

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
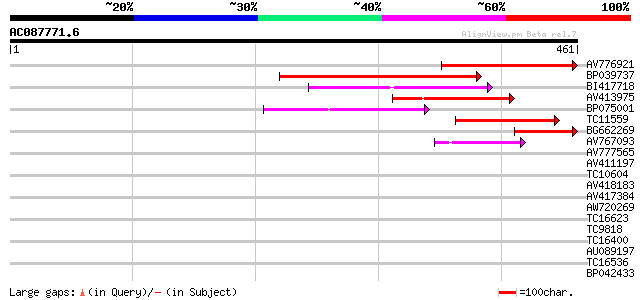
Query= AC087771.6 + phase: 0 /pseudo/partial
(461 letters)
Database: LJGI
28,460 sequences; 14,692,800 total letters
Searching..................................................done
Score E
Sequences producing significant alignments: (bits) Value
AV776921 195 1e-50
BP039737 149 1e-36
BI417718 91 4e-19
AV413975 80 5e-16
BP075001 80 9e-16
TC11559 similar to UP|Q9SXT3 (Q9SXT3) Multidrug resistance prote... 79 1e-15
BG662269 51 4e-07
AV767093 45 2e-05
AV777565 39 0.001
AV411197 39 0.002
TC10604 similar to AAQ89671 (AAQ89671) At4g33460, partial (36%) 39 0.002
AV418183 37 0.005
AV417384 36 0.015
AW720269 35 0.020
TC16623 weakly similar to UP|TAP2_HUMAN (Q03519) Antigen peptide... 32 0.17
TC9818 similar to UP|Q7XA50 (Q7XA50) Sorbitol-like transporter, ... 32 0.17
TC16400 similar to UP|O48783 (O48783) At2g26680 protein, partial... 31 0.49
AU089197 30 0.64
TC16536 similar to UP|O22477 (O22477) Betaine aldehyde dehydroge... 30 1.1
BP042433 29 1.4
>AV776921
Length = 435
Score = 195 bits (496), Expect = 1e-50
Identities = 98/110 (89%), Positives = 103/110 (93%)
Frame = +2
Query: 352 DVCFSYPSRPDVGIFTNLNLDIPAGKIVALVGGSGSGKSTVVSLIERFYEPISGQILLDK 411
DVCFSYPSRPDV I L LDIP+GKIVALVGGSGSGKSTV+SLIERFYEP+SG ILLD
Sbjct: 2 DVCFSYPSRPDVEILNKLCLDIPSGKIVALVGGSGSGKSTVISLIERFYEPLSGDILLDG 181
Query: 412 NDIRELDLKWLRQQIGLVNQEPALFATSIKENILYGKDDATLEELKRAVK 461
NDIR+LDLKWLRQQIGLVNQEPALFATSIKENILYGKD+ATLEELKRAVK
Sbjct: 182 NDIRDLDLKWLRQQIGLVNQEPALFATSIKENILYGKDNATLEELKRAVK 331
>BP039737
Length = 543
Score = 149 bits (375), Expect = 1e-36
Identities = 70/164 (42%), Positives = 107/164 (64%)
Frame = +3
Query: 220 VQAFAGEERAVRSYKAALMKTYVNGRKAGLAKGLGLGSMHCVLFLSWALLVWYTSVVVHK 279
V ++ GE +A+ SY A+ T G KAG+AKGLGLG + + +SWAL+ WY V +
Sbjct: 51 VYSYVGESKALNSYSDAIQNTLKLGYKAGMAKGLGLGCTYGIACMSWALVFWYAGVFIRN 230
Query: 280 NIANGGESFTTMLNVVISGLSLGQAAPDISAFIRAKAAAYPIFEMIERDTVSKKSSKTGR 339
+GG++FT + + ++ G+SLGQ+ ++ AF + KAA Y + E+I++ + G+
Sbjct: 231 GQTDGGKAFTAIFSAIVGGMSLGQSFSNLGAFSKGKAAGYKLMEIIKQKPTIIEDLSDGK 410
Query: 340 KLSKLDGHIQFNDVCFSYPSRPDVGIFTNLNLDIPAGKIVALVG 383
L +++G+I+F DV FSYPSRPDV IF N ++ PAGK VA+VG
