

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
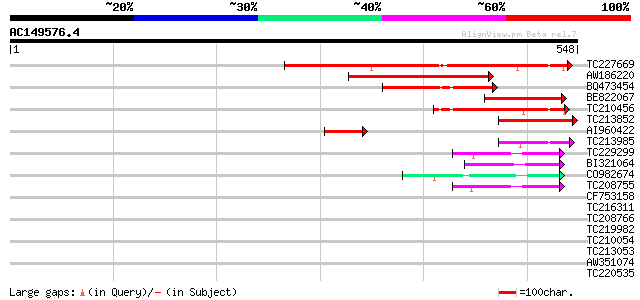
Query= AC149576.4 + phase: 0 /pseudo
(548 letters)
Database: GMGI
63,676 sequences; 37,918,896 total letters
Searching..................................................done
Score E
Sequences producing significant alignments: (bits) Value
TC227669 homologue to UP|Q9AVQ9 (Q9AVQ9) Phosphate transporter, ... 270 9e-73
AW186220 243 2e-64
BQ473454 homologue to GP|24415831|gb phosphate transporter {Medi... 147 2e-35
BE822067 133 2e-31
TC210456 similar to UP|Q9AU01 (Q9AU01) Phosphate transporter 1, ... 131 9e-31
TC213852 120 2e-27
AI960422 64 2e-10
TC213985 homologue to UP|Q9AVR0 (Q9AVR0) Phosphate transporter, ... 60 3e-09
TC229299 similar to UP|GT10_HUMAN (O95528) Solute carrier family... 53 3e-07
BI321064 48 1e-05
CO982674 44 1e-04
TC208755 similar to UP|HXT4_YEAST (P32467) Low-affinity glucose ... 43 4e-04
CF753158 41 0.001
TC216311 UP|Q7XA52 (Q7XA52) Monosaccharide transporter, complete 41 0.001
TC208766 similar to UP|STA_RICCO (Q10710) Sugar carrier protein ... 38 0.010
TC219982 36 0.051
TC210054 homologue to UP|Q7XA52 (Q7XA52) Monosaccharide transpor... 35 0.066
TC213053 similar to UP|Q7XA50 (Q7XA50) Sorbitol-like transporter... 34 0.15
AW351074 similar to GP|9294345|dbj organic anion transporter-lik... 34 0.19
TC220535 similar to UP|HXT5_YEAST (P38695) Probable glucose tran... 34 0.19
>TC227669 homologue to UP|Q9AVQ9 (Q9AVQ9) Phosphate transporter, partial
(55%)
Length = 1226
Score = 270 bits (691), Expect = 9e-73
Identities = 145/290 (50%), Positives = 188/290 (64%), Gaps = 11/290 (3%)
Frame = +3
Query: 266 YTALVEQNVLQAAKDMEKVLDVSMSQITEE-HPLPPATNVAYPLLSREFLWRHGRDLFAC 324
YTALV +N QAA DM KVL V + ++ + + N Y L S+EF RHG L
Sbjct: 12 YTALVAKNAKQAASDMSKVLQVEVEAEEDKLQHMVESENQKYGLFSKEFAKRHGLHLVGT 191
Query: 325 SANWFLLDIVFYSQVLFQSEIYKR--YLNEKDDEDVYQEAFHLARIQAILAVCSTIPGYF 382
+ WFLLDI FYSQ LFQ +I+ ++ D + E + +AR Q ++A+CST+PGY+
Sbjct: 192 TVTWFLLDIAFYSQNLFQKDIFTAIGWIPPAQDMNAIHEVYRIARAQTLIALCSTVPGYW 371
Query: 383 FTVYFIDRVGRVKIQMMGFFFMAVSFFALGFPYYSHWTKGENHDNKGFMVIYGLAFFFAN 442
FTV FID +GR IQ+MGFFFM V FAL PY +HW +NH+N GF+V+Y FFF+N
Sbjct: 372 FTVAFIDIIGRFAIQLMGFFFMTVFMFALAIPY-NHW---KNHNNIGFVVMYSFTFFFSN 539
Query: 443 FGPNTTTFIVPAELFPARFRSTCHGISGAVGKVGAIIGSVGFLWASH----KEKEEGYPK 498
FGPN TTF+VPAE+FPAR RSTCHGIS A GK GAI+G+ GFL+A+ + + GYP
Sbjct: 540 FGPNATTFVVPAEIFPARLRSTCHGISAAAGKAGAIVGAFGFLYAAQSTNPNKVDHGYPT 719
