVYPPPYVPTPPIVKPP 779
Query: 217 MAGTFYRCPGPGEPPFV 233
+ Y P PP V
Sbjct: 780 IV---YPPPYVPLPPVV 821
>TC14555 similar to UP|Q43558 (Q43558) Proline rich protein precursor,
partial (43%)
Length = 1038
Score = 47.4 bits (111), Expect = 3e-06
Identities = 24/48 (50%), Positives = 26/48 (54%), Gaps = 1/48 (2%)
Frame = +2
Query: 156 PPPVSQQPFPYPAYPPSYQAPLP-PPPPVVASTPPSAPPSNVVPALPP 202
PPPV P P PP+ +P P PP V STPP AP S PA PP
Sbjct: 287 PPPVQSSPPPASTPPPAQSSPPPVSSPPPVQSTPPPAPASTPPPASPP 430
Score = 43.5 bits (101), Expect = 4e-05
Identities = 30/93 (32%), Positives = 35/93 (37%), Gaps = 6/93 (6%)
Frame = +2
Query: 156 PPPVSQQP------FPYPAYPPSYQAPLPPPPPVVASTPPSAPPSNVVPALPPAKTNASS 209
PPPV P P PA PP + P PPP A+ PP+ P VPA PA A
Sbjct: 365 PPPVQSTPPPAPASTPPPASPPPFSPPPATPPPPAATPPPALTP---VPATSPAPAPAKV 535
Query: 210 HPQLKCPMAGTFYRCPGPGEPPFVKVGDQVQKG 242
+ P P P P V + G
Sbjct: 536 KSKSPAP-------APAPALAPVVSLSPSEAPG 613
Score = 42.0 bits (97), Expect = 1e-04
Identities = 26/71 (36%), Positives = 30/71 (41%), Gaps = 5/71 (7%)
Frame = +2
Query: 155 QPPPVSQQPFPYPAYPPSYQAPLPP---PPPVVASTPP--SAPPSNVVPALPPAKTNASS 209
QPP P P + PP Q+ PP PPP +S PP S PP P PA T +
Sbjct: 242 QPPAAQSSPPPAQSSPPPVQSSPPPASTPPPAQSSPPPVSSPPPVQSTPPPAPASTPPPA 421
Query: 210 HPQLKCPMAGT 220
P P T
Sbjct: 422 SPPPFSPPPAT 454
Score = 41.6 bits (96), Expect = 2e-04
Identities = 27/70 (38%), Positives = 32/70 (45%), Gaps = 2/70 (2%)
Frame = +2
Query: 153 ALQPPPVSQQPFPYPAYPPSYQAPLPPPPPVVASTPP--SAPPSNVVPALPPAKTNASSH 210
A PPP P P + PP PPP ASTPP S PP + PA PP A++
Sbjct: 317 ASTPPPAQSSPPPVSSPPPVQST----PPPAPASTPPPASPPPFSPPPATPPPP--AATP 478
Query: 211 PQLKCPMAGT 220
P P+ T
Sbjct: 479 PPALTPVPAT 508
Score = 40.4 bits (93), Expect = 4e-04
Identities = 28/78 (35%), Positives = 33/78 (41%), Gaps = 2/78 (2%)
Frame = +2
Query: 156 PPPVSQQPFPYPAYPPSYQAPLPPP--PPVVASTPPSAPPSNVVPALPPAKTNASSHPQL 213
PPPVS P P + PP A PPP PP + P + PP P PPA T +
Sbjct: 347 PPPVSSPP-PVQSTPPPAPASTPPPASPPPFSPPPATPPPPAATP--PPALTPVPATSPA 517
Query: 214 KCPMAGTFYRCPGPGEPP 231
P A + P P P
Sbjct: 518 PAP-AKVKSKSPAPAPAP 568
Score = 37.7 bits (86), Expect = 0.002
Identities = 24/73 (32%), Positives = 32/73 (42%), Gaps = 4/73 (5%)
Frame = +2
Query: 163 PFPYPAYPPSYQAPLPP----PPPVVASTPPSAPPSNVVPALPPAKTNASSHPQLKCPMA 218
P P PP+ Q+ PP PPPV +S PP++ P + PP SS P ++
Sbjct: 224 PTPTTPQPPAAQSSPPPAQSSPPPVQSSPPPASTPPPAQSSPPP----VSSPPPVQSTPP 391
Query: 219 GTFYRCPGPGEPP 231
P P PP
Sbjct: 392 PAPASTPPPASPP 430
Score = 37.4 bits (85), Expect = 0.003
Identities = 22/64 (34%), Positives = 23/64 (35%)
Frame = +2
Query: 153 ALQPPPVSQQPFPYPAYPPSYQAPLPPPPPVVASTPPSAPPSNVVPALPPAKTNASSHPQ 212
A PPP S PF P P PPPP P P PA PAK + S
Sbjct: 401 ASTPPPASPPPFSPP--------PATPPPPAATPPPALTPVPATSPAPAPAKVKSKSPAP 556
Query: 213 LKCP 216
P
Sbjct: 557 APAP 568
Score = 36.2 bits (82), Expect = 0.007
Identities = 23/79 (29%), Positives = 32/79 (40%), Gaps = 3/79 (3%)
Frame = +2
Query: 156 PPPVSQQPFPYPAYPPSYQAPLPP---PPPVVASTPPSAPPSNVVPALPPAKTNASSHPQ 212
P P + QP + PP Q+ PP PP ++ PP+ V + PP ++ P
Sbjct: 224 PTPTTPQPPAAQSSPPPAQSSPPPVQSSPPPASTPPPAQSSPPPVSSPPPVQSTPPPAPA 403
Query: 213 LKCPMAGTFYRCPGPGEPP 231
P A P P PP
Sbjct: 404 STPPPASPPPFSPPPATPP 460
Score = 35.0 bits (79), Expect = 0.016
Identities = 25/77 (32%), Positives = 30/77 (38%), Gaps = 5/77 (6%)
Frame = +2
Query: 160 SQQPFPYPAYPPSYQAPLPPPPPVVASTPPSA--PPSNVVPALPPAKT---NASSHPQLK 214
+Q P P P+ P P PP S+PP A P V + PPA T SS P +
Sbjct: 182 AQSPSNSPTTSPAPPTPTTPQPPAAQSSPPPAQSSPPPVQSSPPPASTPPPAQSSPPPVS 361
