

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
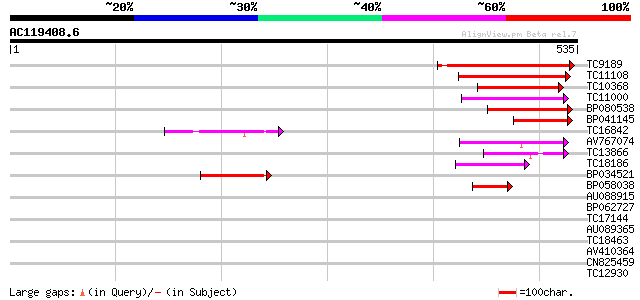
Query= AC119408.6 - phase: 0
(535 letters)
Database: LJGI
28,460 sequences; 14,692,800 total letters
Searching..................................................done
Score E
Sequences producing significant alignments: (bits) Value
TC9189 similar to UP|Q9FFN2 (Q9FFN2) Similarity to limonene cycl... 148 2e-36
TC11108 weakly similar to UP|O64805 (O64805) T1F15.13 protein (F... 124 5e-29
TC10368 111 3e-25
TC11000 homologue to UP|Q940Q8 (Q940Q8) AT5g61840/mac9_140, part... 82 2e-16
BP080538 80 6e-16
BP041145 70 9e-13
TC16842 63 1e-10
AV767074 58 3e-09
TC13866 homologue to UP|Q940Q8 (Q940Q8) AT5g61840/mac9_140, part... 56 2e-08
TC18186 similar to UP|Q8S9K6 (Q8S9K6) At1g21480/F24J8_23, partia... 54 6e-08
BP034521 53 1e-07
BP058038 40 7e-04
AU088915 37 0.006
BP062727 33 0.12
TC17144 33 0.15
AU089365 30 0.98
TC18463 similar to AAR92278 (AAR92278) At4g22580, partial (34%) 30 1.3
AV410364 29 1.7
CN825459 28 2.8
TC12930 homologue to UP|Q84L58 (Q84L58) 1-aminocyclopropane-1-ca... 27 6.3
>TC9189 similar to UP|Q9FFN2 (Q9FFN2) Similarity to limonene cyclase,
partial (30%)
Length = 544
Score = 148 bits (374), Expect = 2e-36
Identities = 67/130 (51%), Positives = 97/130 (74%)
Frame = +3
Query: 404 YKPLPLRVSKKMTYIQHMKSSKYCLCPMGFEVNSPRIVEAIYYECVPVIIADNFVLPLSE 463
Y LP K ++Y ++ SK+CLCP G+EV SPR+VEAIY CVPV+I++++V P S+
Sbjct: 3 YSSLP----KAVSYYDMLRKSKFCLCPSGYEVASPRVVEAIYTGCVPVLISEHYVPPFSD 170
Query: 464 VLDWSAFSVVVAEKDIPRLKDILLSIPMRKYVAMQNNVKMVQKHFLWNPKPIRYDLFHMI 523
VL+W +FS+ V+ KDIPRLK+ILLS+ R+Y+ MQ V +++HF + P R+D+FHM+
Sbjct: 171 VLNWKSFSLEVSVKDIPRLKEILLSVNTRQYIRMQRRVGHIRRHFEIHSPPKRFDVFHMV 350
Query: 524 LHSIWLNKLN 533
LHS+WL +LN
Sbjct: 351 LHSVWLRRLN 380
>TC11108 weakly similar to UP|O64805 (O64805) T1F15.13 protein (Fragment),
partial (10%)
Length = 517
Score = 124 bits (310), Expect = 5e-29
Identities = 49/106 (46%), Positives = 78/106 (73%)
Frame = +1
Query: 424 SKYCLCPMGFEVNSPRIVEAIYYECVPVIIADNFVLPLSEVLDWSAFSVVVAEKDIPRLK 483
SK+C+CP G +VNS RI ++I+Y C+PVI++D + LP +++++W F+VV+ E D+ +LK
Sbjct: 19 SKFCICPGGSQVNSARIADSIHYGCIPVILSDYYDLPFNDIINWRKFAVVLKESDVYQLK 198
Query: 484 DILLSIPMRKYVAMQNNVKMVQKHFLWNPKPIRYDLFHMILHSIWL 529
IL +I ++V + NN+ VQKHF WN P+ YD FHM+++ +WL
Sbjct: 199 QILKNISQAEFVTLHNNLVKVQKHFQWNSPPVGYDAFHMVMYDLWL 336
>TC10368
Length = 561
Score = 111 bits (278), Expect = 3e-25
Identities = 50/81 (61%), Positives = 65/81 (79%)
Frame = +2
Query: 442 EAIYYECVPVIIADNFVLPLSEVLDWSAFSVVVAEKDIPRLKDILLSIPMRKYVAMQNNV 501
EAI YECVPVII+DNFV P EVL+W +F+V+V EKDIP LK+ILLSIP +Y+ + V
Sbjct: 2 EAISYECVPVIISDNFVPPFFEVLNWESFAVIVLEKDIPNLKNILLSIPEPRYLRLLMRV 181