Query: 499 GIGMKASLIILGGVCIVGMFVTYFFTKETMGRSLE----ENEDEQSHHAE 544
GIG+K SLI+LG + GM T E+ G+SLE ENED+ + E
Sbjct: 720 GIGVKNSLIVLGVINFFGMVFT-LLVPESKGKSLEELSGENEDDGAEAIE 866
>AW186220
Length = 425
Score = 243 bits (620), Expect = 2e-64
Identities = 111/140 (79%), Positives = 128/140 (91%)
Frame = +1
Query: 328 WFLLDIVFYSQVLFQSEIYKRYLNEKDDEDVYQEAFHLARIQAILAVCSTIPGYFFTVYF 387
WFL+DIVFYSQVLFQSEIYKRYL++K+D DVYQE FH A IQA++AVCSTIPGYFF++YF
Sbjct: 4 WFLVDIVFYSQVLFQSEIYKRYLDKKEDVDVYQETFHAAWIQAVIAVCSTIPGYFFSMYF 183
Query: 388 IDRVGRVKIQMMGFFFMAVSFFALGFPYYSHWTKGENHDNKGFMVIYGLAFFFANFGPNT 447
ID+ GRVK MMGFFFMA++FF++G PYYS+WT E+H NK FMV+YGLAFFFANFGPNT
Sbjct: 184 IDKWGRVKNPMMGFFFMALAFFSIGIPYYSYWTTKEHHKNKVFMVLYGLAFFFANFGPNT 363
Query: 448 TTFIVPAELFPARFRSTCHG 467
TTFIVPAELFPARFRS+CHG
Sbjct: 364 TTFIVPAELFPARFRSSCHG 423
>BQ473454 homologue to GP|24415831|gb phosphate transporter {Medicago
sativa}, partial (55%)
Length = 426
Score = 147 bits (370), Expect = 2e-35
Identities = 68/111 (61%), Positives = 85/111 (76%)
Frame = +1
Query: 361 EAFHLARIQAILAVCSTIPGYFFTVYFIDRVGRVKIQMMGFFFMAVSFFALGFPYYSHWT 420
E + +AR Q ++A+CST+PGY+FTV ID +GR IQ+MGFFFM V FAL PY+ HW+
Sbjct: 103 EVYKIARAQTLIALCSTVPGYWFTVALIDYMGRFAIQLMGFFFMTVFMFALAIPYH-HWS 279
Query: 421 KGENHDNKGFMVIYGLAFFFANFGPNTTTFIVPAELFPARFRSTCHGISGA 471
+ +N GF+V+Y FFFANFGPN TTF+VPAE+FPAR RSTCHGIS A
Sbjct: 280 EKDNRI--GFVVMYSFTFFFANFGPNATTFVVPAEIFPARLRSTCHGISAA 426
>BE822067
Length = 610
Score = 133 bits (335), Expect = 2e-31
Identities = 62/79 (78%), Positives = 71/79 (89%)
Frame = -1
Query: 460 RFRSTCHGISGAVGKVGAIIGSVGFLWASHKEKEEGYPKGIGMKASLIILGGVCIVGMFV 519
RFRS+CHGISG V VGAIIGSVGFLWASH++KE+GY K IGMK SLIILGGVC++GM +
Sbjct: 610 RFRSSCHGISGXVXXVGAIIGSVGFLWASHRKKEDGYXKXIGMKVSLIILGGVCLLGMVI 431
Query: 520 TYFFTKETMGRSLEENEDE 538
TYFFT+ETMGRSL+ENE E
Sbjct: 430 TYFFTRETMGRSLQENEVE 374
>TC210456 similar to UP|Q9AU01 (Q9AU01) Phosphate transporter 1, partial
(29%)
Length = 617
Score = 131 bits (329), Expect = 9e-31
Identities = 78/148 (52%), Positives = 94/148 (62%), Gaps = 16/148 (10%)
Frame = +2
Query: 410 ALGFPYYSHWTKGENHDNKGFMVIYGLAFFFANFGPNTTTFIVPAELFPARFRSTCHGIS 469
AL PY HWT +N GF VI + FFFANFGPN TTFIVPAE+FPARFRSTCHGIS
Sbjct: 14 ALAVPY-DHWTHKDNRI--GFRVIDAMTFFFANFGPNATTFIVPAEIFPARFRSTCHGIS 184
Query: 470 GAVGKVGAIIGSVGFLW-ASHKEKEEG---YPKGIGMKASLIILGGVCIVGMFVTYFFTK 525
+ GK+GAI+G+ GFL+ A +K+K + YP GIG+K +LI G V I+ + T F