Query: 215 CPMAGTFYRCPGPGEPP 231
P P P P
Sbjct: 362 SPPPVQSTPPPAPASTP 412
Score = 31.6 bits (70), Expect = 0.17
Identities = 25/73 (34%), Positives = 33/73 (44%), Gaps = 17/73 (23%)
Frame = +2
Query: 153 ALQPPPVSQQP---FPYPAYPPS-----------YQAPLPPPPPVVASTPPSAPP---SN 195
A PPP + P P PA P+ AP P PVV+ +P AP S+
Sbjct: 449 ATPPPPAATPPPALTPVPATSPAPAPAKVKSKSPAPAPAPALAPVVSLSPSEAPGPSLSS 628
Query: 196 VVPALPPAKTNAS 208
+ PAL PA ++ S
Sbjct: 629 LSPALSPAASDDS 667
>TC7889 similar to UP|Q40336 (Q40336) Proline-rich cell wall protein,
partial (66%)
Length = 1158
Score = 47.0 bits (110), Expect = 4e-06
Identities = 32/87 (36%), Positives = 35/87 (39%)
Frame = +1
Query: 148 IRKKEALQPPPVSQQPFPYPAYPPSYQAPLPPPPPVVASTPPSAPPSNVVPALPPAKTNA 207
I K + PPV + P PY PP P P PPVV TPP P VVP PP
Sbjct: 52 IVKPPYVPKPPVVKPP-PYVPKPPVVSPPYVPKPPVVPVTPPYVPKPPVVPVTPP----- 213
Query: 208 SSHPQLKCPMAGTFYRCPGPGEPPFVK 234
P Y P +PP VK
Sbjct: 214 ------YVPKPPIVYPPPYVPKPPIVK 276
Score = 40.0 bits (92), Expect = 5e-04
Identities = 20/40 (50%), Positives = 23/40 (57%), Gaps = 4/40 (10%)
Frame = +1
Query: 157 PPVSQQPFPYP--AYPPS--YQAPLPPPPPVVASTPPSAP 192
PP+ P P P PPS + P PPPPPVV S PP+ P
Sbjct: 412 PPIVTPPTPTPPVVTPPSPPTETPCPPPPPVVPSPPPAQP 531
Score = 36.2 bits (82), Expect = 0.007
Identities = 34/114 (29%), Positives = 42/114 (36%), Gaps = 16/114 (14%)
Frame = +1
Query: 156 PPPVSQQPFPYPAYPPSYQAPLPPPPPVVAS----------TPPSAPPSNVVPALPPAKT 205
P P P PY PP + P+ PPP V + PP P VVP+ PP T
Sbjct: 220 PKPPIVYPPPYVPKPPIVKPPIVYPPPYVPTPPIVKPPIVYPPPYVPLPPVVPS-PPIVT 396
Query: 206 NASSHPQLKCPMAGT--FYRCPGPGE----PPFVKVGDQVQKGQVVCIIEAMKL 253
+ P + P T P P PP V Q C I+ +KL
Sbjct: 397 PPTPTPPIVTPPTPTPPVVTPPSPPTETPCPPPPPVVPSPPPAQPTCPIDTLKL 558
Score = 30.8 bits (68), Expect = 0.29
Identities = 20/49 (40%), Positives = 21/49 (42%), Gaps = 1/49 (2%)
Frame = +1
Query: 157 PPVSQQPFPYPAYPPS-YQAPLPPPPPVVASTPPSAPPSNVVPALPPAK 204
PPV PY PP Y P P PP+V PP P VP P K
Sbjct: 187 PPVVPVTPPYVPKPPIVYPPPYVPKPPIV--KPPIVYPPPYVPTPPIVK 327
Score = 27.7 bits (60), Expect = 2.5
Identities = 18/49 (36%), Positives = 19/49 (38%), Gaps = 3/49 (6%)
Frame = +1
Query: 157 PPVSQQPFPYPAYPPSYQAPLPP---PPPVVASTPPSAPPSNVVPALPP 202
PP +P P PP P PP PPP V P PP P P
Sbjct: 169 PPYVPKPPVVPVTPP--YVPKPPIVYPPPYVPKPPIVKPPIVYPPPYVP 309
Score = 27.7 bits (60), Expect = 2.5
Identities = 26/93 (27%), Positives = 34/93 (35%), Gaps = 7/93 (7%)
Frame = +1
Query: 148 IRKKEALQPPPVSQQPF-----PYPAYPPSYQA--PLPPPPPVVASTPPSAPPSNVVPAL 200
+ K + PP V + P PY PP P P PP+V P P V P +
Sbjct: 106 VPKPPVVSPPYVPKPPVVPVTPPYVPKPPVVPVTPPYVPKPPIVYPPPYVPKPPIVKPPI 285
Query: 201 PPAKTNASSHPQLKCPMAGTFYRCPGPGEPPFV 233
+ P +K P+ Y P PP V
Sbjct: 286 VYPPPYVPTPPIVKPPIV---YPPPYVPLPPVV 375
>TC14758 similar to UP|O24601 (O24601) Glycine-rich RNA binding protein 2,
partial (93%)
Length = 725
Score = 46.6 bits (109), Expect = 5e-06
Identities = 29/77 (37%), Positives = 36/77 (46%), Gaps = 1/77 (1%)
Frame = -1
Query: 156 PPPVS-QQPFPYPAYPPSYQAPLPPPPPVVASTPPSAPPSNVVPALPPAKTNASSHPQLK 214
PPP+ ++P P P YPPS + P PPPPP PP PP P PP +P +
Sbjct: 533 PPPLP*REPPPSPPYPPSRRPP*PPPPP-----PPYPPPPPP*PRDPP----PPPYPPSR 381
Query: 215 CPMAGTFYRCPGPGEPP 231
P + P P PP
Sbjct: 380 RPPPPPYPPPPRPPPPP 330
Score = 40.4 bits (93), Expect = 4e-04
Identities = 19/38 (50%), Positives = 21/38 (55%)
Frame = -1
Query: 156 PPPVSQQPFPYPAYPPSYQAPLPPPPPVVASTPPSAPP 193
PPP + P P P YPPS + P PP PP PP PP
Sbjct: 431 PPP*PRDP-PPPPYPPSRRPPPPPYPPPPRPPPPPPPP 321
Score = 37.0 bits (84), Expect = 0.004
Identities = 19/46 (41%), Positives = 24/46 (51%)
