

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC149581.7 + phase: 0
(392 letters)
Database: GMGI
63,676 sequences; 37,918,896 total letters
Searching..................................................done
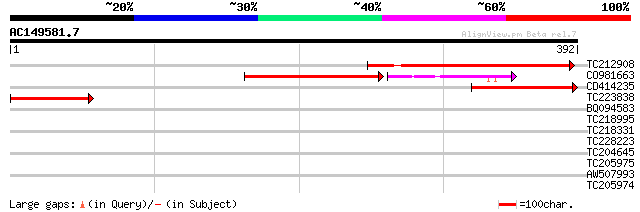
Score E
Sequences producing significant alignments: (bits) Value
TC212908 233 1e-61
CO981663 108 4e-34
CD414235 123 1e-28
TC223838 111 5e-25
BQ094583 33 0.002
TC218995 similar to GB|AAD34703.1|4966372|F3O9 ESTs gb|N38586 an... 30 1.4
TC218331 similar to UP|Q9LRZ7 (Q9LRZ7) Arabidopsis thaliana geno... 29 3.2
TC228223 29 3.2
TC204645 similar to UP|Q9FM07 (Q9FM07) Permease 1, complete 28 7.1
TC205975 homologue to UP|Q94KS2 (Q94KS2) TGF-beta receptor-inter... 28 9.3
AW507993 similar to PIR|T09153|T091 glucose-6-phosphate isomeras... 28 9.3
TC205974 homologue to UP|Q94KS2 (Q94KS2) TGF-beta receptor-inter... 28 9.3
>TC212908
Length = 597
Score = 233 bits (593), Expect = 1e-61
Identities = 109/143 (76%), Positives = 124/143 (86%)
Frame = +1
Query: 248 FHKNIAENRYVKVSSQRGGKEGGFGPENMLDSDHLWSYWTPREDDKEKDHWIEIWGNDGS 307
FH+N+AE+ +VKVSSQR G FGPE MLDS+HLWSYW PR+DD EKDHWIEIW +G
Sbjct: 1 FHQNLAEDCFVKVSSQRRG----FGPEKMLDSEHLWSYWAPRDDDGEKDHWIEIWAKEGK 168
Query: 308 LRFNVIRIQEAIGLGQRIERYEIYVDGKSIIQGTTIGYKRLHRLDGDVVHARVVRIRFIK 367
RFNVIRIQEAIGLGQRI+R+EIYVDGK II+ TT+GY+RLHRL+G VHARVVRIR K
Sbjct: 169 FRFNVIRIQEAIGLGQRIKRHEIYVDGKLIIEATTVGYQRLHRLNGGEVHARVVRIRIRK 348
Query: 368 ARGVPLISSIGLHFDPFWHSRFT 390
ARGVPLISSIGLH+DPFWHS+FT
Sbjct: 349 ARGVPLISSIGLHYDPFWHSKFT 417
>CO981663
Length = 825
Score = 108 bits (271), Expect(2) = 4e-34
Identities = 53/96 (55%), Positives = 64/96 (66%)
Frame = -3
Query: 163 TQYLNTGDPKGTDWLPAECDVSIRPGWFWHKSESPKKLSDLLDIYYKSVGRNCVLLLNVP 222
+QY + GD DW+PA CDVSIRP WFWH SE PK LL+I KSVGRNC LLLNV
Sbjct: 808 SQYNSEGDQFCPDWIPATCDVSIRPSWFWHASEQPKSARSLLEI**KSVGRNCQLLLNVT 629
Query: 223 PNTTGLISENDAHRLKEFRSAIDTIFHKNIAENRYV 258
PN++GLIS+ D L+EF +IF N+A N +
Sbjct: 628 PNSSGLISDEDIQILQEFTELRRSIFSHNLAINSVI 521
Score = 53.9 bits (128), Expect(2) = 4e-34
Identities = 34/95 (35%), Positives = 56/95 (58%), Gaps = 6/95 (6%)
Frame = -2
Query: 262 SQRGG-KEGGFGPENMLDSDHLWSYWTPREDDKEKDHWIEIWGNDGSLRFNVIRIQEAIG 320
S RGG +E F P N+L+ + +++YW P E+ + WI + FN++++QE I
Sbjct: 512 SIRGGIQETHFSPYNVLE-EGIYTYWAPGENQTK---WILYIDLQELVSFNILQLQEPIN 345
Query: 321 LGQRIERYE---IYVDG--KSIIQGTTIGYKRLHR 350
+GQR+ + + +DG ++ GTTIGY+RL R