Sbjct: 411 CLDEVNGNIEFKDVTFSYPSRPDVIIFRNFSIFFPAGKTVAVVG 542
>BI417718
Length = 515
Score = 90.9 bits (224), Expect = 4e-19
Identities = 55/149 (36%), Positives = 81/149 (53%)
Frame = +2
Query: 244 GRKAGLAKGLGLGSMHCVLFLSWALLVWYTSVVVHKNIANGGESFTTMLNVVISGLSLGQ 303
G + GL G+G G +LF +A + V +A+ + F + + ++ + + +
Sbjct: 77 GIQRGLISGIGFGVSFFLLFSVYATTFHVGARFVGAGMASFSDVFQVLFALTMAAIGISR 256
Query: 304 AAPDISAFIRAKAAAYPIFEMIERDTVSKKSSKTGRKLSKLDGHIQFNDVCFSYPSRPDV 363
AP+ S +AK IFE+I+R + ++G L G I+F V F YPSRPD+
Sbjct: 257 RAPNSS---KAKIVTASIFEIIDRKSKIDPCDESGSTLDSTKGKIEFCHVSFKYPSRPDI 427
Query: 364 GIFTNLNLDIPAGKIVALVGGSGSGKSTV 392
IF +L+L I AG VALVG SGSGKSTV
Sbjct: 428 QIFPDLSLTIHAGTTVALVGESGSGKSTV 514
>AV413975
Length = 343
Score = 80.5 bits (197), Expect = 5e-16
Identities = 41/99 (41%), Positives = 63/99 (63%)
Frame = +2
Query: 312 IRAKAAAYPIFEMIERDTVSKKSSKTGRKLSKLDGHIQFNDVCFSYPSRPDVGIFTNLNL 371
++A A+ +F++++R + KS T L DG ++ +DV FSYPSRP + + +
Sbjct: 44 MKAAGASRRVFQIMDRVSSMSKSG-TKCPLGDQDGEVELDDVWFSYPSRPSHPVLKGITM 220
Query: 372 DIPAGKIVALVGGSGSGKSTVVSLIERFYEPISGQILLD 410
+ G VALVG SG GK+T+ +LIERFY+P G+ILL+
Sbjct: 221 KLHPGSKVALVGPSGGGKTTIANLIERFYDPTKGKILLN 337
>BP075001
Length = 403
Score = 79.7 bits (195), Expect = 9e-16
Identities = 44/135 (32%), Positives = 73/135 (53%)
Frame = +2
Query: 207 GEIAEEVIGNVRTVQAFAGEERAVRSYKAALMKTYVNGRKAGLAKGLGLGSMHCVLFLSW 266
G +AE+ I ++RTV + E++ + + +AL K+ G K G KG+ +GS+ +L+ +W
Sbjct: 2 GSVAEQCISSIRTVYPYVSEKQTLNRFSSALEKSMELGIKQGQTKGVVIGSIG-LLYATW 178
Query: 267 ALLVWYTSVVVHKNIANGGESFTTMLNVVISGLSLGQAAPDISAFIRAKAAAYPIFEMIE 326
A W SV+V +GG+ F + ++ G+SL A P ++ I A A I EMI+
Sbjct: 179 AFQSWAGSVLVRTKGESGGQVFVAQICIIWGGISLMSALPKLAFIIEASVAGTRIIEMID 358
Query: 327 RDTVSKKSSKTGRKL 341
R + + GR L
Sbjct: 359 RKPSINANGEKGRLL 403
>TC11559 similar to UP|Q9SXT3 (Q9SXT3) Multidrug resistance protein
(Fragment), partial (41%)
Length = 460
Score = 79.3 bits (194), Expect = 1e-15
Identities = 38/85 (44%), Positives = 57/85 (66%)
Frame = +3
Query: 363 VGIFTNLNLDIPAGKIVALVGGSGSGKSTVVSLIERFYEPISGQILLDKNDIRELDLKWL 422
V I ++LDIP G IV ++G SGSGKST++ + R +EP S + LD DI LD+ L
Sbjct: 129 VPILKGIHLDIPKGVIVGVIGPSGSGKSTLLRALNRLWEPPSASVFLDARDICHLDVLSL 308