Frame = -1
Query: 156 PPPVSQQPFPYPAYPPSYQAPLPPPPPVVASTPPSAPPSNVVPALP 201
P P S++P P P PP P PPPP AS PS+ +P +P
Sbjct: 398 PYPPSRRPPPPPYPPPPRPPPPPPPPRAWASFTVMLRPSSSLPFIP 261
Score = 35.4 bits (80), Expect = 0.012
Identities = 18/46 (39%), Positives = 19/46 (41%)
Frame = -1
Query: 157 PPVSQQPFPYPAYPPSYQAPLPPPPPVVASTPPSAPPSNVVPALPP 202
PP P+P P P P PP PP PP PP P PP
Sbjct: 464 PPPPPPPYPPPPPP*PRDPPPPPYPPSRRPPPPPYPPPPRPPPPPP 327
Score = 29.6 bits (65), Expect = 0.65
Identities = 23/68 (33%), Positives = 24/68 (34%)
Frame = -1
Query: 164 FPYPAYPPSYQAPLPPPPPVVASTPPSAPPSNVVPALPPAKTNASSHPQLKCPMAGTFYR 223
FP P PP PP PP S P PP P PP P P + R
Sbjct: 548 FPPPPPPPLP*REPPPSPPYPPSRRPP*PPPPPPPYPPPPPP*PRDPPPPPYPPS---RR 378
Query: 224 CPGPGEPP 231
P P PP
Sbjct: 377 PPPPPYPP 354
Score = 28.9 bits (63), Expect = 1.1
Identities = 11/20 (55%), Positives = 13/20 (65%)
Frame = -1
Query: 174 QAPLPPPPPVVASTPPSAPP 193
Q P PPPPP+ PP +PP
Sbjct: 551 QFPPPPPPPLP*REPPPSPP 492
>TC14279 similar to UP|Q09085 (Q09085) Hydroxyproline-rich glycoprotein
(HRGP) (Fragment), partial (57%)
Length = 941
Score = 46.2 bits (108), Expect = 7e-06
Identities = 29/75 (38%), Positives = 32/75 (42%)
Frame = +1
Query: 156 PPPVSQQPFPYPAYPPSYQAPLPPPPPVVASTPPSAPPSNVVPALPPAKTNASSHPQLKC 215
PPP S P PY P +P PPPP AS PP +P PA P S P +K
Sbjct: 340 PPPTSSPPPPYHYVSPPPPSPSPPPPYHYASPPPPSPS----PA--PTYIYKSPPPPVKL 501
Query: 216 PMAGTFYRCPGPGEP 230
P Y P P P
Sbjct: 502 PPPPYHYTSPPPPSP 546
Score = 42.4 bits (98), Expect = 1e-04
Identities = 28/82 (34%), Positives = 38/82 (46%), Gaps = 7/82 (8%)
Frame = +1
Query: 156 PPPVSQQPFPYPAYPPS---YQAPLP----PPPPVVASTPPSAPPSNVVPALPPAKTNAS 208
PPP +P+ Y + PP Y++P P PPPP +PP PPS+ + PP S
Sbjct: 28 PPPPVPKPYYYKSPPPPPYYYKSPPPPSPSPPPPYYYQSPP--PPSH---SPPPPYYYKS 192
Query: 209 SHPQLKCPMAGTFYRCPGPGEP 230
P P +Y+ P P P
Sbjct: 193 PPPPSPSPPPPYYYKSPPPPSP 258
Score = 42.4 bits (98), Expect = 1e-04
Identities = 25/70 (35%), Positives = 30/70 (42%)
Frame = +1
Query: 161 QQPFPYPAYPPSYQAPLPPPPPVVASTPPSAPPSNVVPALPPAKTNASSHPQLKCPMAGT 220
+ P P P P Y PPPPP +PP PPS P+ PP S P P
Sbjct: 16 KSPPPPPPVPKPYYYKSPPPPPYYYKSPP--PPS---PSPPPPYYYQSPPPPSHSPPPPY 180
Query: 221 FYRCPGPGEP 230
+Y+ P P P
Sbjct: 181 YYKSPPPPSP 210
Score = 42.0 bits (97), Expect = 1e-04
Identities = 29/79 (36%), Positives = 34/79 (42%), Gaps = 4/79 (5%)
Frame = +1
Query: 156 PPPVSQQPFPYPAYPPSYQAPLP----PPPPVVASTPPSAPPSNVVPALPPAKTNASSHP 211
PPP P PY YQ+P P PPPP +PP PPS P+ PP S P
Sbjct: 100 PPPSPSPPPPY-----YYQSPPPPSHSPPPPYYYKSPP--PPS---PSPPPPYYYKSPPP 249
Query: 212 QLKCPMAGTFYRCPGPGEP 230
P +Y+ P P P
Sbjct: 250 PSPSPPPPYYYKSPPPPSP 306
Score = 42.0 bits (97), Expect = 1e-04
Identities = 31/79 (39%), Positives = 35/79 (44%), Gaps = 3/79 (3%)
Frame = +1
Query: 156 PPPVSQQPFPYPAYPPSYQAPLP-PPPPVVASTPPSAPPSNVVPALPPAKTNASSHPQLK 214
PPP S P P P Y S P P PPPP +PP PPS P+ PP S P
Sbjct: 145 PPPPSHSP-PPPYYYKSPPPPSPSPPPPYYYKSPP--PPS---PSPPPPYYYKSPPPPSP 306
Query: 215 CPMAGTFYRCPGP--GEPP 231
P +Y+ P P PP
Sbjct: 307 IPHPPYYYKSPPPPTSSPP 363
Score = 40.8 bits (94), Expect = 3e-04
Identities = 28/84 (33%), Positives = 34/84 (40%), Gaps = 12/84 (14%)
Frame = +1
Query: 156 PPPVSQQPFPYPAYPPSYQAPLP--------PPPPVVASTPP----SAPPSNVVPALPPA 203
PPP P PY P +P+P PPPP + PP S PP + P+ PP
Sbjct: 244 PPPSPSPPPPYYYKSPPPPSPIPHPPYYYKSPPPPTSSPPPPYHYVSPPPPS--PSPPPP 417
Query: 204 KTNASSHPQLKCPMAGTFYRCPGP 227
AS P P Y+ P P
Sbjct: 418 YHYASPPPPSPSPAPTYIYKSPPP 489
Score = 38.9 bits (89), Expect = 0.001
Identities = 26/85 (30%), Positives = 29/85 (33%), Gaps = 10/85 (11%)