Sbjct: 344 MGQRVIEFHLEALSLDGVWMRVVYGTTIGYQRLLR 240
>CD414235
Length = 296
Score = 123 bits (309), Expect = 1e-28
Identities = 58/73 (79%), Positives = 65/73 (88%)
Frame = -2
Query: 320 GLGQRIERYEIYVDGKSIIQGTTIGYKRLHRLDGDVVHARVVRIRFIKARGVPLISSIGL 379
GLGQRI+R+EIYVDGK II+ TT+GYKRLHRLDG VHAR VRIR KARGVPLISSIGL
Sbjct: 289 GLGQRIKRHEIYVDGKLIIKATTVGYKRLHRLDGGEVHARGVRIRIRKARGVPLISSIGL 110
Query: 380 HFDPFWHSRFTAA 392
H+DPFWHS+FT +
Sbjct: 109 HYDPFWHSKFTVS 71
>TC223838
Length = 478
Score = 111 bits (278), Expect = 5e-25
Identities = 49/58 (84%), Positives = 53/58 (90%)
Frame = +3
Query: 1 MILTAKHHDGFCLWPSKYTKHSVISSKWQNGKGDVVKEFVNAATDKGIDVGIYLSPWD 58
MILTAKHHDGFCLWPSKYT HSVISS WQ GKGDVV++FVNAAT +GIDV +YLSPWD
Sbjct: 303 MILTAKHHDGFCLWPSKYTDHSVISSPWQGGKGDVVQDFVNAATAQGIDVXVYLSPWD 476
>BQ094583
Length = 422
Score = 32.7 bits (73), Expect(2) = 0.002
Identities = 12/15 (80%), Positives = 13/15 (86%)
Frame = +3
Query: 1 MILTAKHHDGFCLWP 15
+ILT KHHDGF LWP
Sbjct: 312 VILTVKHHDGFFLWP 356
Score = 26.2 bits (56), Expect(2) = 0.002
Identities = 10/15 (66%), Positives = 12/15 (79%)
Frame = +2
Query: 18 YTKHSVISSKWQNGK 32
YT +SV SS W+NGK
Sbjct: 362 YTNYSVRSSNWRNGK 406
>TC218995 similar to GB|AAD34703.1|4966372|F3O9 ESTs gb|N38586 and gb|N38613
come from this gene. {Arabidopsis thaliana;} , partial
(33%)
Length = 759
Score = 30.4 bits (67), Expect = 1.4
Identities = 14/40 (35%), Positives = 20/40 (50%)
Frame = -3
Query: 152 RTSLSIGASNITQYLNTGDPKGTDWLPAECDVSIRPGWFW 191
R+S G+ IT +++G K T CD+ I P W W
Sbjct: 133 RSSCWYGSVGITDRISSGRSKSTTEPVPCCDIPIHPVWLW 14
>TC218331 similar to UP|Q9LRZ7 (Q9LRZ7) Arabidopsis thaliana genomic DNA,
chromosome 3, TAC clone:K13N2, partial (16%)
Length = 685
Score = 29.3 bits (64), Expect = 3.2
Identities = 30/107 (28%), Positives = 40/107 (37%), Gaps = 10/107 (9%)
Frame = -1
Query: 207 YYKSVGRNCVLLLNVPPNTTGLISENDAH--------RLKEFRSAIDTIFHKNIAENRYV 258
Y K + N L P +SE AH R K R + H I ++
Sbjct: 451 YSKFMKGNIKKKLRERPTREFNVSELMAHTGSPGA*GRRKSHRM*TPSATHSAI*STLFL 272
Query: 259 KVSSQRGGKEGGFGPENMLDSDHLWSYWTPR--EDDKEKDHWIEIWG 303
K+SS E G G ++ H W TPR EK W+E+ G
Sbjct: 271 KLSSLSCSTEDGVGLALAREARHKWVPETPRFALGMAEKKRWVELSG 131
>TC228223
Length = 849
Score = 29.3 bits (64), Expect = 3.2
Identities = 17/63 (26%), Positives = 26/63 (40%), Gaps = 9/63 (14%)
Frame = -3
Query: 247 IFHKNIAENRYVKVSSQRGGKE----GGFGPENMLDSDHLWS-----YWTPREDDKEKDH 297