Sbjct: 185 SSSGKLGAIVGAFGFLYLAQNKDKSKADARYPAGIGVKNALIA*GVVNILRFYFT-FLVP 361
Query: 526 ETMGRSLEE------------NEDEQSH 541
E G+SLEE E EQSH
Sbjct: 362 EANGKSLEEMSSENDEDVGTQEESEQSH 445
>TC213852
Length = 411
Score = 120 bits (300), Expect = 2e-27
Identities = 57/76 (75%), Positives = 65/76 (85%)
Frame = +3
Query: 473 GKVGAIIGSVGFLWASHKEKEEGYPKGIGMKASLIILGGVCIVGMFVTYFFTKETMGRSL 532
GKVGAIIGSVGFLWASHK+KE GYPKGIGM+ +LIILG VC++GM VTY FT+ETMGRSL
Sbjct: 36 GKVGAIIGSVGFLWASHKKKENGYPKGIGMEVTLIILGVVCLLGMLVTYLFTRETMGRSL 215
Query: 533 EENEDEQSHHAEEYND 548
EENE E +H E +D
Sbjct: 216 EENEVESVNHEVEEHD 263
>AI960422
Length = 130
Score = 63.5 bits (153), Expect = 2e-10
Identities = 29/42 (69%), Positives = 33/42 (78%)
Frame = +2
Query: 305 AYPLLSREFLWRHGRDLFACSANWFLLDIVFYSQVLFQSEIY 346
AYPLLS E+L +HG DLFACS WFL+DI YSQVL SE+Y
Sbjct: 5 AYPLLSWEYLRKHGPDLFACSFTWFLVDIAVYSQVLVHSELY 130
>TC213985 homologue to UP|Q9AVR0 (Q9AVR0) Phosphate transporter, partial
(15%)
Length = 583
Score = 60.1 bits (144), Expect = 3e-09
Identities = 33/78 (42%), Positives = 47/78 (59%), Gaps = 4/78 (5%)
Frame = +2
Query: 473 GKVGAIIGSVGFLWASHKEK----EEGYPKGIGMKASLIILGGVCIVGMFVTYFFTKETM 528
GK GAI+G+ GFL+A+ + + GYP GIG+K SLI+LG + +GM T E
Sbjct: 71 GKAGAIVGAFGFLYAAQSKDPSKTDAGYPTGIGIKNSLIMLGVINFIGMLFT-LLVPEAK 247
Query: 529 GRSLEENEDEQSHHAEEY 546
G+SLEE E + + E+
Sbjct: 248 GKSLEELSGENNENDAEH 301
>TC229299 similar to UP|GT10_HUMAN (O95528) Solute carrier family 2,
facilitated glucose transporter, member 10 (Glucose
transporter type 10), partial (6%)
Length = 1060
Score = 53.1 bits (126), Expect = 3e-07
Identities = 33/110 (30%), Positives = 51/110 (46%), Gaps = 2/110 (1%)
Frame = +2
Query: 429 GFMVIYGLAFFFANFGPN--TTTFIVPAELFPARFRSTCHGISGAVGKVGAIIGSVGFLW 486
G+ + GLA + F P T ++V +E++P R+R C GI+ + +I S FL
Sbjct: 521 GWAALIGLALYIIFFSPGMGTVPWVVNSEIYPLRYRGVCGGIASTTVWISNLIVSESFL- 697
Query: 487 ASHKEKEEGYPKGIGMKASLIILGGVCIVGMFVTYFFTKETMGRSLEENE 536
K +G + ++ G V IV +F F ET G +EE E
Sbjct: 698 --------SLTKALGTAWTFMMFGIVAIVAIFFVIIFVPETKGVPMEEVE 823
>BI321064
Length = 284
Score = 47.8 bits (112), Expect = 1e-05
Identities = 29/97 (29%), Positives = 47/97 (47%)
Frame = +3
Query: 440 FANFGPNTTTFIVPAELFPARFRSTCHGISGAVGKVGAIIGSVGFLWASHKEKEEGYPKG 499
F + G +T ++V +E++P R+R C GI+ V +I S FL +
Sbjct: 9 FFSPGMDTVPWVVNSEIYPLRYRGVCGGIASTTCWVSNLIVSQSFLTLT---------VA 161
Query: 500 IGMKASLIILGGVCIVGMFVTYFFTKETMGRSLEENE 536
IG + ++ G V ++G+F F ET G +EE E