Frame = +1
Query: 156 PPPVSQQPFPYPAYPPSYQAPLPPPPPVVASTPPSAP----------PSNVVPALPPAKT 205
PPP P PY P +P PPPP S PP +P P + PP
Sbjct: 196 PPPSPSPPPPYYYKSPPPPSPSPPPPYYYKSPPPPSPIPHPPYYYKSPPPPTSSPPPPYH 375
Query: 206 NASSHPQLKCPMAGTFYRCPGPGEP 230
S P P Y P P P
Sbjct: 376 YVSPPPPSPSPPPPYHYASPPPPSP 450
Score = 38.9 bits (89), Expect = 0.001
Identities = 32/94 (34%), Positives = 37/94 (39%), Gaps = 13/94 (13%)
Frame = +1
Query: 154 LQPPPVSQQPFP---YPAYPPSYQAPLP------PPPPVVASTPP----SAPPSNVVPAL 200
+ PPP S P P Y + PP +P P PPPPV PP S PP + PA
Sbjct: 379 VSPPPPSPSPPPPYHYASPPPPSPSPAPTYIYKSPPPPVKLPPPPYHYTSPPPPSPSPA- 555
Query: 201 PPAKTNASSHPQLKCPMAGTFYRCPGPGEPPFVK 234
P S P K P + P PP K
Sbjct: 556 -PTYIYKSPPPPTKSPPPPVYIYASPP--PPIYK 648
Score = 35.4 bits (80), Expect = 0.012
Identities = 28/79 (35%), Positives = 31/79 (38%), Gaps = 4/79 (5%)
Frame = +1
Query: 156 PPPVSQQPFPYPAYPPSYQAPLP----PPPPVVASTPPSAPPSNVVPALPPAKTNASSHP 211
PPP P P P P Y++P P PPPP +PP PPS P P S P
Sbjct: 193 PPP----PSPSPPPPYYYKSPPPPSPSPPPPYYYKSPP--PPS---PIPHPPYYYKSPPP 345
Query: 212 QLKCPMAGTFYRCPGPGEP 230
P Y P P P
Sbjct: 346 PTSSPPPPYHYVSPPPPSP 402
>TC14326 weakly similar to UP|EXLP_TOBAC (Q03211) Pistil-specific
extensin-like protein precursor (PELP), partial (11%)
Length = 630
Score = 45.8 bits (107), Expect = 9e-06
Identities = 30/84 (35%), Positives = 38/84 (44%), Gaps = 3/84 (3%)
Frame = +2
Query: 154 LQPPPVSQQPFPYPAYPPSYQAPL-PPPPPVVASTPPSAPPSNVVPALP--PAKTNASSH 210
++ PPV P PPSY P+ P PP+V S P AP +V + P P A +
Sbjct: 323 VKSPPVKAPTPPIVKSPPSYPPPVKAPSPPIVKSPPVKAPTPPIVKSPPYYPPPVKAPTP 502
Query: 211 PQLKCPMAGTFYRCPGPGEPPFVK 234
PQ+K P P PP VK
Sbjct: 503 PQVKTP----------PPHPPVVK 544
Score = 45.4 bits (106), Expect = 1e-05
Identities = 25/71 (35%), Positives = 33/71 (46%), Gaps = 9/71 (12%)
Frame = +2
Query: 157 PPVSQQPFPYPAYPPSYQAPLPP---------PPPVVASTPPSAPPSNVVPALPPAKTNA 207
PP+ + P P+YPP +AP PP P P + +PP PP P P KT
Sbjct: 353 PPIVKSP---PSYPPPVKAPSPPIVKSPPVKAPTPPIVKSPPYYPPPVKAPTPPQVKTPP 523
Query: 208 SSHPQLKCPMA 218
P +K P+A
Sbjct: 524 PHPPVVKPPVA 556
Score = 40.0 bits (92), Expect = 5e-04
Identities = 33/88 (37%), Positives = 41/88 (46%), Gaps = 5/88 (5%)
Frame = +2
Query: 152 EALQPPPVSQQPFPYPAYPPSYQAPLPP---PPPVVASTPP--SAPPSNVVPALPPAKTN 206
+A PP V P+ +PP +AP PP PPV A TPP +PPS P P+
Sbjct: 248 KAPSPPIVKSPPY----FPPPVKAPSPPIVKSPPVKAPTPPIVKSPPSYPPPVKAPSPPI 415
Query: 207 ASSHPQLKCPMAGTFYRCPGPGEPPFVK 234
S P +K P + P P PP VK
Sbjct: 416 VKS-PPVKAP-TPPIVKSP-PYYPPPVK 490
Score = 38.5 bits (88), Expect = 0.001
Identities = 30/81 (37%), Positives = 36/81 (44%), Gaps = 3/81 (3%)
Frame = +2
Query: 157 PPVSQQPFPYPAYPPSYQAPLPPP-PPVVASTPP--SAPPSNVVPALPPAKTNASSHPQL 213
PP P PP +AP PPP PPV A +PP +PP P P+ S P +
Sbjct: 164 PPTPSTPAVTVPTPPLVKAPTPPPQPPVKAPSPPIVKSPPYFPPPVKAPSPPIVKS-PPV 340
Query: 214 KCPMAGTFYRCPGPGEPPFVK 234
K P + P P PP VK
Sbjct: 341 KAP-TPPIVKSP-PSYPPPVK 397
Score = 37.4 bits (85), Expect = 0.003
Identities = 31/89 (34%), Positives = 38/89 (41%), Gaps = 11/89 (12%)
Frame = +2
Query: 157 PPVSQQPFPYPAYPPSYQAPLPP--------PPPVVASTPP--SAPPSNVVPALPPAKTN 206
PP+ + P P P P +AP PP PPPV A +PP +PP P P K+
Sbjct: 203 PPLVKAPTPPPQ--PPVKAPSPPIVKSPPYFPPPVKAPSPPIVKSPPVK-APTPPIVKSP 373
Query: 207 ASSHPQLKCPMAGTFYRCPGPG-EPPFVK 234
S P +K P P PP VK
Sbjct: 374 PSYPPPVKAPSPPIVKSPPVKAPTPPIVK 460
Score = 36.2 bits (82), Expect = 0.007
Identities = 22/71 (30%), Positives = 27/71 (37%)
Frame = +2
Query: 154 LQPPPVSQQPFPYPAYPPSYQAPLPPPPPVVASTPPSAPPSNVVPALPPAKTNASSHPQL 213