Query: 502 KMVQKHFLWNPKPIRYDLFHM 522
K V++HFLW+ P++YD+FHM
Sbjct: 182 KKVKQHFLWHKNPVQYDIFHM 244
>TC11000 homologue to UP|Q940Q8 (Q940Q8) AT5g61840/mac9_140, partial (33%)
Length = 637
Score = 82.4 bits (202), Expect = 2e-16
Identities = 42/103 (40%), Positives = 62/103 (59%), Gaps = 2/103 (1%)
Frame = +3
Query: 427 CLCPMGFEVNSPRIVEAIYYECVPVIIADNFVLPLSEVLDWSAFSVVVAEKDIPRLKDIL 486
CLCP+G+ SPR+VEA+ + C+PVIIAD+ VLP ++ + W V V EKD+P+L IL
Sbjct: 3 CLCPLGWAPWSPRLVEAVVFGCIPVIIADDIVLPFADAIPWEEIGVFVDEKDVPQLDTIL 182
Query: 487 LSIPMRKYVAMQNNV-KMVQKHFLWNPKPIR-YDLFHMILHSI 527
SIP + Q + K + P+P + D FH +L+ +
Sbjct: 183 TSIPPEVILRKQRLLANPSMKQAMLFPQPAQPGDTFHQVLNGL 311
>BP080538
Length = 387
Score = 80.5 bits (197), Expect = 6e-16
Identities = 35/80 (43%), Positives = 55/80 (68%)
Frame = -1
Query: 452 IIADNFVLPLSEVLDWSAFSVVVAEKDIPRLKDILLSIPMRKYVAMQNNVKMVQKHFLWN 511
IIA+ + LP ++VL+W +FSV+V DIP LK IL I +Y+ +Q+NV V+KHF W+
Sbjct: 387 IIANYYDLPFADVLNWKSFSVIVTTLDIPLLKKILKGISSDEYLMLQSNVLKVRKHFQWH 208
Query: 512 PKPIRYDLFHMILHSIWLNK 531
P +D F+M+++ +WL +
Sbjct: 207 SPPHDFDAFYMVMYELWLRR 148
>BP041145
Length = 507
Score = 70.1 bits (170), Expect = 9e-13
Identities = 28/56 (50%), Positives = 41/56 (73%)
Frame = -3
Query: 476 EKDIPRLKDILLSIPMRKYVAMQNNVKMVQKHFLWNPKPIRYDLFHMILHSIWLNK 531
E D+ +LKDIL SIP +++VA+ N+ VQKHF WN P+R D FHM+++ +WL +
Sbjct: 502 ETDVYKLKDILRSIPEKQFVALNQNLVKVQKHFNWNTPPVRLDAFHMVMYELWLRR 335
>TC16842
Length = 563
Score = 62.8 bits (151), Expect = 1e-10
Identities = 40/114 (35%), Positives = 62/114 (54%), Gaps = 2/114 (1%)
Frame = +3
Query: 147 PSGKQTDIRLITPTEALVYARKEIDHVTSVNEDPDLYAPLFRNVSVFKRSYELMETVLKV 206
PS + L+ + + + R + D V D D + ++ + VFK +Y M KV
Sbjct: 240 PSPQVPIANLVGNPQVVEHNRDDDDAV-----DDDEFGDVYHSPRVFKLNYAEMVKNFKV 404
Query: 207 YIYRDGSRPIFHNP--SLKGIYASEGWFMKLMQENKQFVTKDPERAHLFYLPYS 258
YIY DG F+ L G YASEG+F + ++E++ F T DP++AHLF++P S
Sbjct: 405 YIYPDGDPKTFYQTPRKLTGKYASEGYFFQNIRESR-FRTLDPDQAHLFFIPIS 563
>AV767074
Length = 481
Score = 58.2 bits (139), Expect = 3e-09
Identities = 32/106 (30%), Positives = 55/106 (51%), Gaps = 3/106 (2%)
Frame = -1
Query: 425 KYCLCPMGFEVNSPRIVEAIYYECVPVIIADNFVLPLSEVLDWSAFSVVVAEKDIPR--- 481
K+CL P G +S R+ +AI CVPVI++D LP + +D+S FS+ + K+ +
Sbjct: 481 KFCLHPAGDTPSSCRLFDAIVSHCVPVIVSDQIELPFEDEIDYSQFSLFFSFKEALQPGY 302
Query: 482 LKDILLSIPMRKYVAMQNNVKMVQKHFLWNPKPIRYDLFHMILHSI 527
+ D L P K+ M +K + H+ + P + D +M+ +
Sbjct: 301 MIDQLRKFPKDKWSEMWRQLKNISHHYEFQYPPKKEDAVNMLWRQV 164
>TC13866 homologue to UP|Q940Q8 (Q940Q8) AT5g61840/mac9_140, partial (28%)
Length = 542
Score = 55.8 bits (133), Expect = 2e-08
Identities = 32/85 (37%), Positives = 48/85 (55%), Gaps = 5/85 (5%)
