

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC149493.2 + phase: 0
(332 letters)
Database: GMGI
63,676 sequences; 37,918,896 total letters
Searching..................................................done
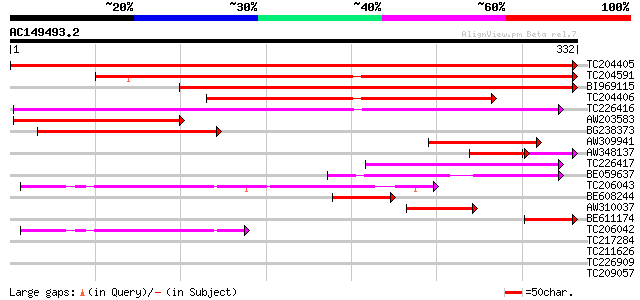
Score E
Sequences producing significant alignments: (bits) Value
TC204405 UP|Q6RIB6 (Q6RIB6) Cytosolic malate dehydrogenase , co... 630 0.0
TC204591 homologue to UP|Q6RIB6 (Q6RIB6) Cytosolic malate dehydr... 506 e-144
BI969115 similar to GP|10140741|gb cytoplasmic malate dehydrogen... 384 e-107
TC204406 homologue to UP|Q6RIB6 (Q6RIB6) Cytosolic malate dehydr... 319 1e-87
TC226416 homologue to UP|MDHP_MEDSA (O48902) Malate dehydrogenas... 236 8e-63
AW203583 similar to GP|10140741|gb cytoplasmic malate dehydrogen... 168 4e-42
BG238373 similar to GP|10334493|em cytosolic malate dehydrogenas... 157 6e-39
AW309941 similar to SP|O48905|MDHC_ Malate dehydrogenase cytopl... 99 3e-21
AW348137 homologue to GP|10798652|emb malate dehydrogenase {Nico... 67 6e-15
TC226417 similar to UP|MDHP_MEDSA (O48902) Malate dehydrogenase ... 77 1e-14
BE059637 homologue to SP|O48902|MDHP Malate dehydrogenase [NADP]... 75 5e-14
TC206043 homologue to UP|O81278 (O81278) Nodule-enhanced malate ... 67 1e-11
BE608244 similar to GP|10798652|emb malate dehydrogenase {Nicoti... 63 2e-10
AW310037 similar to GP|27462764|gb| malate dehydrogenase {Lupinu... 61 8e-10
BE611174 59 3e-09
TC206042 homologue to UP|O81278 (O81278) Nodule-enhanced malate ... 52 3e-07
TC217284 similar to UP|Q71G33 (Q71G33) COP8-like protein, partia... 29 2.6
TC211626 homologue to UP|Q9LH76 (Q9LH76) Arabidopsis thaliana ge... 28 5.8
TC226909 similar to PIR|H96764|H96764 protein RING zinc finger p... 28 5.8
TC209057 similar to UP|Q94AC4 (Q94AC4) At2g25730/F3N11.18, parti... 28 7.5
>TC204405 UP|Q6RIB6 (Q6RIB6) Cytosolic malate dehydrogenase , complete
Length = 1503
Score = 630 bits (1624), Expect = 0.0
Identities = 318/332 (95%), Positives = 326/332 (97%)
Frame = +2
Query: 1 MAKDPVRVLVTGAAGQIGYALVPMIARGVMLGPDQPVILHMLDIAPAAESLNGVKMELVD 60
MAKDPVRVLVTGAAGQIGYALVPMIARGVMLGPDQPVILHMLDI PAAESLNGVKMELVD
Sbjct: 101 MAKDPVRVLVTGAAGQIGYALVPMIARGVMLGPDQPVILHMLDIPPAAESLNGVKMELVD 280
Query: 61 AAFPLLKGVVATTDVVEACTGVNIAVMVGGFPRKEGMERKDVMSKNVSIYKSQASALEKH 120
AAFPLLKGVVATTDVVEACTGVNIAVMVGGFPRKEGMERKDVM+KNVSIYKSQASALEKH
Sbjct: 281 AAFPLLKGVVATTDVVEACTGVNIAVMVGGFPRKEGMERKDVMTKNVSIYKSQASALEKH 460
Query: 121 AAANCKVLVVANPANTNALILKEFAPSIPERNISCLTRLDHNRALGQISERLNVQVSDVK 180
AAANCKVLVVANPANTNALILKEFAPSIPE+NISCLTRLDHNRALGQISERL VQVSDVK
Sbjct: 461 AAANCKVLVVANPANTNALILKEFAPSIPEKNISCLTRLDHNRALGQISERLTVQVSDVK 640
Query: 181 NVIIWGNHSSTQYPDVNHATVNTPAGEKPVRELVSDDAWLNGEFISTVQQRGAAIIKARK 240