Query: 423 RQQIGLVNQEPALFATSIKENILYG 447
R+++G++ Q PALF ++ +N+ YG
Sbjct: 309 RRKVGMLFQLPALFEGTVADNVRYG 383
>BG662269
Length = 283
Score = 51.2 bits (121), Expect = 4e-07
Identities = 25/52 (48%), Positives = 38/52 (73%), Gaps = 1/52 (1%)
Frame = +2
Query: 411 KNDIRELDLKWLRQQIGLVNQEPALFATSIKENILYGK-DDATLEELKRAVK 461
+++I+ L +KWLRQQ+G+V+QEP LF +I+ NI YGK +AT E+ A +
Sbjct: 11 RDEIQTLQVKWLRQQMGMVSQEPVLFNETIRANIAYGKGGNATEAEIIAAAE 166
>AV767093
Length = 603
Score = 45.4 bits (106), Expect = 2e-05
Identities = 25/75 (33%), Positives = 45/75 (59%), Gaps = 1/75 (1%)
Frame = +1
Query: 346 GHIQFNDVCFSYPSRPDVG-IFTNLNLDIPAGKIVALVGGSGSGKSTVVSLIERFYEPIS 404
G I+ N + Y RP+ + ++L + G+ + +VG +GSGKST++ ++ R EP +
Sbjct: 379 GSIELNSLQVRY--RPNTPLVLKGISLTVQGGEKIGVVGRTGSGKSTLIQVLFRLIEPSA 552
Query: 405 GQILLDKNDIRELDL 419
G+I++D +I L L
Sbjct: 553 GKIIIDGINICTLGL 597
>AV777565
Length = 308
Score = 39.3 bits (90), Expect = 0.001
Identities = 17/29 (58%), Positives = 21/29 (71%)
Frame = -2
Query: 8 ERKKEHKVSMLKLFSFADSYDYVLMFIGS 36
E+KKE + +LFSFAD YDY+LM GS
Sbjct: 88 EKKKEQSLPFYQLFSFADKYDYMLMISGS 2
>AV411197
Length = 391
Score = 38.9 bits (89), Expect = 0.002
Identities = 15/34 (44%), Positives = 26/34 (76%)
Frame = +3
Query: 428 LVNQEPALFATSIKENILYGKDDATLEELKRAVK 461
L+ QEP +F+T+I+ENI+Y + +A+ E+K A +
Sbjct: 54 LIQQEPIIFSTTIRENIIYARHNASEAEMKEAAR 155
>TC10604 similar to AAQ89671 (AAQ89671) At4g33460, partial (36%)
Length = 498
Score = 38.5 bits (88), Expect = 0.002
Identities = 19/59 (32%), Positives = 36/59 (60%), Gaps = 2/59 (3%)
Frame = +2
Query: 355 FSYPSRP--DVGIFTNLNLDIPAGKIVALVGGSGSGKSTVVSLIERFYEPISGQILLDK 411
FS+ +R DV + + +L IP+G+ L+G +G GKST++ ++ P SG + +++
Sbjct: 164 FSFTARQTNDVRVLRDCSLRIPSGQFWMLLGPNGCGKSTLLKILAGLLAPTSGTVYVNE 340
>AV418183
Length = 298
Score = 37.4 bits (85), Expect = 0.005
Identities = 17/36 (47%), Positives = 25/36 (69%)
Frame = +3
Query: 414 IRELDLKWLRQQIGLVNQEPALFATSIKENILYGKD 449
+ E+ + L ++I +V+QEP LF SI+ENI YG D
Sbjct: 12 LAEISHRHLHRKISIVSQEPTLFNCSIEENIAYGFD 119
>AV417384
Length = 380
Score = 35.8 bits (81), Expect = 0.015
Identities = 16/44 (36%), Positives = 27/44 (61%)
Frame = +2
Query: 365 IFTNLNLDIPAGKIVALVGGSGSGKSTVVSLIERFYEPISGQIL 408
