

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC149576.6 - phase: 0
(597 letters)
Database: GMGI
63,676 sequences; 37,918,896 total letters
Searching..................................................done
Score E
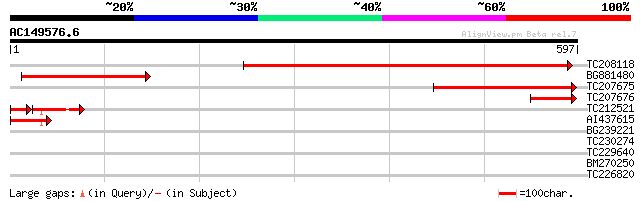
Sequences producing significant alignments: (bits) Value
TC208118 similar to UP|Q9FL94 (Q9FL94) Gb|AAC61821.1, partial (36%) 474 e-134
BG881480 246 2e-65
TC207675 weakly similar to UP|Q9LDF8 (Q9LDF8) Gb|AAF35944.1 (At3... 218 6e-57
TC207676 similar to UP|NU1M_PARTE (P05513) NADH-ubiquinone oxido... 79 4e-15
TC212521 57 1e-11
AI437615 46 4e-05
BG239221 39 0.009
TC230274 30 4.0
TC229640 similar to UP|Q6NMI3 (Q6NMI3) At1g31780, partial (52%) 30 4.0
BM270250 29 5.2
TC226820 similar to GB|AAP37788.1|30725532|BT008429 At5g53620 {A... 28 8.8
>TC208118 similar to UP|Q9FL94 (Q9FL94) Gb|AAC61821.1, partial (36%)
Length = 1369
Score = 474 bits (1221), Expect = e-134
Identities = 241/350 (68%), Positives = 287/350 (81%), Gaps = 4/350 (1%)
Frame = +3
Query: 247 YPNQKMFHPLPPSLGPGVYLGAVERATSFITDDLWYGIFAGTNPETFVRADGAFIPFAED 306
YPNQKMFHPLPP+LGPGVYLGAVERATSFITD+LWYGIFAG NPETFVRADGAFIPFA+D
Sbjct: 3 YPNQKMFHPLPPTLGPGVYLGAVERATSFITDELWYGIFAGINPETFVRADGAFIPFADD 182
Query: 307 FNMNNVITSIRGVGDIGEVHRIDLQSPINSLIGRQVIKVGRSSGLTTGTIMAYALEYNDE 366
F+M+ V TS+RGVGDIG+V IDLQ+PI+SLIG+QV+KVGRSSGLTTG ++AYALEYNDE
Sbjct: 183 FDMSTVTTSVRGVGDIGDVKIIDLQAPISSLIGKQVVKVGRSSGLTTGVVLAYALEYNDE 362
Query: 367 KGICFLTDFLVVGENQQTFDLEGDSGSLILLTGQNREKPRPVGIIWGGTANRGRLKLRVG 426
KGICFLTD LVVGENQQTFDLEGDSGSLI+L G N EKPRP+GIIWGGTANRGRLKL+VG
Sbjct: 363 KGICFLTDLLVVGENQQTFDLEGDSGSLIMLKGDNGEKPRPIGIIWGGTANRGRLKLKVG 542
Query: 427 QPPENWTSGVDLGRLLDLLELDLVTTNETLQDSGQEQMNGSTAGIGSTVGESSPT--VPI 484
QPPENWTSGVDLGRLL+LLELDL+TT+E LQ + QEQ S IGSTVG+SSP V
Sbjct: 543 QPPENWTSGVDLGRLLNLLELDLITTDEGLQVAVQEQRAVSATVIGSTVGDSSPPDGVLP 722
Query: 485 KEKLEESFEPFCLNMEHVP--VEEPSTIVKPSLRPCEFHIRNEIETVPNVEHQFIRTSFA 542
K+K E+ +EP L ++ +P V S +KPS+ EF + + I+ P++EHQFI SF
Sbjct: 723 KDKAEDKYEPLGLQIQSIPLGVVPSSQDMKPSIMETEFKLEDGIKVGPSIEHQFI-PSFI 899
Query: 543 GKSPVHQSFLKEDMQFKSLSELRNEPDEDNFVSLHLGEPEAKRRKHSNSS 592
G+SP+H++ +++ ++LS LRN DED VSL LG+ EAKRR+ S+
Sbjct: 900 GRSPLHKNSIQDRTATENLSSLRNNCDEDLCVSLQLGDNEAKRRRSEAST 1049
>BG881480
Length = 413
Score = 246 bits (629), Expect = 2e-65
Identities = 124/137 (90%), Positives = 129/137 (93%), Gaps = 1/137 (0%)
Frame = +3
Query: 13 SGSTQSEESALDLERNYYGHPS-SSPLHMQTFAVGVQHSEGNAAYFSWPTLNRWNDAAED 71
