

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC149493.3 - phase: 0 /pseudo
(473 letters)
Database: GMGI
63,676 sequences; 37,918,896 total letters
Searching..................................................done
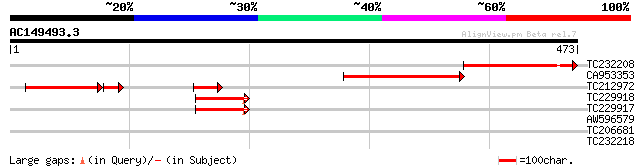
Score E
Sequences producing significant alignments: (bits) Value
TC232208 similar to UP|Q9S7F0 (Q9S7F0) F1K23.20 (F3M18.1), parti... 179 2e-45
CA953353 179 2e-45
TC212972 85 2e-19
TC229918 60 3e-09
TC229917 60 3e-09
AW596579 homologue to GP|9843649|emb SRPK4 {Arabidopsis thaliana... 29 5.2
TC206681 similar to UP|Q8H0P8 (Q8H0P8) RNA-binding protein, part... 28 6.8
TC232218 similar to GB|AAP13396.1|30023726|BT006288 At1g68330 {A... 28 6.8
>TC232208 similar to UP|Q9S7F0 (Q9S7F0) F1K23.20 (F3M18.1), partial (15%)
Length = 863
Score = 179 bits (455), Expect = 2e-45
Identities = 80/95 (84%), Positives = 88/95 (92%)
Frame = +1
Query: 379 QGDCTHTLVIRDMRLIHADDVQNWAVYPIVTFQLKIRFQKCGVCKIFRATKVTVDDKWTP 438
QGDCTHTLVIRDMRLIH +DV N AVYPI+TFQLK+RFQKC VCKIFRATKVTVDDKWTP
Sbjct: 502 QGDCTHTLVIRDMRLIHPEDVHNRAVYPIITFQLKLRFQKCNVCKIFRATKVTVDDKWTP 681
Query: 439 DNPCYFCDECFSLLHLAEDGGSPMYTDFIEYDYNH 473
+NPCYFCDECFSLLH A+D G+ +YTDF+EYDYNH
Sbjct: 682 ENPCYFCDECFSLLHQADD-GTLLYTDFVEYDYNH 783
>CA953353
Length = 422
Score = 179 bits (454), Expect = 2e-45
Identities = 84/101 (83%), Positives = 89/101 (87%)
Frame = +1
Query: 279 KKGQHDPSGYFLIEDVFYTDLRDPSAIDLTRPILDWLQNSKEEAQKKWEYIINGELQQKQ 338
K GQHDPSGYFLIEDVF DLRDPSAIDLTRPILDWL++SKEEAQKKWEYII GELQ+KQ
Sbjct: 13 KAGQHDPSGYFLIEDVFCPDLRDPSAIDLTRPILDWLRDSKEEAQKKWEYIITGELQKKQ 192
Query: 339 KAIVGEASVSHLPRFASFEMHKIHFCDLGFRLGAGYLYCHQ 379
KAI+GE S S LP F S EMHKI FCDL F+LGAGYLYCHQ
Sbjct: 193 KAIMGEKSASQLPHFRSIEMHKIRFCDLSFQLGAGYLYCHQ 315
>TC212972
Length = 808
Score = 85.1 bits (209), Expect(2) = 2e-19
Identities = 43/65 (66%), Positives = 51/65 (78%), Gaps = 1/65 (1%)
Frame = +2
Query: 14 DPSISIPRGGPIYVSNMSGPIVRVPLFQDSILTQLHSLQSELPPDSNH-DISVDDLKVFT 72
D SI IPRGGPIYV NM GP RVP FQ +L++L +L++EL D +H DISVDDLK+FT
Sbjct: 152 DLSIPIPRGGPIYVPNMVGPSTRVPHFQTCLLSELRNLEAELSEDFSHLDISVDDLKIFT 331
Query: 73 EDDLM 77
EDDLM
Sbjct: 332 EDDLM 346
Score = 42.0 bits (97), Expect = 6e-04
Identities = 19/24 (79%), Positives = 23/24 (95%)
Frame = +1
Query: 154 MLQSNCKEKVEEVVRIKQKQEEDK 177
+L+S+C EKVE+VVRIKQKQEEDK
Sbjct: 478 VLESDCIEKVEQVVRIKQKQEEDK 549
Score = 28.9 bits (63), Expect(2) = 2e-19
Identities = 11/17 (64%), Positives = 16/17 (93%)
Frame = +3
Query: 79 MALKQVFQGRDNNQDPP 95
MALK+VFQGR+N+++ P
Sbjct: 351 MALKEVFQGRENDENHP 401
>TC229918
Length = 770