Frame = +2
Query: 448 CVPVIIADNFVLPLSEVLDWSAFSVVVAEKDIPRLKDILLSIP----MRKYVAMQNNVKM 503
C+PVIIAD+ VLP ++ + W V V E+D+P+L IL SIP +RK + N
Sbjct: 5 CIPVIIADDIVLPFADAIPWEDIGVFVDEEDVPKLDTILTSIPPEIILRKQRLLAN---P 175
Query: 504 VQKHFLWNPKPIR-YDLFHMILHSI 527
K + P+P + D FH +L+ +
Sbjct: 176 SMKQAMLFPQPAQPGDAFHQVLNGL 250
>TC18186 similar to UP|Q8S9K6 (Q8S9K6) At1g21480/F24J8_23, partial (32%)
Length = 606
Score = 53.9 bits (128), Expect = 6e-08
Identities = 29/71 (40%), Positives = 40/71 (55%), Gaps = 1/71 (1%)
Frame = +3
Query: 421 MKSSKYCLCPMGFEVNSPRIVEAIYYECVPVIIADNFVLPLSEVLDWSAFSVVVAEKDI- 479
+++SK+CL P G + R E+ + ECVPVII+D LP V+D+S S+ I
Sbjct: 3 LRNSKFCLAPRGESSWTLRFYESFFVECVPVIISDQIELPFQNVIDYSQISIKWPSNRIG 182
Query: 480 PRLKDILLSIP 490
P L L SIP
Sbjct: 183 PELLQYLESIP 215
>BP034521
Length = 507
Score = 53.1 bits (126), Expect = 1e-07
Identities = 29/69 (42%), Positives = 43/69 (62%), Gaps = 2/69 (2%)
Frame = +1
Query: 181 DLYAPLFRNVSVFKRSYELMETVLKVYIYRDGS-RPIFHNP-SLKGIYASEGWFMKLMQE 238
D + +F + F+R Y+ ME KV++Y DG FH+P L G YASEG+F K ++E
Sbjct: 301 DSSSDVFHSAEAFRRDYQKMEEEFKVFVYPDGDPDTYFHSPRKLTGKYASEGYFFKNIRE 480
Query: 239 NKQFVTKDP 247
++ F+T DP
Sbjct: 481 SR-FLTLDP 504
>BP058038
Length = 552
Score = 40.4 bits (93), Expect = 7e-04
Identities = 15/38 (39%), Positives = 26/38 (67%)
Frame = +3
Query: 437 SPRIVEAIYYECVPVIIADNFVLPLSEVLDWSAFSVVV 474
+PR+VEA+ Y +P+I+A++ VLP ++ + W V V
Sbjct: 279 NPRLVEAVVYGRIPIIVAEDIVLPFADAIPWVEIGVFV 392
>AU088915
Length = 116
Score = 37.4 bits (85), Expect = 0.006
Identities = 17/35 (48%), Positives = 23/35 (65%)
Frame = +1
Query: 331 CNADLSERIFIEGRDVSLPETTIRAPRRPLRYLGG 365
CNAD+ + F+ G+DVSLPET + P R +GG
Sbjct: 4 CNADVKQG-FVFGKDVSLPETLVHTASHPARDIGG 105
>BP062727
Length = 478
Score = 33.1 bits (74), Expect = 0.12
Identities = 10/27 (37%), Positives = 18/27 (66%)
Frame = +2
Query: 448 CVPVIIADNFVLPLSEVLDWSAFSVVV 474
C+P+++AD+ VLP ++ + W V V
Sbjct: 308 CIPIVVADDIVLPFADAIPWEEICVFV 388
>TC17144
Length = 602
Score = 32.7 bits (73), Expect = 0.15
Identities = 16/67 (23%), Positives = 33/67 (48%), Gaps = 3/67 (4%)
Frame = +3
Query: 448 CVPVIIADNFVLPLSEVLDWSAFSVVVAEKDIPR---LKDILLSIPMRKYVAMQNNVKMV 504
C+P I++D LP +LD+ ++ V+ D + L L + + +Q N+
Sbjct: 39 CIPAIVSDELELPFEGILDYRKIALFVSSIDALKPGWLLKFLKDVRPAQIKELQPNLAQY 218
Query: 505 QKHFLWN 511
+HF+++
Sbjct: 219 SRHFVFS 239
>AU089365
Length = 583
Score = 30.0 bits (66), Expect = 0.98
Identities = 11/26 (42%), Positives = 15/26 (57%)
Frame = +3
Query: 404 YKPLPLRVSKKMTYIQHMKSSKYCLC 429
Y+PLP R K+ + H+ S CLC
Sbjct: 393 YQPLPTRSLLKLVLLHHLTGSSLCLC 470
>TC18463 similar to AAR92278 (AAR92278) At4g22580, partial (34%)
