

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
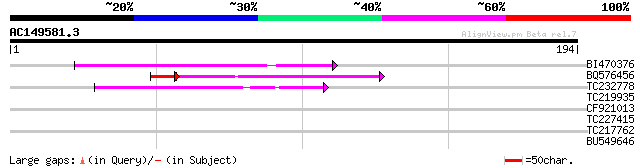
Query= AC149581.3 + phase: 0
(194 letters)
Database: GMGI
63,676 sequences; 37,918,896 total letters
Searching..................................................done
Score E
Sequences producing significant alignments: (bits) Value
BI470376 weakly similar to GP|14090338|dbj P0638D12.5 {Oryza sat... 49 1e-06
BQ576456 45 5e-06
TC232778 similar to UP|Q9LNG5 (Q9LNG5) F21D18.16, partial (5%) 46 1e-05
TC219935 similar to UP|P93081 (P93081) 3-hydroxy-3-methylglutary... 29 1.2
CF921013 27 4.5
TC227415 similar to UP|Q9LVD9 (Q9LVD9) Gb|AAC28507.1, partial (89%) 27 7.7
TC217762 similar to UP|SOX_ARATH (Q9SJA7) Potential sarcosine ox... 27 7.7
BU549646 27 7.7
>BI470376 weakly similar to GP|14090338|dbj P0638D12.5 {Oryza sativa
(japonica cultivar-group)}, partial (5%)
Length = 429
Score = 49.3 bits (116), Expect = 1e-06
Identities = 31/90 (34%), Positives = 46/90 (50%)
Frame = -1
Query: 23 LHDLIYTGYSTVTHAMICAMCERWHTETSSFHLLVGEMTIPLDDMHNLLHIPIHGRILDH 82
L+ ++ V A++ A ERW ET +FHL GE TI L D+ LL IP+ GR L
Sbjct: 378 LYPIMKLAQLKVNGALVNAFIERWRPETHTFHLKYGEATITLQDVSVLLGIPVDGRPLIG 199
Query: 83 DEAMNQDCAIDLMIRLLGMSEVDVRVEVRS 112
+ ++ +L LLG+ D ++ S
Sbjct: 198 NTNIDW---FELFHELLGVMPDDAAIDGNS 118
>BQ576456
Length = 424
Score = 45.1 bits (105), Expect(2) = 5e-06
Identities = 26/71 (36%), Positives = 40/71 (55%)
Frame = +1
Query: 58 GEMTIPLDDMHNLLHIPIHGRILDHDEAMNQDCAIDLMIRLLGMSEVDVRVEVRSESAGH 117
GE+TI LDDM LLH+PI G L E + D A+ L++ LL +S + R ++ +
Sbjct: 31 GELTITLDDMACLLHLPITG-ALHRFEPLGVDEAVLLLMELLEVSREEARAKIVRAHRAY 207
Query: 118 ISYPTLKRVYE 128
+ L+ VY+
Sbjct: 208 VRLSWLREVYQ 240
Score = 21.6 bits (44), Expect(2) = 5e-06
Identities = 8/10 (80%), Positives = 10/10 (100%)
Frame = +3
Query: 49 ETSSFHLLVG 58
ET++FHLLVG
Sbjct: 3 ETNTFHLLVG 32
>TC232778 similar to UP|Q9LNG5 (Q9LNG5) F21D18.16, partial (5%)
Length = 746
Score = 45.8 bits (107), Expect = 1e-05
Identities = 32/80 (40%), Positives = 41/80 (51%)
Frame = -1
Query: 30 GYSTVTHAMICAMCERWHTETSSFHLLVGEMTIPLDDMHNLLHIPIHGRILDHDEAMNQD 89
GY + A+I A ERW ET +FHL GE TI L D+ LL + G L + N D
Sbjct: 506 GYLKINAALISAFIERWRPETHTFHLRCGEATITLQDVLILLGLRTDGAPL--IGSTNLD 333
Query: 90 CAIDLMIRLLGMSEVDVRVE 109
A DL LLG+ + +E
Sbjct: 332 WA-DLCEELLGVRPQEGEIE 276
>TC219935 similar to UP|P93081 (P93081) 3-hydroxy-3-methylglutaryl coenzyme A
reductase , partial (9%)
Length = 414
Score = 29.3 bits (64), Expect = 1.2
Identities = 17/62 (27%), Positives = 33/62 (52%), Gaps = 3/62 (4%)