I ++N+ + G + L G +GSGKST + ++ F P +G+IL
Sbjct: 245 ILRHVNVSLHDGGALVLTGANGSGKSTFLRMLAGFSRPSAGEIL 376
>AW720269
Length = 524
Score = 35.4 bits (80), Expect = 0.020
Identities = 29/90 (32%), Positives = 48/90 (53%), Gaps = 6/90 (6%)
Frame = +3
Query: 363 VGIFTNLNLDIPAGKIVALVGGSGSGKSTVVSLI-----ERFYEPISGQILLDKNDIREL 417
V I +++ + +IVA+VG SG+GKST++ +I +R ++P + I ND
Sbjct: 48 VNILKSVSFVARSSEIVAVVGPSGTGKSTLLRIIAGRVKDRDFDPKTISI----NDHPMT 215
Query: 418 DLKWLRQQIGLVNQEPALF-ATSIKENILY 446
LR+ G V QE L ++KE +L+
Sbjct: 216 SPAQLRKICGFVAQEDNLLPLLTVKETLLF 305
>TC16623 weakly similar to UP|TAP2_HUMAN (Q03519) Antigen peptide
transporter 2 (APT2) (Peptide transporter TAP2) (Peptide
transporter PSF2) (Peptide supply factor 2) (PSF-2)
(Peptide transporter involved in antigen processing 2),
partial (4%)
Length = 530
Score = 32.3 bits (72), Expect = 0.17
Identities = 28/109 (25%), Positives = 51/109 (46%), Gaps = 2/109 (1%)
Frame = +1
Query: 312 IRAKAAAYPIFEMIERDTVSKKSSKTGRKLSKLDGHIQFNDVCFSYPSRPDVGIFTNLNL 371
+R ++ P F + R V+ ++ T S L +Q ND+ + LNL
Sbjct: 67 LRLRSPPKPTFVTLRRTCVAATAA-TDTSQSPL---LQVNDLKAKIVGEHK-DLLKGLNL 231
Query: 372 DIPAGKIVALVGGSGSGKSTVVSLI--ERFYEPISGQILLDKNDIRELD 418
I G++ AL+G +GSGKST ++ YE G+ + ++ +++
Sbjct: 232 TINRGEVHALMGANGSGKSTFAKVLVGHPDYEVTGGRAVYKGENLLDME 378
>TC9818 similar to UP|Q7XA50 (Q7XA50) Sorbitol-like transporter, partial
(62%)
Length = 1162
Score = 32.3 bits (72), Expect = 0.17
Identities = 21/76 (27%), Positives = 38/76 (49%), Gaps = 9/76 (11%)
Frame = +2
Query: 29 YVLMFIGSI---GAIVHGASVPIFFIFFGKLINVIGLAY------LFPKEASHKVAKSKC 79
Y ++F G+I GA++ G S +F+ FG+ I IG+ Y ++ E S ++
Sbjct: 440 YTIVFAGAIFFVGALLMGFSPNYWFLMFGRFIAGIGIGYALMIAPVYTAEVSPASSRGFL 619
Query: 80 SNIIFILDRGGLLDAY 95
++ + GG+L Y
Sbjct: 620 TSFPEVFINGGILLGY 667
>TC16400 similar to UP|O48783 (O48783) At2g26680 protein, partial (58%)
Length = 1053
Score = 30.8 bits (68), Expect = 0.49
Identities = 13/35 (37%), Positives = 21/35 (59%)
Frame = +1
Query: 149 FLHYISRFIAGFTIGFVRVWQISLVTLSIVPAIAL 183
FL RF+ GFT+ +R+W +SL L ++ + L
Sbjct: 130 FLRICRRFVKGFTLIELRIWLLSLRLLHLIVLVTL 234
>AU089197
Length = 658
Score = 30.4 bits (67), Expect = 0.64