Length = 555
Score = 29.6 bits (65), Expect = 1.3
Identities = 36/120 (30%), Positives = 49/120 (40%), Gaps = 10/120 (8%)
Frame = +2
Query: 197 YELMETVLKVYIYRDGSRPIFHNPSLKGIYASEGWFMKLMQENKQF----VTKDPERAHL 252
Y L+E + Y+ G HN S Y ++ ++L+ + +T DP A+
Sbjct: 173 YPLLEDLCP-YLANHGLGQKTHNRS-HSWYRTDPSMLELIFHRRMLEYPCLTADPTAANA 346
Query: 253 FYLPYSARQMEVT-LYVP-----GSHDLKPLSIFLRDYVNKIAAKYPFWNRTHGSDHFLV 306
YLPY A + LY P H L L FLR N W R G DHFL+
Sbjct: 347 IYLPYYASFDSLRYLYGPEYNSSADHGLH-LFRFLRHDDNP-----EIWERHMGHDHFLI 508
>AV410364
Length = 370
Score = 29.3 bits (64), Expect = 1.7
Identities = 9/24 (37%), Positives = 16/24 (66%)
Frame = +2
Query: 280 FLRDYVNKIAAKYPFWNRTHGSDH 303
+ ++ + I +YP+WNR+ G DH
Sbjct: 104 YYKNAYHHIVEQYPYWNRSSGRDH 175
>CN825459
Length = 696
Score = 28.5 bits (62), Expect = 2.8
Identities = 30/118 (25%), Positives = 41/118 (34%), Gaps = 4/118 (3%)
Frame = +1
Query: 87 LQNVHLTLPNSAVSPKLSNNFVQSVSVEADSINTEARRKKDRNLANTSTTVTTSFPRGRV 146
LQN T P + S +A EARR++ N + +V +F V
Sbjct: 10 LQNEFFTAKPLPCDPSTLPKYPPSKEFDAKLREEEARRRRASNKGHGQESVGRNFRESNV 189
Query: 147 PSGKQTDIRLITPTEALVYARKEIDHVTSVNEDPDLYAPLF----RNVSVFKRSYELM 200
+ L E K I + ED D PL R +SVF S + M
Sbjct: 190 VPASDANAELQASMEKRQEQSKCISEKFNPEEDGDYGFPLEPAKPRALSVFSHSGQSM 363
>TC12930 homologue to UP|Q84L58 (Q84L58) 1-aminocyclopropane-1-carboxylic
acid oxidase, partial (36%)
Length = 422
Score = 27.3 bits (59), Expect = 6.3
Identities = 20/82 (24%), Positives = 36/82 (43%), Gaps = 9/82 (10%)
Frame = +1
Query: 260 RQMEVTLYVPGSHDLKPLSIFLRDYVNKIAAKYPFWNRTHGSDHFLV------ACHEWGP 313
R+++ Y+ SH P I Y+N +A +++ +G + C EWG
Sbjct: 1 RKLKQQCYISLSHTCSPSLITTSLYLNPMAIPVIDFSKLNGEERAKTMAQIHNGCEEWGF 180
Query: 314 YTVTEH---EELARNTLKALCN 332
+ + H EEL +K +C+
Sbjct: 181 FQLINHGIPEELLER-VKKVCS 243
Database: LJGI
Posted date: Jul 30, 2004 11:16 AM
Number of letters in database: 14,692,800
Number of sequences in database: 28,460
Lambda K H
0.323 0.137 0.418
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 9,616,935
Number of Sequences: 28460
Number of extensions: 133567
Number of successful extensions: 744
Number of sequences better than 10.0: 43
Number of HSP's better than 10.0 without gapping: 739
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 741
length of query: 535
length of database: 4,897,600
effective HSP length: 95
effective length of query: 440
effective length of database: 2,193,900
effective search space: 965316000
effective search space used: 965316000
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 16 ( 7.5 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (22.0 bits)
S2: 57 (26.6 bits)
Medicago: description of AC119408.6