++ PPV P PP Y P+ P P TPP PP P P T +
Sbjct: 416 VKSPPVKAPTPPIVKSPPYYPPPVKAPTPPQVKTPPPHPPVVKPPVAPIPSTPIVKSMKD 595
Query: 214 KCPMAGTFYRC 224
P+ YRC
Sbjct: 596 CIPLCN--YRC 622
Score = 34.3 bits (77), Expect = 0.027
Identities = 21/53 (39%), Positives = 25/53 (46%), Gaps = 2/53 (3%)
Frame = +2
Query: 152 EALQPPPVSQQPFPYPAYPPSYQAPLPPPPPVVASTPP--SAPPSNVVPALPP 202
+A PP V P P PP ++P PPPV A TPP PP + PP
Sbjct: 395 KAPSPPIVKSPPVKAPT-PPIVKSPPYYPPPVKAPTPPQVKTPPPHPPVVKPP 550
Score = 29.6 bits (65), Expect = 0.65
Identities = 26/81 (32%), Positives = 32/81 (39%), Gaps = 3/81 (3%)
Frame = +2
Query: 157 PPVSQQPFPYPAYPPSYQAPLPPPPPVVASTPPSAPPSNVV--PALPPAKTNASSHPQLK 214
P V P P PS A P PP+V + P+ PP V P+ P K+ P +K
Sbjct: 131 PSVPAPPSPVKPPTPSTPAVTVPTPPLVKA--PTPPPQPPVKAPSPPIVKSPPYFPPPVK 304
Query: 215 CPMAGTFYRCPGPG-EPPFVK 234
P P PP VK
Sbjct: 305 APSPPIVKSPPVKAPTPPIVK 367
Score = 29.3 bits (64), Expect = 0.86
Identities = 22/58 (37%), Positives = 26/58 (43%)
Frame = +2
Query: 148 IRKKEALQPPPVSQQPFPYPAYPPSYQAPLPPPPPVVASTPPSAPPSNVVPALPPAKT 205
I K PPPV PP + P PP PPVV PP AP +P+ P K+
Sbjct: 452 IVKSPPYYPPPVKAPT------PPQVKTP-PPHPPVV--KPPVAP----IPSTPIVKS 586
>AU251550
Length = 329
Score = 45.1 bits (105), Expect = 2e-05
Identities = 33/93 (35%), Positives = 34/93 (36%), Gaps = 17/93 (18%)
Frame = +2
Query: 156 PPPVSQQPFPYPAYPPSYQ------------APLPPPPPV--VASTPPSAPPSNVVPAL- 200
PP P P P PP Y AP PPPPP PP PPS PA+
Sbjct: 74 PPGYHNPPPPGPPQPPGYYNPPPPGPPRDPYAPAPPPPPFGHHHPPPPHGPPSPPPPAMF 253
Query: 201 --PPAKTNASSHPQLKCPMAGTFYRCPGPGEPP 231
PP PQ P PGPG PP
Sbjct: 254 YQPPPPPGPPGPPQPPHP--------PGPGPPP 328
Score = 41.2 bits (95), Expect = 2e-04
Identities = 27/83 (32%), Positives = 30/83 (35%), Gaps = 8/83 (9%)
Frame = +2
Query: 157 PPVSQQPFPYPAYPPSYQAPLPPPPPV---VASTPPSAPPSNVVPALPPAKTNASSHP-- 211
PP P P PP Y P PP PP + PP PP + PP HP
Sbjct: 35 PPQPGPPGPPGPPPPGYHNPPPPGPPQPPGYYNPPPPGPPRDPYAPAPPPPPFGHHHPPP 214
Query: 212 ---QLKCPMAGTFYRCPGPGEPP 231
P FY+ P P PP
Sbjct: 215 PHGPPSPPPPAMFYQPPPPPGPP 283
>TC8260 similar to UP|Q94ES8 (Q94ES8) Nodule extensin (Fragment), partial
(79%)
Length = 1017
Score = 44.7 bits (104), Expect = 2e-05
Identities = 21/57 (36%), Positives = 29/57 (50%), Gaps = 1/57 (1%)
Frame = +3
Query: 156 PPPVSQQPFPYPA-YPPSYQAPLPPPPPVVASTPPSAPPSNVVPALPPAKTNASSHP 211
PPP +P+ YP+ PP ++ P P P PV S PP P + + PP + HP
Sbjct: 267 PPPPPHKPYKYPSPPPPPHKYPHPHPHPVYHSPPPPPPKKHYKYSSPPPPVHTYPHP 437
Score = 38.5 bits (88), Expect = 0.001
Identities = 26/90 (28%), Positives = 35/90 (38%), Gaps = 13/90 (14%)
Frame = +3
Query: 151 KEALQPPPVSQQPFPYPAYP-------------PSYQAPLPPPPPVVASTPPSAPPSNVV 197
K + PPPV P P+P Y P Y +P PPPPP ++PP V
Sbjct: 393 KYSSPPPPVHTYPHPHPVYHSPPPPVHTYPHPHPVYHSP-PPPPPHEKPYKYASPPPPPV 569
Query: 198 PALPPAKTNASSHPQLKCPMAGTFYRCPGP 227
P ++ P P +Y+ P P
Sbjct: 570 HTYPHPVYHSPPPPVHSPPPPHYYYKSPPP 659
Score = 38.5 bits (88), Expect = 0.001
Identities = 23/58 (39%), Positives = 27/58 (45%), Gaps = 11/58 (18%)
Frame = +3
Query: 156 PPPVSQQPFPYPAY-----PPSYQAPL----PPPPPVVASTPP--SAPPSNVVPALPP 202
PPPV P P+P Y PP ++ P PPPPPV P +PP V PP
Sbjct: 459 PPPVHTYPHPHPVYHSPPPPPPHEKPYKYASPPPPPVHTYPHPVYHSPPPPVHSPPPP 632
Score = 35.8 bits (81), Expect = 0.009
Identities = 28/75 (37%), Positives = 30/75 (39%), Gaps = 14/75 (18%)
Frame = +3
Query: 156 PPPVSQQPFPY----PAY---PPSYQAPL------PPPPPVVASTPPSAPPSNVVPALPP 202
PPPV P PY P Y PP Y+ P PPPPP PS PP PP
Sbjct: 162 PPPVHSPPPPYEKPHPVYHSPPPPYEKPHPVYHSPPPPPPHKPYKYPSPPP-------PP 320
Query: 203 AK-TNASSHPQLKCP 216