NVIIWGNHSSTQYPDVNHATV T AGEKPVREL++DDAWLNGEFI+TVQQRGAAIIKARK
Sbjct: 641 NVIIWGNHSSTQYPDVNHATVATSAGEKPVRELIADDAWLNGEFITTVQQRGAAIIKARK 820
Query: 241 LSSALSAASAACDHIRDWVLGTPQGTFVSMGVYSDGSYNVPSGLIYSFPVTCANGEWKIV 300
LSSALSAASAACDHIRDWVLGTPQGT+VSMGVYSDGSYNVP+GLIYSFPVTCANGEW IV
Sbjct: 821 LSSALSAASAACDHIRDWVLGTPQGTWVSMGVYSDGSYNVPAGLIYSFPVTCANGEWAIV 1000
Query: 301 QGLSIDEFSRKKLDLTAEELSEEKNLAYSCLS 332
QGLSIDEFSRKKLDLTAEELSEEK LAYSCL+
Sbjct: 1001QGLSIDEFSRKKLDLTAEELSEEKALAYSCLN 1096
>TC204591 homologue to UP|Q6RIB6 (Q6RIB6) Cytosolic malate dehydrogenase ,
partial (84%)
Length = 1604
Score = 506 bits (1303), Expect = e-144
Identities = 266/327 (81%), Positives = 275/327 (83%), Gaps = 45/327 (13%)
Frame = +2
Query: 51 LNGVKMELVDAAFPLLKG------------------------------------------ 68
LNGVKMELVDAAFPLLKG
Sbjct: 2 LNGVKMELVDAAFPLLKGLTYLC*RDCCINIMNNISNFDLTGLF**LI*SSRLLNPRILF 181
Query: 69 ---VVATTDVVEACTGVNIAVMVGGFPRKEGMERKDVMSKNVSIYKSQASALEKHAAANC 125
VVATTDVVEACTGVNIAVMVGGFPRKEGMERKDVM+KNVSIYKSQASALEKHAAANC
Sbjct: 182 ASGVVATTDVVEACTGVNIAVMVGGFPRKEGMERKDVMTKNVSIYKSQASALEKHAAANC 361
Query: 126 KVLVVANPANTNALILKEFAPSIPERNISCLTRLDHNRALGQISERLNVQVSDVKNVIIW 185
KVLVVANPANTNALILKEFAPSIPE+NISCLTRLDHNRALGQISERLN+QVSDVKNVIIW
Sbjct: 362 KVLVVANPANTNALILKEFAPSIPEKNISCLTRLDHNRALGQISERLNIQVSDVKNVIIW 541
Query: 186 GNHSSTQYPDVNHATVNTPAGEKPVRELVSDDAWLNGEFISTVQQRGAAIIKARKLSSAL 245
GNHSSTQYPDVNHATV GEKPVREL++DDAWLNGEFI+TVQQRGAAIIKARKLSSAL
Sbjct: 542 GNHSSTQYPDVNHATV----GEKPVRELIADDAWLNGEFITTVQQRGAAIIKARKLSSAL 709
Query: 246 SAASAACDHIRDWVLGTPQGTFVSMGVYSDGSYNVPSGLIYSFPVTCANGEWKIVQGLSI 305
SAASAACDHIRDWVLGTPQGT+VSMGVYSDGSYNVP+GLIYSFPVTCANGEW IVQGL I
Sbjct: 710 SAASAACDHIRDWVLGTPQGTWVSMGVYSDGSYNVPAGLIYSFPVTCANGEWTIVQGLPI 889
Query: 306 DEFSRKKLDLTAEELSEEKNLAYSCLS 332
DEFSRKKLDLTAEELSEEK LAYSCL+
Sbjct: 890 DEFSRKKLDLTAEELSEEKALAYSCLN 970
>BI969115 similar to GP|10140741|gb cytoplasmic malate dehydrogenase {Oryza
sativa} [Oryza sativa (japonica cultivar-group)],
partial (70%)
Length = 790
Score = 384 bits (987), Expect = e-107
Identities = 190/233 (81%), Positives = 208/233 (88%)
Frame = -1
Query: 100 KDVMSKNVSIYKSQASALEKHAAANCKVLVVANPANTNALILKEFAPSIPERNISCLTRL 159
KDVM VSI K+QASALE+HAA +CKVLVVANPANTNAL KE APSIPE+NI+CLTRL
Sbjct: 790 KDVMXXXVSIIKAQASALEQHAATDCKVLVVANPANTNALXXKEXAPSIPEKNITCLTRL 611
Query: 160 DHNRALGQISERLNVQVSDVKNVIIWGNHSSTQYPDVNHATVNTPAGEKPVRELVSDDAW 219
DHNRALGQISERLNV VSDVKNVI+WGNHSSTQYPDVNHATV T +GEKPVRELV DD W
Sbjct: 610 DHNRALGQISERLNVLVSDVKNVIVWGNHSSTQYPDVNHATVTTNSGEKPVRELVVDDNW 431
Query: 220 LNGEFISTVQQRGAAIIKARKLSSALSAASAACDHIRDWVLGTPQGTFVSMGVYSDGSYN 279
LN EFI+TVQQRGAAIIKARK SSALSAASAACDHIRDWVLGTP+G +VSMGVYSDGSY