Score = 59.7 bits (143), Expect = 3e-09
Identities = 33/48 (68%), Positives = 39/48 (80%), Gaps = 3/48 (6%)
Frame = +2
Query: 156 QSNCKEKVEEVVRIKQKQEEDKAQVRLHSFQLVFLIFI---HFIMCKI 200
QS+ EKVE+VVRIKQKQEEDKA V+LHSF+LV L F+ HF+ CKI
Sbjct: 377 QSDRIEKVEQVVRIKQKQEEDKASVKLHSFELVLLSFMWLFHFV-CKI 517
>TC229917
Length = 1013
Score = 59.7 bits (143), Expect = 3e-09
Identities = 33/48 (68%), Positives = 39/48 (80%), Gaps = 3/48 (6%)
Frame = +1
Query: 156 QSNCKEKVEEVVRIKQKQEEDKAQVRLHSFQLVFLIFI---HFIMCKI 200
QS+ EKVE+VVRIKQKQEEDKA V+LHSF+LV L F+ HF+ CKI
Sbjct: 634 QSDRIEKVEQVVRIKQKQEEDKASVKLHSFELVLLSFMWLFHFV-CKI 774
>AW596579 homologue to GP|9843649|emb SRPK4 {Arabidopsis thaliana}, partial
(22%)
Length = 362
Score = 28.9 bits (63), Expect = 5.2
Identities = 14/42 (33%), Positives = 20/42 (47%)
Frame = -2
Query: 215 LQDQSICQQINQNSKDDVS*VHKFRQKGQHTGSSGAHTCAGF 256
+ + I QQI + DD V K +H+G +G H C F
Sbjct: 262 MDEIKILQQIAEGDPDDKKCVVKLLDHFKHSGPNGQHVCMVF 137
>TC206681 similar to UP|Q8H0P8 (Q8H0P8) RNA-binding protein, partial (60%)
Length = 1222
Score = 28.5 bits (62), Expect = 6.8
Identities = 11/36 (30%), Positives = 19/36 (52%)
Frame = -3
Query: 374 YLYCHQGDCTHTLVIRDMRLIHADDVQNWAVYPIVT 409
+LYC GD H ++ + + VQ W ++ +VT
Sbjct: 611 FLYCQMGDLQHIKYLKHLGVEVQSQVQAWKLHLLVT 504
>TC232218 similar to GB|AAP13396.1|30023726|BT006288 At1g68330 {Arabidopsis
thaliana;} , partial (16%)
Length = 977
Score = 28.5 bits (62), Expect = 6.8
Identities = 15/47 (31%), Positives = 30/47 (62%), Gaps = 1/47 (2%)
Frame = -3
Query: 383 THTLVIRDMRLIHADDVQNWAVYPIVTF-QLKIRFQKCGVCKIFRAT 428
+ +L R + LIH DD+Q+ ++P+ + ++++RF+ C I R+T
Sbjct: 324 SQSLSPRGISLIHGDDLQHEPMHPLCSLDRVRMRFEG*FPCWISRST 184
Database: GMGI
Posted date: Oct 22, 2004 4:58 PM
Number of letters in database: 37,918,896
Number of sequences in database: 63,676
Lambda K H
0.334 0.146 0.467
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 22,383,333
Number of Sequences: 63676
Number of extensions: 342459
Number of successful extensions: 2520
Number of sequences better than 10.0: 16
Number of HSP's better than 10.0 without gapping: 2505
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 2518
length of query: 473
length of database: 12,639,632
effective HSP length: 101
effective length of query: 372
effective length of database: 6,208,356
effective search space: 2309508432
effective search space used: 2309508432
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 15 ( 7.2 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 39 (21.6 bits)
S2: 61 (28.1 bits)
Medicago: description of AC149493.3