K + HP P
Sbjct: 321 HKYPHPHPHPVYHSP 365
Score = 35.8 bits (81), Expect = 0.009
Identities = 25/82 (30%), Positives = 31/82 (37%), Gaps = 3/82 (3%)
Frame = +3
Query: 153 ALQPPPVSQQPFPYPAYPPSYQAPLPP---PPPVVASTPPSAPPSNVVPALPPAKTNASS 209
A PP +P+ Y + PP +P PP P PV S PP + V PP
Sbjct: 111 AYSSPPPPPKPYSYHSPPPPVHSPPPPYEKPHPVYHSPPPPYEKPHPVYHSPPP------ 272
Query: 210 HPQLKCPMAGTFYRCPGPGEPP 231
P Y+ P P PP
Sbjct: 273 ------PPPHKPYKYPSPPPPP 320
Score = 32.3 bits (72), Expect = 0.10
Identities = 24/79 (30%), Positives = 32/79 (40%), Gaps = 18/79 (22%)
Frame = +3
Query: 156 PPPVSQQPFPYPAYPP---SYQAPLP----PPPPV--------VASTPPSAPPSN---VV 197
PPP ++ + Y + PP +Y P P PPPPV V +PP PP
Sbjct: 366 PPPPPKKHYKYSSPPPPVHTYPHPHPVYHSPPPPVHTYPHPHPVYHSPPPPPPHEKPYKY 545
Query: 198 PALPPAKTNASSHPQLKCP 216
+ PP + HP P
Sbjct: 546 ASPPPPPVHTYPHPVYHSP 602
Score = 31.6 bits (70), Expect = 0.17
Identities = 27/96 (28%), Positives = 34/96 (35%), Gaps = 20/96 (20%)
Frame = +3
Query: 156 PPPVSQQPFPYPAYPPSYQAPLPPPP--------------------PVVASTPPSAPPSN 195
PPP + P P+P P Y +P PPPP PV S P PP +
Sbjct: 309 PPPPHKYPHPHPH--PVYHSPPPPPPKKHYKYSSPPPPVHTYPHPHPVYHSPP---PPVH 473
Query: 196 VVPALPPAKTNASSHPQLKCPMAGTFYRCPGPGEPP 231
P P + P + P Y+ P PP
Sbjct: 474 TYPHPHPVYHSPPPPPPHEKP-----YKYASPPPPP 566
Score = 28.1 bits (61), Expect = 1.9
Identities = 13/37 (35%), Positives = 15/37 (40%), Gaps = 11/37 (29%)
Frame = +3
Query: 156 PPPVSQQPFPYPAY-----------PPSYQAPLPPPP 181
PPP +P+P Y PP Y PPPP
Sbjct: 552 PPPPPVHTYPHPVYHSPPPPVHSPPPPHYYYKSPPPP 662
>TC15655 similar to UP|Q8LE99 (Q8LE99) Transketolase-like protein, partial
(17%)
Length = 575
Score = 44.7 bits (104), Expect = 2e-05
Identities = 28/77 (36%), Positives = 36/77 (46%)
Frame = +3
Query: 155 QPPPVSQQPFPYPAYPPSYQAPLPPPPPVVASTPPSAPPSNVVPALPPAKTNASSHPQLK 214
QPPP S P PS +APLP PPP + +PPS +N P+ P+ T
Sbjct: 45 QPPPPSLSP------KPSSRAPLPSPPPRASPSPPSPASANPPPSPSPSPTLQ------- 185
Query: 215 CPMAGTFYRCPGPGEPP 231
P+ F P P +PP
Sbjct: 186 -PLTAAF---PPPSKPP 224
Score = 35.8 bits (81), Expect = 0.009
Identities = 23/57 (40%), Positives = 29/57 (50%), Gaps = 5/57 (8%)
Frame = +3
Query: 153 ALQPPPVSQQPFPYPAYPPSYQAPLPPPPPVVASTPPSAPPSNVVPAL-----PPAK 204
+L P P S+ P P P PP +A PP P A+ PPS PS + L PP+K
Sbjct: 60 SLSPKPSSRAPLPSP--PP--RASPSPPSPASANPPPSPSPSPTLQPLTAAFPPPSK 218
>TC15253 weakly similar to UP|Q9ZT16 (Q9ZT16) Arabinogalactan-protein
(ATAGP4) (AT5G10430/F12B17_220), partial (42%)
Length = 575
Score = 44.7 bits (104), Expect = 2e-05
Identities = 28/78 (35%), Positives = 32/78 (40%), Gaps = 2/78 (2%)
Frame = +3
Query: 156 PPPVSQQPFPYPAYPPSYQAPLPPPPPVVASTPPSAPPSNVVPALPP--AKTNASSHPQL 213
PPP + P P PA PP P P P A+ PSA P P P A T ++S P
Sbjct: 192 PPPPAAAPAPPPATPPPAATPAPTTTPPAATPAPSASPPAPTPTASPTGAPTPSASSPPA 371
Query: 214 KCPMAGTFYRCPGPGEPP 231
P PGP P
Sbjct: 372 PIPSGPA--SGPGPAAGP 419
Score = 39.7 bits (91), Expect = 6e-04
Identities = 31/79 (39%), Positives = 34/79 (42%), Gaps = 6/79 (7%)
Frame = +3
Query: 159 VSQQPFPYPAYPPSYQAPLPPPPPVVASTPPSA-PPSNVVPA---LPPAKTNA-SSHPQL 213
V+Q P P P+ PPPPP A PP A PP PA PPA T A S+ P
Sbjct: 141 VAQAPGAAPTQAPT---TTPPPPPAAAPAPPPATPPPAATPAPTTTPPAATPAPSASPPA 311
Query: 214 KCPMAG-TFYRCPGPGEPP 231
P A T P PP
Sbjct: 312 PTPTASPTGAPTPSASSPP 368
Score = 34.7 bits (78), Expect = 0.020
Identities = 26/80 (32%), Positives = 36/80 (44%), Gaps = 9/80 (11%)
Frame = +1
Query: 147 MIRKKEALQPPPVSQ---QPFPYPAYPPSYQAPLPPPPPVVASTPPSAPPSNVVPA---- 199
++ + L PPP Q QP P P +PP PL PP A APP +
Sbjct: 265 LLHQLPPLLPPPPLQLPLQPLPPPVHPP-LPLPLHPPQSHPAQHQAQAPPLDQAQTALIH 441