Sbjct: 162 IGTAWTFMLFGFVALIGIFFVLIFVPETKGVPVEEVE 272
>CO982674
Length = 703
Score = 44.3 bits (103), Expect = 1e-04
Identities = 46/160 (28%), Positives = 62/160 (38%), Gaps = 3/160 (1%)
Frame = -1
Query: 380 GYFFTVYFIDRVGRVKIQMMGFFFMAVSFF--ALGFPYYSHWTKGENHDNKG-FMVIYGL 436
G +Y IDR GR K+ + + MA+S F A G + G N G M I+
Sbjct: 589 GALCALYLIDREGRQKLLIGSYLGMAISMFLVASGIIFPLDEQLGNNLSILGTIMYIFSF 410
Query: 437 AFFFANFGPNTTTFIVPAELFPARFRSTCHGISGAVGKVGAIIGSVGFLWASHKEKEEGY 496
A G T I+ EL R R G S + V + + FL K
Sbjct: 409 AI-----GAGPVTGIIIPELSSTRTRGKIMGFSFSTHWVCNFVVGLFFLELVDK------ 263
Query: 497 PKGIGMKASLIILGGVCIVGMFVTYFFTKETMGRSLEENE 536
G+ G + ++ Y+F ET GRSLEE E
Sbjct: 262 ---FGVAPVYASFGAISLLAATFAYYFIVETKGRSLEEIE 152
>TC208755 similar to UP|HXT4_YEAST (P32467) Low-affinity glucose transporter
HXT4 (Low-affinity glucose transporter LGT1), partial
(3%)
Length = 701
Score = 42.7 bits (99), Expect = 4e-04
Identities = 27/110 (24%), Positives = 47/110 (42%), Gaps = 2/110 (1%)
Frame = +3
Query: 429 GFMVIYGLAFFFANFGP--NTTTFIVPAELFPARFRSTCHGISGAVGKVGAIIGSVGFLW 486
G++ I GLA + A F P + V +E++P +R C G+S V V +I FL
Sbjct: 21 GWLAILGLALYIAFFSPGMGPVPWTVNSEVYPEEYRGICGGMSATVNWVSNLIVVQSFL- 197
Query: 487 ASHKEKEEGYPKGIGMKASLIILGGVCIVGMFVTYFFTKETMGRSLEENE 536
+G + +I+ + ++ + ET G + +E E
Sbjct: 198 --------SVAAAVGTGPTFLIIAIIAVLAFMFVVVYVPETKGLTFDEVE 323
>CF753158
Length = 473
Score = 41.2 bits (95), Expect = 0.001
Identities = 22/51 (43%), Positives = 30/51 (58%)
Frame = -3
Query: 496 YPKGIGMKASLIILGGVCIVGMFVTYFFTKETMGRSLEENEDEQSHHAEEY 546
YP GIG+K SLI+LG + +GM T E G+SLEE E + + E+
Sbjct: 387 YPTGIGIKNSLIMLGVINFIGMLFT-LLVPEAKGKSLEELSGENNENDAEH 238
>TC216311 UP|Q7XA52 (Q7XA52) Monosaccharide transporter, complete
Length = 1925
Score = 41.2 bits (95), Expect = 0.001
Identities = 52/212 (24%), Positives = 86/212 (40%), Gaps = 10/212 (4%)
Frame = +1
Query: 333 IVFYSQVLFQSEIYKRYLNEKDDEDVYQEAFHLARIQAILAVCSTIPGYFFTVYFIDRVG 392
I+FY+ VLF S + KDD A + A++ + ++Y +D+ G
Sbjct: 1009 IMFYAPVLFSS------IGFKDDA---------ALMSAVITGVVNVVATCVSIYGVDKWG 1143
Query: 393 RVKIQMMGFFFM----AVSFFALGFPYYSHWTKGENHDNKGFMVIYGLAFFFANF----G 444
R + + G M AV A+G + + G+ +V+ + + + F G
Sbjct: 1144 RRALFLEGGVQMLICQAVVAAAIGAKFGTDGNPGDLPKWYAIVVVLFICIYVSAFAWSWG 1323
Query: 445 PNTTTFIVPAELFPARFRSTCHGISGAVGKVGA-IIGSVGFLWASHKEKEEGYPKGIGMK 503
P ++VP+E+FP RS I+ +V + +I V H MK
Sbjct: 1324 P--LGWLVPSEIFPLEIRSAAQSINVSVNMLFTFLIAQVFLTMLCH------------MK 1461
