

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
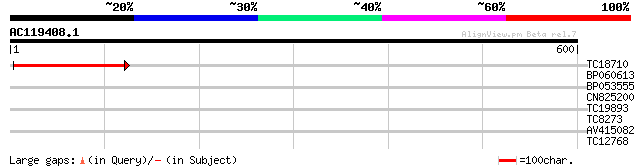
Query= AC119408.1 + phase: 0 /pseudo
(600 letters)
Database: LJGI
28,460 sequences; 14,692,800 total letters
Searching..................................................done
Score E
Sequences producing significant alignments: (bits) Value
TC18710 103 8e-23
BP060613 30 1.1
BP053555 30 1.1
CN825200 23 2.8
TC19893 similar to UP|Q7XA86 (Q7XA86) At3g51390, partial (12%) 28 4.3
TC8273 UP|BAD11134 (BAD11134) Glutamate-rich protein, complete 28 5.6
AV415082 27 9.5
TC12768 similar to UP|Q9FH10 (Q9FH10) Serine/threonine protein k... 27 9.5
>TC18710
Length = 843
Score = 103 bits (257), Expect = 8e-23
Identities = 53/123 (43%), Positives = 75/123 (60%), Gaps = 1/123 (0%)
Frame = +2
Query: 5 RIKYILPNITNDFQSTFVPGRLITDNSLIVFETFHYIRKTRTNNNGYVGIKLDIAKAYDS 64
R+K +P++ + QS F+ GR I DN LIV E FH I + + IKLD+ KAYD
Sbjct: 419 RLKDAMPDLISPMQSGFIQGRQIQDNLLIVQEAFHAINRPGALGRNHSIIKLDMNKAYDR 598
Query: 65 LKWNFIEDTLTAMGFPINLINTIMLCIRCVTFSILINGQPTEPFSPQRGIRQGDPF-PLI 123
++W F+E +L A GF N + IM+ + V+++ ING PQRG+RQGDPF P +
Sbjct: 599 VEWKFLESSLLAFGFSTNWVKMIMILVSGVSYNYKINGVVGPKLLPQRGLRQGDPFSPYL 778
Query: 124 FLY 126
FL+
Sbjct: 779 FLF 787
>BP060613
Length = 378
Score = 30.0 bits (66), Expect = 1.1
Identities = 32/106 (30%), Positives = 46/106 (43%)
Frame = +2
Query: 306 FGGVALIVKRRFTGPKEKIFLNPDIRVEWASKFLEILIWPCLPNKSRDFRLIKNPLSPEF 365
FGGV L FTG LN ++ SK L+ I P P++ +F I+ +
Sbjct: 44 FGGV-LRGHEVFTGGNGTF*LN*KMKEGLVSKTLKYRIKPS*PSRLGEFCTIQMHFGFKS 220
Query: 366 SKLNISLTKTFSRLI*VIAQVMLGGAYSILFGLFRKGVAGK*EMGK 411
K SLT F R + + G AYS++ +G E+GK
Sbjct: 221 *KPYTSLTMIFYRPLNELVLPGFGLAYSMVGSFSSSKASGNWEVGK 358
>BP053555
Length = 516
Score = 30.0 bits (66), Expect = 1.1
Identities = 18/61 (29%), Positives = 35/61 (56%)
Frame = +3
Query: 475 LLIWPMMMRLCGILIKMANIRCTVVIKLYRTEKLLLLPSLLIATLRLIYGKNCGPYTLFP 534
++ W +++L I++ N+R LY+ + L+++ +LIAT L+ N P+ LFP
Sbjct: 216 MIFWDQLLKLN---IQLLNLRRYSNTLLYQIK*LVVMLLILIATFSLV-KINLAPFILFP 383
Query: 535 Y 535
+
Sbjct: 384 F 386
>CN825200
Length = 620
Score = 23.5 bits (49), Expect(2) = 2.8
Identities = 15/43 (34%), Positives = 21/43 (47%), Gaps = 4/43 (9%)
Frame = -1
Query: 496 CTVVIKLYRTEK----LLLLPSLLIATLRLIYGKNCGPYTLFP 534
C ++I L T LLL+ LL+ + L+ CG TL P
Sbjct: 209 CQILINLCLTTSFIPLLLLVFLLLVVVVLLLCRSGCGTRTLRP 81
Score = 23.5 bits (49), Expect(2) = 2.8
Identities = 11/32 (34%), Positives = 17/32 (52%)