SGSTQSEESALDLER+YYGHP+ SSP +Q FA G QHSE NAAYFSWPTL+RWNDAAED
Sbjct: 3 SGSTQSEESALDLERSYYGHPNPSSPSPLQPFAGGAQHSESNAAYFSWPTLSRWNDAAED 182
Query: 72 RANYFGNLQKGVLPETLGRLPSGQQATTLLELMTIRAFHSKILRRFSLGTAIGFRIRGGV 131
RANYFGNLQKGVLPETLGRLP+GQQATTLLELMTIRAFHSKILRRFSLGTAIGFRIRGGV
Sbjct: 183 RANYFGNLQKGVLPETLGRLPTGQQATTLLELMTIRAFHSKILRRFSLGTAIGFRIRGGV 362
Query: 132 LTDIPAILVFVAHKVHR 148
LTDIPAILVFVA KVHR
Sbjct: 363 LTDIPAILVFVARKVHR 413
>TC207675 weakly similar to UP|Q9LDF8 (Q9LDF8) Gb|AAF35944.1 (At3g12950),
partial (6%)
Length = 841
Score = 218 bits (555), Expect = 6e-57
Identities = 113/151 (74%), Positives = 124/151 (81%), Gaps = 1/151 (0%)
Frame = +2
Query: 447 LDLVTTNETLQDSGQEQMNGSTAGIGSTVGESSPTVPIKEKLEESFEPFCLNMEHVPVE- 505
LDL+TTNE LQ + EQ NGS AGI STVGESSPTVPIKEKLEESFEPFCLN+ VE
Sbjct: 2 LDLITTNEALQAAVLEQRNGSAAGIDSTVGESSPTVPIKEKLEESFEPFCLNIPLAQVED 181
Query: 506 EPSTIVKPSLRPCEFHIRNEIETVPNVEHQFIRTSFAGKSPVHQSFLKEDMQFKSLSELR 565
EPS V PS+RPCEFHI++EIE PNVEHQFI S+AGKSP QS+LKEDM+ KSL+ELR
Sbjct: 182 EPSQRVNPSIRPCEFHIKSEIEIAPNVEHQFI-PSYAGKSPARQSYLKEDMELKSLAELR 358
Query: 566 NEPDEDNFVSLHLGEPEAKRRKHSNSSLSLK 596
N PDEDNFVSLHLGEPE KRRK SNSS +K
Sbjct: 359 NGPDEDNFVSLHLGEPEMKRRKLSNSSFCIK 451
>TC207676 similar to UP|NU1M_PARTE (P05513) NADH-ubiquinone oxidoreductase
chain 1 , partial (6%)
Length = 433
Score = 79.3 bits (194), Expect = 4e-15
Identities = 38/48 (79%), Positives = 42/48 (87%)
Frame = +3
Query: 549 QSFLKEDMQFKSLSELRNEPDEDNFVSLHLGEPEAKRRKHSNSSLSLK 596
QS+LKEDM+ KSL+ELRN PDEDNFVSLHLGEPE KRRK SNSS +K
Sbjct: 15 QSYLKEDMELKSLAELRNGPDEDNFVSLHLGEPEMKRRKISNSSFCIK 158
>TC212521
Length = 381
Score = 57.4 bits (137), Expect(2) = 1e-11
Identities = 33/57 (57%), Positives = 37/57 (64%), Gaps = 3/57 (5%)
Frame = +3
Query: 25 LERNYYGH---PSSSPLHMQTFAVGVQHSEGNAAYFSWPTLNRWNDAAEDRANYFGN 78
LERN H PS SP +Q FA QH E +AAYFSW +R NDAAE+RANYF N
Sbjct: 216 LERNCCSHSNLPSLSPPTLQPFASAGQHCESSAAYFSWS--SRLNDAAEERANYFLN 380
Score = 30.4 bits (67), Expect(2) = 1e-11
Identities = 15/23 (65%), Positives = 16/23 (69%)
Frame = +2
Query: 1 MNRNRLGLSAHHSGSTQSEESAL 23
M R RL + H SGST SEESAL
Sbjct: 143 MERARLNMRGHCSGSTPSEESAL 211
>AI437615
Length = 310
Score = 46.2 bits (108), Expect = 4e-05
Identities = 27/47 (57%), Positives = 29/47 (61%), Gaps = 3/47 (6%)
Frame = +1
Query: 1 MNRNRLGLSAHHSGSTQSEESALDLERNYYGH---PSSSPLHMQTFA 44
M R RL + SGST SEESALDLERN H PS SP +Q FA
Sbjct: 157 MERTRLNMRGRCSGSTPSEESALDLERNCCSHSNLPSLSPPTLQPFA 297