Query: 200 --LPPAKTNASSHPQLKCPM 217
LPP ++++S P L P+
Sbjct: 442 HHLPPLHSHSTS-PSLLAPL 498
Score = 33.9 bits (76), Expect = 0.035
Identities = 23/68 (33%), Positives = 26/68 (37%), Gaps = 9/68 (13%)
Frame = +3
Query: 153 ALQPPPVSQQPFPYPAYPPSYQAPLPPPPPVVASTP---------PSAPPSNVVPALPPA 203
A P P + P PA S AP P P A TP PS P S PA P
Sbjct: 243 AATPAPTTTPPAATPAPSASPPAPTPTASPTGAPTPSASSPPAPIPSGPASGPGPAAGPG 422
Query: 204 KTNASSHP 211
+A + P
Sbjct: 423 PNSADTPP 446
Score = 31.6 bits (70), Expect = 0.17
Identities = 24/72 (33%), Positives = 29/72 (39%), Gaps = 6/72 (8%)
Frame = +3
Query: 155 QPPPVSQQPFPYPAYPPSYQA-PLPPP--PPVVASTPPSAPPSNVVP---ALPPAKTNAS 208
Q P + P PP A P PPP PP A+ P+ P P A PPA T +
Sbjct: 147 QAPGAAPTQAPTTTPPPPPAAAPAPPPATPPPAATPAPTTTPPAATPAPSASPPAPTPTA 326
Query: 209 SHPQLKCPMAGT 220
S P A +
Sbjct: 327 SPTGAPTPSASS 362
Score = 26.2 bits (56), Expect = 7.2
Identities = 18/63 (28%), Positives = 24/63 (37%)
Frame = +1
Query: 155 QPPPVSQQPFPYPAYPPSYQAPLPPPPPVVASTPPSAPPSNVVPALPPAKTNASSHPQLK 214
Q P +Q P + LP PP++ PP PP + L P HP L
Sbjct: 181 QQPHHHRQQQHLPHLQQLHHQQLPLLPPLLHQLPPLLPPPPLQLPLQPLPPPV--HPPLP 354
Query: 215 CPM 217
P+
Sbjct: 355 LPL 363
>TC19964 similar to UP|Q40363 (Q40363) NuM1 protein, partial (11%)
Length = 562
Score = 44.3 bits (103), Expect = 3e-05
Identities = 22/49 (44%), Positives = 27/49 (54%)
Frame = -2
Query: 154 LQPPPVSQQPFPYPAYPPSYQAPLPPPPPVVASTPPSAPPSNVVPALPP 202
L PPP+ +P P PP PLPPP P +++ PP PP P LPP
Sbjct: 285 LPPPPLPPRPPRSPPRPPLPNRPLPPPRP-LSNRPPRPPPLLPPPLLPP 142
>TC19908 similar to GB|CAA60022.1|791150|VURNEXT26 extensin-like protein
{Vigna unguiculata;} , partial (49%)
Length = 472
Score = 43.5 bits (101), Expect = 4e-05
Identities = 25/59 (42%), Positives = 29/59 (48%), Gaps = 3/59 (5%)
Frame = +2
Query: 156 PPPVSQQPFPYPAYPPSYQAPLPPPPPVVASTPP---SAPPSNVVPALPPAKTNASSHP 211
PPP P PY P +P PPPP + S PP S PP V + PP +SSHP
Sbjct: 35 PPPSPSPPPPYVYKSPPPPSPSPPPPYIYKSPPPPSHSPPPPYVYKSPPPP---SSSHP 202
Score = 37.4 bits (85), Expect = 0.003
Identities = 25/72 (34%), Positives = 31/72 (42%)
Frame = +2
Query: 156 PPPVSQQPFPYPAYPPSYQAPLPPPPPVVASTPPSAPPSNVVPALPPAKTNASSHPQLKC 215
PPP P+ Y + PP +P PPPP V S PP + P+ PP S P
Sbjct: 5 PPP----PYVYKSPPPP--SPSPPPPYVYKSPPPPS------PSPPPPYIYKSPPPPSHS 148
Query: 216 PMAGTFYRCPGP 227
P Y+ P P
Sbjct: 149 PPPPYVYKSPPP 184
>TC17113 weakly similar to UP|Q9LFU8 (Q9LFU8) Proline-rich protein, partial
(25%)
Length = 1077
Score = 43.5 bits (101), Expect = 4e-05
Identities = 25/61 (40%), Positives = 27/61 (43%), Gaps = 6/61 (9%)
Frame = +1
Query: 155 QPPPVSQQPFPYPAYPP------SYQAPLPPPPPVVASTPPSAPPSNVVPALPPAKTNAS 208
QPPP Q P P +PP + P PPPPP PP PP P PP T S
Sbjct: 673 QPPPSKQPPSPLFPFPPIVIPGLTPPPPPPPPPPFFPPFPPLFPP----PGRPPISTKNS 840
Query: 209 S 209
S
Sbjct: 841 S 843
Score = 43.1 bits (100), Expect = 6e-05
Identities = 29/84 (34%), Positives = 36/84 (42%), Gaps = 7/84 (8%)
Frame = +1
Query: 156 PPPVSQQPFPYPAYPPSYQAPL--PPPPPVVAS---TPPSAPPSNVVPALPPAKTNASS- 209
PPP+ PF P+ PP + P P PPP++ + PPS PP P PP S
Sbjct: 526 PPPLLPDPFQPPSPPPLFPNPFQPPSPPPLIPNPFQPPPSPPPFIPNPFQPPPSKQPPSP 705
Query: 210 -HPQLKCPMAGTFYRCPGPGEPPF 232
P + G P P PPF
Sbjct: 706 LFPFPPIVIPGLTPPPPPPPPPPF 777
Score = 42.4 bits (98), Expect = 1e-04
Identities = 27/77 (35%), Positives = 32/77 (41%)
Frame = +1
Query: 155 QPPPVSQQPFPYPAYPPSYQAPLPPPPPVVASTPPSAPPSNVVPALPPAKTNASSHPQLK 214
QPPP+ PF P P+ P PPP PPS PP P PP+ +P
Sbjct: 457 QPPPLIPNPFQPPPLIPNPFQPSPPPLLPDPFQPPSPPPLFPNPFQPPSPPPLIPNPFQP 636