Frame = -1
Query: 466 TKIK*NVSLLLIWPMMMRLCGILIKMANIRCT 497
TKI S L +W + M +L +++I CT
Sbjct: 425 TKILQKGSCLRLWHLSMTSLSVLCSISHITCT 330
>TC19893 similar to UP|Q7XA86 (Q7XA86) At3g51390, partial (12%)
Length = 513
Score = 28.1 bits (61), Expect = 4.3
Identities = 20/74 (27%), Positives = 35/74 (47%), Gaps = 2/74 (2%)
Frame = -1
Query: 292 LKACVRRLRKLFVHFGGVALIVKRRFTGPKEKIFLNPDIRVEWASKFLEILIW--PCLPN 349
++ACVR L++ GG L + T E ++++ + EW F W P + N
Sbjct: 219 IRACVRFGLFLYLLRGG*WLHTEEASTSTAEVVYISTNSFTEWMHSF-----WN*P*IRN 55
Query: 350 KSRDFRLIKNPLSP 363
S + LI++ + P
Sbjct: 54 PSSPYELIRSLICP 13
>TC8273 UP|BAD11134 (BAD11134) Glutamate-rich protein, complete
Length = 923
Score = 27.7 bits (60), Expect = 5.6
Identities = 23/59 (38%), Positives = 35/59 (58%), Gaps = 6/59 (10%)
Frame = -2
Query: 150 LPLMPQLYLIYFMLM--IVSYFV-GPNPRRLVLL*TF---FRFIKRLLDRKLIWINQRW 202
L L+P L L +F+ + +V + V G PRRLV+L F F+F+ ++L+ L QRW
Sbjct: 253 LLLLPLLVLRWFLPLGVLVQWLVLGQEPRRLVVLWLFPGMFQFVPQVLE--LAVQPQRW 83
>AV415082
Length = 418
Score = 26.9 bits (58), Expect = 9.5
Identities = 14/28 (50%), Positives = 19/28 (67%), Gaps = 2/28 (7%)
Frame = -3
Query: 335 ASKFLEILIWPCLPNKS--RDFRLIKNP 360
+SKFL IL+ PC P S + RL+K+P
Sbjct: 233 SSKFLPILLIPCPPMNS*KKTTRLLKHP 150
>TC12768 similar to UP|Q9FH10 (Q9FH10) Serine/threonine protein kinase-like,
partial (18%)
Length = 597
Score = 26.9 bits (58), Expect = 9.5
Identities = 8/12 (66%), Positives = 11/12 (91%)
Frame = -1
Query: 343 IWPCLPNKSRDF 354
+WPCLP+KS+ F
Sbjct: 579 LWPCLPSKSKSF 544
Database: LJGI
Posted date: Jul 30, 2004 11:16 AM
Number of letters in database: 14,692,800
Number of sequences in database: 28,460
Lambda K H
0.344 0.154 0.513
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 11,119,902
Number of Sequences: 28460
Number of extensions: 165081
Number of successful extensions: 1604
Number of sequences better than 10.0: 16
Number of HSP's better than 10.0 without gapping: 1589
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 1603
length of query: 600
length of database: 4,897,600
effective HSP length: 95
effective length of query: 505
effective length of database: 2,193,900
effective search space: 1107919500
effective search space used: 1107919500
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 15 ( 7.5 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 38 (21.6 bits)
S2: 58 (26.9 bits)
Medicago: description of AC119408.1