Identities = 23/77 (29%), Positives = 37/77 (47%), Gaps = 1/77 (1%)
Frame = +1
Query: 345 DGHIQFNDVCFSY-PSRPDVGIFTNLNLDIPAGKIVALVGGSGSGKSTVVSLIERFYEPI 403
DG I+ + + Y P+ P V ++ K + + +GSGKST++ + P+
Sbjct: 349 DGMIELHKLQVQYDPAAPIV--LKDVTFIFSGQKKIGIXXRTGSGKSTLLQALF*IVXPL 522
Query: 404 SGQILLDKNDIRELDLK 420
SG IL+D I L K
Sbjct: 523 SGCILIDGVXIPRLACK 573
>TC16536 similar to UP|O22477 (O22477) Betaine aldehyde dehydrogenase ,
partial (71%)
Length = 1097
Score = 29.6 bits (65), Expect = 1.1
Identities = 21/66 (31%), Positives = 31/66 (46%)
Frame = +2
Query: 148 NFLHYISRFIAGFTIGFVRVWQISLVTLSIVPAIALAGGCYAYVTIGLIAKVRKAYVRAG 207
NF Y+ R G +G + W ++ + A ALA GC A + +A V + G
Sbjct: 449 NFKSYVLREPIG-VVGLITPWNYPMLMATWKVAPALAAGCAAILKPSELASV--TCLELG 619
Query: 208 EIAEEV 213
EI +EV
Sbjct: 620 EICKEV 637
>BP042433
Length = 534
Score = 29.3 bits (64), Expect = 1.4
Identities = 30/116 (25%), Positives = 52/116 (43%), Gaps = 7/116 (6%)
Frame = +2
Query: 300 SLGQAAPDISAFIRAKAAAYPIFEMI----ERDTVSKKSS---KTGRKLSKLDGHIQFND 352
SLG + R A I+E++ E V +KSS + R +I+F+
Sbjct: 101 SLGTLSISARRLNRLSGYADRIYELMSVSRELHLVDEKSSLQRQGSRNCISEANYIEFSG 280
Query: 353 VCFSYPSRPDVGIFTNLNLDIPAGKIVALVGGSGSGKSTVVSLIERFYEPISGQIL 408
V P+ + +L L + G + + G +GSGKS++ ++ + ISG I+
Sbjct: 281 VKVVTPTGNV--LVDDLTLRVEPGSNLLITGPNGSGKSSLFRVLGGLWPMISGHIV 442
Database: LJGI
Posted date: Jul 30, 2004 11:16 AM
Number of letters in database: 14,692,800
Number of sequences in database: 28,460
Lambda K H
0.327 0.142 0.428
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 7,577,391
Number of Sequences: 28460
Number of extensions: 105432
Number of successful extensions: 672
Number of sequences better than 10.0: 51
Number of HSP's better than 10.0 without gapping: 672
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 672
length of query: 461
length of database: 4,897,600
effective HSP length: 93
effective length of query: 368
effective length of database: 2,250,820
effective search space: 828301760
effective search space used: 828301760
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 15 ( 7.1 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 40 (21.7 bits)
S2: 57 (26.6 bits)
Medicago: description of AC087771.6