>BG239221
Length = 334
Score = 38.5 bits (88), Expect = 0.009
Identities = 15/15 (100%), Positives = 15/15 (100%)
Frame = +2
Query: 279 DLWYGIFAGTNPETF 293
DLWYGIFAGTNPETF
Sbjct: 290 DLWYGIFAGTNPETF 334
>TC230274
Length = 800
Score = 29.6 bits (65), Expect = 4.0
Identities = 16/38 (42%), Positives = 22/38 (57%), Gaps = 1/38 (2%)
Frame = +3
Query: 180 GAPAPTPKEQ-LYTELADGLRGSDSCVGSGSQVASQET 216
G+P P++Q ++ + DG DSC S VASQET
Sbjct: 183 GSPLIQPQDQPIWQDSCDGKVKEDSCTSSDVGVASQET 296
>TC229640 similar to UP|Q6NMI3 (Q6NMI3) At1g31780, partial (52%)
Length = 1495
Score = 29.6 bits (65), Expect = 4.0
Identities = 23/63 (36%), Positives = 29/63 (45%), Gaps = 1/63 (1%)
Frame = +1
Query: 454 ETLQDSGQEQM-NGSTAGIGSTVGESSPTVPIKEKLEESFEPFCLNMEHVPVEEPSTIVK 512
+T+ DSG +Q N S G G T ES + E PF L +E V +PS IV
Sbjct: 184 DTITDSGPKQFSNNSENGSGKT--ESDLMFVLDRIFEGVCRPFKLRVEQVLQSQPSLIVS 357
Query: 513 PSL 515
L
Sbjct: 358 YKL 366
>BM270250
Length = 191
Score = 29.3 bits (64), Expect = 5.2
Identities = 15/38 (39%), Positives = 22/38 (57%), Gaps = 2/38 (5%)
Frame = -1
Query: 562 SELRNEP--DEDNFVSLHLGEPEAKRRKHSNSSLSLKN 597
++ RN+P D N ++LH+ + KR KH SS S N
Sbjct: 119 NQARNKPHFDTKNILTLHMKIGDQKRLKHQGSSCSTTN 6
>TC226820 similar to GB|AAP37788.1|30725532|BT008429 At5g53620 {Arabidopsis
thaliana;} , partial (18%)
Length = 2036
Score = 28.5 bits (62), Expect = 8.8
Identities = 11/25 (44%), Positives = 17/25 (68%)
Frame = +3
Query: 138 ILVFVAHKVHRQWLNHVQCLPAALE 162
+ +F + KVHR+WL +Q LP L+
Sbjct: 1119 VTIFSSVKVHRKWLLQLQLLPKLLQ 1193
Database: GMGI
Posted date: Oct 22, 2004 4:58 PM
Number of letters in database: 37,918,896
Number of sequences in database: 63,676
Lambda K H
0.317 0.136 0.404
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 25,584,926
Number of Sequences: 63676
Number of extensions: 356914
Number of successful extensions: 1545
Number of sequences better than 10.0: 22
Number of HSP's better than 10.0 without gapping: 1533
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 1539
length of query: 597
length of database: 12,639,632
effective HSP length: 103
effective length of query: 494
effective length of database: 6,081,004
effective search space: 3004015976
effective search space used: 3004015976
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 16 ( 7.3 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.6 bits)
S2: 62 (28.5 bits)
Medicago: description of AC149576.6