Query: 215 CPMAGTFYRCPGPGEPP 231
P F P P +PP
Sbjct: 637 PPSPPPF--IPNPFQPP 681
Score = 39.7 bits (91), Expect = 6e-04
Identities = 28/88 (31%), Positives = 33/88 (36%), Gaps = 10/88 (11%)
Frame = +1
Query: 156 PPPVSQQPFPYPAYPPSYQAP------LPPPP----PVVASTPPSAPPSNVVPALPPAKT 205
PPP+ PF P P +Q P PPP P S PP P P+ PP
Sbjct: 403 PPPLIPNPFQPPLVPNPFQPPPLIPNPFQPPPLIPNPFQPSPPPLLPDPFQPPSPPPLFP 582
Query: 206 NASSHPQLKCPMAGTFYRCPGPGEPPFV 233
N P + F P P PPF+
Sbjct: 583 NPFQPPSPPPLIPNPFQ--PPPSPPPFI 660
Score = 35.8 bits (81), Expect = 0.009
Identities = 23/55 (41%), Positives = 24/55 (42%), Gaps = 7/55 (12%)
Frame = +1
Query: 155 QPPPVSQQPFPYPAY-PPSYQAPLP--PPPPVV----ASTPPSAPPSNVVPALPP 202
QPPP P P PPS Q P P P PP+V PP PP P PP
Sbjct: 631 QPPPSPPPFIPNPFQPPPSKQPPSPLFPFPPIVIPGLTPPPPPPPPPPFFPPFPP 795
Score = 29.6 bits (65), Expect = 0.65
Identities = 18/41 (43%), Positives = 24/41 (57%)
Frame = +1
Query: 156 PPPVSQQPFPYPAYPPSYQAPLPPPPPVVASTPPSAPPSNV 196
PPP PF +P +PP + P P PP+ ST S+P S+V
Sbjct: 751 PPPPPPPPF-FPPFPPLF--PPPGRPPI--STKNSSP*SSV 858
>TC15072
Length = 721
Score = 43.1 bits (100), Expect = 6e-05
Identities = 29/89 (32%), Positives = 36/89 (39%), Gaps = 4/89 (4%)
Frame = +1
Query: 153 ALQPPPVSQQPFPYPAYPPSYQAPLPPPPPVVASTPPSAPPSNVV----PALPPAKTNAS 208
++ PPP P Y YPP Y A PPPPP ++ S NV P +PP ++
Sbjct: 1 SMPPPPQPTYPPSY-GYPPYYNAVPPPPPPAPVTSEQSPSFGNVPWASNPMVPPPPSDNQ 177
Query: 209 SHPQLKCPMAGTFYRCPGPGEPPFVKVGD 237
S P R P PP GD
Sbjct: 178 SQP-----------RATNPPLPPASSAGD 231
>TC9115 similar to PIR|T00425|T00425 photolyase/blue-light receptor (PHR2)
[imported] - Arabidopsis thaliana
{Arabidopsis thaliana;}, partial (55%)
Length = 1018
Score = 43.1 bits (100), Expect = 6e-05
Identities = 25/63 (39%), Positives = 33/63 (51%), Gaps = 2/63 (3%)
Frame = +1
Query: 156 PPPVSQQPFP--YPAYPPSYQAPLPPPPPVVASTPPSAPPSNVVPALPPAKTNASSHPQL 213
PPP ++P P P PP+ APL P P ST P+ P S P+ A T+AS+ +L
Sbjct: 298 PPPHPRRPNPPSNPPSPPTPSAPLTP*APTAPSTLPTPPHSAAPPSSGSATTSASATTRL 477
Query: 214 KCP 216
P
Sbjct: 478 STP 486
Score = 34.3 bits (77), Expect = 0.027
Identities = 16/35 (45%), Positives = 18/35 (50%)
Frame = +1
Query: 167 PAYPPSYQAPLPPPPPVVASTPPSAPPSNVVPALP 201
P PPS + PPPP PPS PPS P+ P
Sbjct: 262 PKPPPSPTSHSPPPPHPRRPNPPSNPPSPPTPSAP 366
Score = 26.2 bits (56), Expect = 7.2
Identities = 20/72 (27%), Positives = 28/72 (38%), Gaps = 9/72 (12%)
Frame = -2
Query: 168 AYPPSYQA---------PLPPPPPVVASTPPSAPPSNVVPALPPAKTNASSHPQLKCPMA 218
AYPP+ +A P PP +PPS+PP + P P H + C
Sbjct: 867 AYPPTEKANPQHGTKCSPKSTSPP---HSPPSSPPQSSPPPSP-------HHAKPLCEHT 718
Query: 219 GTFYRCPGPGEP 230
++ P P P
Sbjct: 717 QRQHQQPSPAPP 682
Database: LJGI
Posted date: Jul 30, 2004 11:16 AM
Number of letters in database: 14,692,800
Number of sequences in database: 28,460
Lambda K H
0.315 0.133 0.392
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 6,503,066
Number of Sequences: 28460
Number of extensions: 147467
Number of successful extensions: 8476
Number of sequences better than 10.0: 987
Number of HSP's better than 10.0 without gapping: 3158
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 5621
length of query: 302
length of database: 4,897,600
effective HSP length: 90
effective length of query: 212
effective length of database: 2,336,200
effective search space: 495274400
effective search space used: 495274400
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 16 ( 7.3 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 42 (22.0 bits)
S2: 55 (25.8 bits)
Medicago: description of AC119411.2