++HKN+ EN Y+K R + G P+ + LW ++ KE H
Sbjct: 286 MYHKNLDENPYIKNKKDRAQRPY*PCGKISPQPPCHALLLWQQQ*GHHYQENSQSKEAPH 107
Query: 298 WIE 300
W E
Sbjct: 106 WKE 98
>TC204645 similar to UP|Q9FM07 (Q9FM07) Permease 1, complete
Length = 1930
Score = 28.1 bits (61), Expect = 7.1
Identities = 15/41 (36%), Positives = 24/41 (57%), Gaps = 2/41 (4%)
Frame = -1
Query: 181 CDVSIRPGWFWHKSESPKKLSDLLDIYYK--SVGRNCVLLL 219
C+VSIRP + +S S + L ++ YK S+ C+LL+
Sbjct: 382 CNVSIRPSNYCRQSSSKQSLQQCVNSSYKK*SLDNFCLLLI 260
>TC205975 homologue to UP|Q94KS2 (Q94KS2) TGF-beta receptor-interacting
protein 1, partial (35%)
Length = 546
Score = 27.7 bits (60), Expect = 9.3
Identities = 12/28 (42%), Positives = 15/28 (52%)
Frame = +1
Query: 333 DGKSIIQGTTIGYKRLHRLDGDVVHARV 360
DGKS G GY RLH D D + ++
Sbjct: 262 DGKSFSSGGEDGYVRLHHFDPDYFNIKI 345
>AW507993 similar to PIR|T09153|T091 glucose-6-phosphate isomerase (EC
5.3.1.9) precursor chloroplast - spinach, partial (13%)
Length = 456
Score = 27.7 bits (60), Expect = 9.3
Identities = 13/32 (40%), Positives = 14/32 (43%), Gaps = 7/32 (21%)
Frame = -3
Query: 173 GTDWLPAECD-------VSIRPGWFWHKSESP 197
GT W P + SIR GW WH S P
Sbjct: 448 GTRWAPEQSP*ASSAPPFSIRRGWRWHPSRGP 353
>TC205974 homologue to UP|Q94KS2 (Q94KS2) TGF-beta receptor-interacting
protein 1, partial (57%)
Length = 783
Score = 27.7 bits (60), Expect = 9.3
Identities = 12/28 (42%), Positives = 15/28 (52%)
Frame = +3
Query: 333 DGKSIIQGTTIGYKRLHRLDGDVVHARV 360
DGKS G GY RLH D D + ++
Sbjct: 483 DGKSFSSGGEDGYVRLHHFDPDYFNIKI 566
Database: GMGI
Posted date: Oct 22, 2004 4:58 PM
Number of letters in database: 37,918,896
Number of sequences in database: 63,676
Lambda K H
0.319 0.138 0.442
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 19,580,297
Number of Sequences: 63676
Number of extensions: 272867
Number of successful extensions: 1420
Number of sequences better than 10.0: 24
Number of HSP's better than 10.0 without gapping: 1406
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 1418
length of query: 392
length of database: 12,639,632
effective HSP length: 99
effective length of query: 293
effective length of database: 6,335,708
effective search space: 1856362444
effective search space used: 1856362444
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 16 ( 7.4 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.7 bits)
S2: 60 (27.7 bits)
Medicago: description of AC149581.7