Query: 504 ASLIILGGVCIVGM-FVTYFFTKETMGRSLEE 534
L + ++ M F YFF ET G +EE
Sbjct: 1462 FGLFLFFAFFVLIMTFFVYFFLPETKGIPIEE 1557
>TC208766 similar to UP|STA_RICCO (Q10710) Sugar carrier protein A, partial
(55%)
Length = 1202
Score = 38.1 bits (87), Expect = 0.010
Identities = 55/210 (26%), Positives = 86/210 (40%), Gaps = 8/210 (3%)
Frame = +1
Query: 333 IVFYSQVLFQSEIYKRYLNEKDDEDVYQEAFHLARIQAILAVCSTIPGYFFTVYFIDRVG 392
I+FY+ VLFQS + D + A +LA + F ++ +DR+G
Sbjct: 280 ILFYAPVLFQS------MGFGGDASLISSAL----TGGVLASST-----FISIATVDRLG 414
Query: 393 RVKIQMMGFFFMA----VSFFALGFPYYSHWTKGENHDNKGF--MVIYGLAFFFANFGPN 446
R + + G M + LG + + + +KGF +V+ + F FG +
Sbjct: 415 RRVLLVSGGLQMITCQIIVAIILGVKFGA-----DQELSKGFSILVVVVICLFVVAFGWS 579
Query: 447 --TTTFIVPAELFPARFRSTCHGISGAVGKVGAIIGSVGFLWASHKEKEEGYPKGIGMKA 504
+ VP+E+FP RS GI+ AV + I + FL K
Sbjct: 580 WGPLGWTVPSEIFPLEIRSAGQGITVAVNLLFTFIIAQAFLALLCSFK---------FGI 732
Query: 505 SLIILGGVCIVGMFVTYFFTKETMGRSLEE 534
L G + I+ +FV Y F ET G +EE
Sbjct: 733 FLFFAGWITIMTIFV-YLFLPETKGIPIEE 819
>TC219982
Length = 590
Score = 35.8 bits (81), Expect = 0.051
Identities = 27/109 (24%), Positives = 43/109 (38%), Gaps = 13/109 (11%)
Frame = +2
Query: 450 FIVPAELFPARFRSTCHGISGAVGKVGAIIGSVGFLWASHKEKEEGYPKGIGMKASLIIL 509
++V +E++P R+R C G++ V +I + FL + IG + +I
Sbjct: 23 WVVNSEIYPLRYRGICGGMASTSNWVSNLIVAQSFL---------SLTQAIGTSWTFMIF 175
Query: 510 GGVCIVGMFVTYFFTKETMGRSLEENE-------------DEQSHHAEE 545
+ I + F ET G +EE E S HAEE
Sbjct: 176 IFITIAAIIFVIIFVPETKGLPMEEVEKMLEGRDLNFKFWQRSSRHAEE 322
>TC210054 homologue to UP|Q7XA52 (Q7XA52) Monosaccharide transporter, partial
(31%)
Length = 620
Score = 35.4 bits (80), Expect = 0.066
Identities = 34/107 (31%), Positives = 48/107 (44%), Gaps = 2/107 (1%)
Frame = +2
Query: 430 FMVIYGLAFFFANFGPNTTTFIVPAELFPARFRSTCHGISGAVGKVGA-IIGSVGFLWAS 488
F+ IY AF ++ +GP ++VP+E+FP RS I+ +V +I V
Sbjct: 122 FICIYVSAFAWS-WGP--LGWLVPSEIFPLEIRSAAQSINVSVNMFFTFLIAQVFLTMLC 292
Query: 489 HKEKEEGYPKGIGMKASLIILGGVCIVGM-FVTYFFTKETMGRSLEE 534
H MK L IL ++ M F YFF ET G +EE
Sbjct: 293 H------------MKFGLFILFAFFVLIMTFFIYFFLPETKGIPIEE 397
>TC213053 similar to UP|Q7XA50 (Q7XA50) Sorbitol-like transporter, partial
(27%)
Length = 440
Score = 34.3 bits (77), Expect = 0.15
Identities = 41/161 (25%), Positives = 65/161 (39%), Gaps = 9/161 (5%)
Frame = +1
Query: 339 VLFQSEIYKRYLNEKDDEDVYQE-AFHLARIQAILAVCSTIPGYFFTVYFIDRVGRVKIQ 397
VL+ EI+K+ E D E + A A+ IL + +DRVGR +