Frame = -2
Query: 95 MIRLLGMSEVDVRVEVRSESAGHISYPTLKRVYEHHLTEARRLEEPQTREKLQE---RGR 151
+++LLG+S + R+ S+ ++ +PT E R++E P+ EKL + G+
Sbjct: 242 VLQLLGLSAI---AGGRNGSSANVHFPTRTESTECRCRRRRKMENPEAEEKLGDWKGNGK 72
Query: 152 KR 153
+R
Sbjct: 71 RR 66
>CF921013
Length = 598
Score = 27.3 bits (59), Expect = 4.5
Identities = 15/43 (34%), Positives = 22/43 (50%), Gaps = 5/43 (11%)
Frame = +3
Query: 20 GFDLHDLIYTGYSTVTHAMICAMCERWH-----TETSSFHLLV 57
GF + + ++GY V I MC + H +T +FHLLV
Sbjct: 444 GFWIQEFRHSGYD*VFRDKIIVMCPKTHLFC*FRKTRTFHLLV 572
>TC227415 similar to UP|Q9LVD9 (Q9LVD9) Gb|AAC28507.1, partial (89%)
Length = 1748
Score = 26.6 bits (57), Expect = 7.7
Identities = 28/98 (28%), Positives = 38/98 (38%), Gaps = 5/98 (5%)
Frame = -2
Query: 68 HNLLHIPIHG---RILDHDEAMNQDCAIDLMIRLLGMSEVDVRVEVRSESAGHISYPT-- 122
H HG R L +E++ C RLL ++R+ VR S H YPT
Sbjct: 580 HRFAEYVYHGQPLRRLVREESLTAAC------RLLTSRTFELRMLVRIASPRHSPYPT*G 419
Query: 123 LKRVYEHHLTEARRLEEPQTREKLQERGRKRAIGKWSW 160
+R + RR R RG +R G W+W
Sbjct: 418 RRRTFGCRCCRGRR------RRVRGCRGGRRRFG-WTW 326
>TC217762 similar to UP|SOX_ARATH (Q9SJA7) Potential sarcosine oxidase ,
partial (58%)
Length = 1312
Score = 26.6 bits (57), Expect = 7.7
Identities = 15/41 (36%), Positives = 22/41 (53%)
Frame = +3
Query: 113 ESAGHISYPTLKRVYEHHLTEARRLEEPQTREKLQERGRKR 153
+S+GH+ P+L++ RRLE + QERGR R
Sbjct: 468 KSSGHVPNPSLQK--------RRRLEGQHQGHRHQERGRHR 566
>BU549646
Length = 693
Score = 26.6 bits (57), Expect = 7.7
Identities = 13/29 (44%), Positives = 17/29 (57%)
Frame = +1
Query: 124 KRVYEHHLTEARRLEEPQTREKLQERGRK 152
K V+ HL+ A R+EE T +LQE K
Sbjct: 523 KLVWYLHLSPAVRIEETTTNSQLQENSHK 609
Database: GMGI
Posted date: Oct 22, 2004 4:58 PM
Number of letters in database: 37,918,896
Number of sequences in database: 63,676
Lambda K H
0.323 0.139 0.433
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 8,665,121
Number of Sequences: 63676
Number of extensions: 114835
Number of successful extensions: 727
Number of sequences better than 10.0: 16
Number of HSP's better than 10.0 without gapping: 726
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 727
length of query: 194
length of database: 12,639,632
effective HSP length: 92
effective length of query: 102
effective length of database: 6,781,440
effective search space: 691706880
effective search space used: 691706880
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 16 ( 7.5 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (22.0 bits)
S2: 56 (26.2 bits)
Medicago: description of AC149581.3


