

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
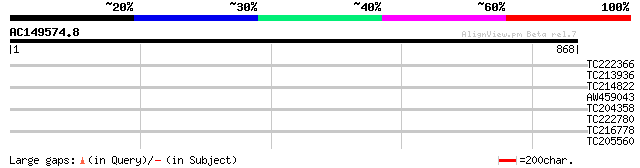
Query= AC149574.8 + phase: 0 /pseudo
(868 letters)
Database: GMGI
63,676 sequences; 37,918,896 total letters
Searching..................................................done
Score E
Sequences producing significant alignments: (bits) Value
TC222366 30 3.5
TC213936 similar to UP|Q9SEG2 (Q9SEG2) Protein kinase PK4, parti... 30 4.5
TC214822 UP|Q8S3F8 (Q8S3F8) S-adenosylmethionine decarboxylase, ... 30 5.9
AW459043 similar to GP|6164957|gb| vacuolar H+-ATP synthase 16kD... 29 7.8
TC204358 homologue to UP|Q8S3F8 (Q8S3F8) S-adenosylmethionine de... 29 7.8
TC222780 similar to UP|Q6L5C2 (Q6L5C2) 'unknow protein, contains... 29 7.8
TC216778 UP|O24431 (O24431) Calmodulin-like domain protein kinas... 29 7.8
TC205560 similar to GB|BAA97361.1|8843813|AB023042 Ca2+-transpor... 29 7.8
>TC222366
Length = 860
Score = 30.4 bits (67), Expect = 3.5
Identities = 24/85 (28%), Positives = 37/85 (43%), Gaps = 1/85 (1%)
Frame = -1
Query: 674 TKTLEWGQACTSLSSRYNGYGFVLS*STQTRINWLCRCRLF-IRSSSW*ITNRLFVYKWK 732
T+ +W S+ NG + +S S + I WL LF +S+ I + F YKW+
Sbjct: 848 TQAFDWKFTENSVHLTINGENYNISVSNRISIIWLELPSLFSYQSALVAILDDKFKYKWE 669
Query: 733 YNNFMEICETDNNSNIIKSRGTFSI 757
N + D N + G FS+
Sbjct: 668 EN*SNRTQKKDENQDQYTRLGYFSV 594
>TC213936 similar to UP|Q9SEG2 (Q9SEG2) Protein kinase PK4, partial (24%)
Length = 501
Score = 30.0 bits (66), Expect = 4.5
Identities = 19/49 (38%), Positives = 25/49 (50%), Gaps = 2/49 (4%)
Frame = +1
Query: 216 CSNKIECS-CWTISYCKRVSTTHEAWQT-NRFQR*KSSCKKRS*KKRRP 262
CS C CW +++ +R TT AW T R R*+ K RS* + P
Sbjct: 43 CSGSTSCEGCWELAHSRR-CTTRRAWTTRTRAWR*RP*AKTRS*TEASP 186
>TC214822 UP|Q8S3F8 (Q8S3F8) S-adenosylmethionine decarboxylase, complete
Length = 1826
Score = 29.6 bits (65), Expect = 5.9
Identities = 12/39 (30%), Positives = 21/39 (53%)
Frame = +2
Query: 92 CGVCSNFSTTTH*DGSSKKVGNICWI*ISIYYKVSRTHD 130
C + + FS H DG+SK C++ + Y + R+H+
Sbjct: 1406 CFLPTEFSVAVHVDGASKSFEQTCFLDVKGYCREERSHE 1522
>AW459043 similar to GP|6164957|gb| vacuolar H+-ATP synthase 16kDa
proteolipid subunit {Dendrobium crumenatum}, partial
(94%)
Length = 469
Score = 29.3 bits (64), Expect = 7.8
Identities = 13/43 (30%), Positives = 20/43 (46%), Gaps = 4/43 (9%)
Frame = -1
Query: 83 YFPSANIWMCGVCSNFSTTT----H*DGSSKKVGNICWI*ISI 121
Y P+ +W+ G+CSN T H G + K *+S+
Sbjct: 400 YHPTTTLWLLGICSNTGVATNANVHASGEASKTAREAAA*VSV 272
>TC204358 homologue to UP|Q8S3F8 (Q8S3F8) S-adenosylmethionine decarboxylase,
partial (61%)
Length = 953
Score = 29.3 bits (64), Expect = 7.8
Identities = 12/39 (30%), Positives = 21/39 (53%)
Frame = +3
Query: 92 CGVCSNFSTTTH*DGSSKKVGNICWI*ISIYYKVSRTHD 130
C + + FS H DG+SK C++ + Y + R+H+
Sbjct: 432 CFLPTEFSVAVHVDGASKLFDQTCFLDVKGYCREERSHE 548
>TC222780 similar to UP|Q6L5C2 (Q6L5C2) 'unknow protein, contains
tetratricopeptide (TPR) domain', partial (10%)
Length = 1034
Score = 29.3 bits (64), Expect = 7.8
Identities = 12/31 (38%), Positives = 16/31 (50%)
Frame = -3
Query: 199 CIH*SKKCD*IIHTSC*CSNKIECSCWTISY 229
C H ++K +H SC C EC CW +Y
Sbjct: 885 CSHLNQK----VHRSCCCC*DYECECWCANY 805
>TC216778 UP|O24431 (O24431) Calmodulin-like domain protein kinase isoenzyme
gamma , complete
Length = 2377
Score = 29.3 bits (64), Expect = 7.8
Identities = 18/50 (36%), Positives = 20/50 (40%)
Frame = -1
Query: 14 CSTCSYTKWTCRIIY*AYTVNCKTTSHEM*TPNFHLGTCNSACCNIDSHP 63
CS S KWTCR CK SH++ L S C SHP
Sbjct: 808 CSQVSSWKWTCRHTAIRSNSQCKDMSHQI-----ALYPTPSQVCRAASHP 674
>TC205560 similar to GB|BAA97361.1|8843813|AB023042 Ca2+-transporting
ATPase-like protein {Arabidopsis thaliana;} , partial
(27%)
Length = 1356
Score = 29.3 bits (64), Expect = 7.8
Identities = 15/30 (50%), Positives = 19/30 (63%), Gaps = 4/30 (13%)
Frame = -1
Query: 769 IHDSAHPKEL----WFILRKNGCNNNLRR* 794
IHD HPK+L W I R+N C+NN+ *
Sbjct: 390 IHDKIHPKKLHCI*WNITRRN-CSNNINH* 304
Database: GMGI
Posted date: Oct 22, 2004 4:58 PM
Number of letters in database: 37,918,896
Number of sequences in database: 63,676
Lambda K H
0.372 0.163 0.676
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 46,838,006
Number of Sequences: 63676
Number of extensions: 773411
Number of successful extensions: 9500
Number of sequences better than 10.0: 17
Number of HSP's better than 10.0 without gapping: 3784
Number of HSP's successfully gapped in prelim test: 324
Number of HSP's that attempted gapping in prelim test: 5522
Number of HSP's gapped (non-prelim): 4482
length of query: 868
length of database: 12,639,632
effective HSP length: 106
effective length of query: 762
effective length of database: 5,889,976
effective search space: 4488161712
effective search space used: 4488161712
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 14 ( 7.5 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 36 (22.0 bits)
S2: 63 (28.9 bits)
Medicago: description of AC149574.8