Sbjct: 1 VLYSPEIFKKAGLESDGEQLLATVAVGFAKTVFILVA----------TFLLDRVGRRPLL 150
Query: 398 MMGFFFMAVSFFALGFPY--------YSHWTKGENHDNKGFMVIYGLAFFFANFGPNTTT 449
+ M S LG W G + MV+ ++ F GP T
Sbjct: 151 LTSVGGMVFSLLTLGLSLTVIDHSRAVLKWAIGLSIG----MVLSYVSTFSVGAGP--IT 312
Query: 450 FIVPAELFPARFRSTCHGISGAVGKVGAIIGSVGFLWASHK 490
++ +E+FP R R+ + V +V + + S+ FL S+K
Sbjct: 313 WVYSSEIFPLRLRAQGAAMGVVVNRVTSGVISMTFLSLSNK 435
>AW351074 similar to GP|9294345|dbj organic anion transporter-like protein
{Arabidopsis thaliana}, partial (36%)
Length = 645
Score = 33.9 bits (76), Expect = 0.19
Identities = 29/116 (25%), Positives = 46/116 (39%), Gaps = 4/116 (3%)
Frame = -2
Query: 368 IQAILAVCSTIPGYFFTVYFIDRVGRVKIQMMGFFFMAVSFFALGFPYYSHWTKGENHDN 427
+ +L + +P + T +DR GR + + +F FF L G N
Sbjct: 512 LNVMLNSVAEMPAFTITAVLLDRFGRKPLTVATMWFSG--FFCL---------MGSLVGN 366
Query: 428 KGFM----VIYGLAFFFANFGPNTTTFIVPAELFPARFRSTCHGISGAVGKVGAII 479
G ++ G+ F G FI AELFP R+ G + ++GAI+
Sbjct: 365 VGVWKMVRMVCGVLGIFGMAGTYNLLFIYTAELFPTVVRNAALGCTTQASQMGAIL 198
>TC220535 similar to UP|HXT5_YEAST (P38695) Probable glucose transporter
HXT5, partial (3%)
Length = 1007
Score = 33.9 bits (76), Expect = 0.19
Identities = 20/84 (23%), Positives = 37/84 (43%)
Frame = +2
Query: 454 AELFPARFRSTCHGISGAVGKVGAIIGSVGFLWASHKEKEEGYPKGIGMKASLIILGGVC 513
+E++P R+R C G++ V +I + FL + IG ++ +I +
Sbjct: 623 SEIYPLRYRGICGGMASTSNWVSNLIVAQSFL---------SLTQAIGTSSTFMIFIFIT 775
Query: 514 IVGMFVTYFFTKETMGRSLEENED 537
+ + F ET G +EE E+
Sbjct: 776 VAAIVFVIIFVPETKGLPIEEVEN 847
Database: GMGI
Posted date: Oct 22, 2004 4:58 PM
Number of letters in database: 37,918,896
Number of sequences in database: 63,676
Lambda K H
0.342 0.150 0.499
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 26,538,792
Number of Sequences: 63676
Number of extensions: 426423
Number of successful extensions: 3865
Number of sequences better than 10.0: 49
Number of HSP's better than 10.0 without gapping: 3769
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 3845
length of query: 548
length of database: 12,639,632
effective HSP length: 102
effective length of query: 446
effective length of database: 6,144,680
effective search space: 2740527280
effective search space used: 2740527280
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 15 ( 7.4 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 39 (22.0 bits)
S2: 61 (28.1 bits)
Medicago: description of AC149576.4