Sbjct: 430 LNNEFITTVQQRGAAIIKARKQSSALSAASAACDHIRDWVLGTPKGEWVSMGVYSDGSYG 251
Query: 280 VPSGLIYSFPVTCANGEWKIVQGLSIDEFSRKKLDLTAEELSEEKNLAYSCLS 332
+P+GLIYSFPVTC G+W IVQGL ID+FSR+K+D TA+EL EEK LA SCL+
Sbjct: 250 IPTGLIYSFPVTCERGDWNIVQGLKIDQFSREKMDKTAQELIEEKTLAKSCLN 92
>TC204406 homologue to UP|Q6RIB6 (Q6RIB6) Cytosolic malate dehydrogenase ,
partial (50%)
Length = 498
Score = 319 bits (818), Expect = 1e-87
Identities = 159/170 (93%), Positives = 166/170 (97%)
Frame = +1
Query: 116 ALEKHAAANCKVLVVANPANTNALILKEFAPSIPERNISCLTRLDHNRALGQISERLNVQ 175
ALEKHAAANCKVLVVANPANTNALILKEFAPSIPE+NISCLTRLDHNRALGQISERLN+Q
Sbjct: 1 ALEKHAAANCKVLVVANPANTNALILKEFAPSIPEKNISCLTRLDHNRALGQISERLNIQ 180
Query: 176 VSDVKNVIIWGNHSSTQYPDVNHATVNTPAGEKPVRELVSDDAWLNGEFISTVQQRGAAI 235
VSDVKNVIIWGNHSSTQYPDVNHATV GEKPVREL++DDAWLNGEFI+TVQQRGAAI
Sbjct: 181 VSDVKNVIIWGNHSSTQYPDVNHATV----GEKPVRELIADDAWLNGEFITTVQQRGAAI 348
Query: 236 IKARKLSSALSAASAACDHIRDWVLGTPQGTFVSMGVYSDGSYNVPSGLI 285
IKARKLSSALSAASAACDHIRDWVLGTPQGT+VSMGVYSDGSYNVP+GLI
Sbjct: 349 IKARKLSSALSAASAACDHIRDWVLGTPQGTWVSMGVYSDGSYNVPAGLI 498
>TC226416 homologue to UP|MDHP_MEDSA (O48902) Malate dehydrogenase [NADP],
chloroplast precursor (NADP-MDH) , partial (96%)
Length = 1767
Score = 236 bits (603), Expect = 8e-63
Identities = 134/324 (41%), Positives = 193/324 (59%), Gaps = 2/324 (0%)
Frame = +3
Query: 3 KDPVRVLVTGAAGQIGYALVPMIARGVMLGPDQPVILHMLDIAPAAESLNGVKMELVDAA 62
K + + V+GAAG I L+ +A G + GPDQP+ L +L + ++L GV MEL D+
Sbjct: 450 KKLINIAVSGAAGMIANHLLFKLASGEVFGPDQPIALKLLGSERSIQALEGVAMELEDSL 629
Query: 63 FPLLKGVVATTDVVEACTGVNIAVMVGGFPRKEGMERKDVMSKNVSIYKSQASALEKHAA 122
FPLL+ V D E A+++G PR GMER D++ N IY +Q AL A+
Sbjct: 630 FPLLREVSIGIDPYEVFQDAEWALLIGAKPRGPGMERADLLDINGQIYAAQGRALNAVAS 809
Query: 123 ANCKVLVVANPANTNALILKEFAPSIPERNISCLTRLDHNRALGQISERLNVQVSDVKNV 182
N KV+VV NP NTNALI + AP+IP +N LTRLD NRA Q++ + V V NV
Sbjct: 810 RNVKVIVVGNPCNTNALICLKNAPNIPAKNFHALTRLDENRAKCQLALKAGVFYDKVSNV 989
Query: 183 IIWGNHSSTQYPDVNHATVNTPAGEKPVRELVSDDAWLNGEFISTVQQRGAAIIKARKLS 242
IWGNHS+TQ PD +A ++ PV+E+V D WL EF VQ+RG A+I+ S
Sbjct: 990 TIWGNHSTTQVPDFLNARID----GLPVKEVVKDQKWLEEEFTEKVQKRGGALIQKWGRS 1157
Query: 243 SALSAASAACDHIRDWVLGTPQGTFVSMGVYSDGS-YNVPSGLIYSFPV-TCANGEWKIV 300
SA S + + D IR V TP+G + S GVYSDG+ Y + G+++S P + +G++++V
Sbjct: 1158SAASTSVSIVDAIRSLVTPTPEGDWFSSGVYSDGNPYGIAEGIVFSMPCRSKGDGDYELV 1337
Query: 301 QGLSIDEFSRKKLDLTAEELSEEK 324
+ + D++ ++++ T EL EK
Sbjct: 1338KDVIFDDYLQQRIAKTEAELLAEK 1409
>AW203583 similar to GP|10140741|gb cytoplasmic malate dehydrogenase {Oryza
sativa} [Oryza sativa (japonica cultivar-group)],
partial (30%)
Length = 423
Score = 168 bits (425), Expect = 4e-42
Identities = 81/100 (81%), Positives = 90/100 (90%)
Frame = +1
Query: 3 KDPVRVLVTGAAGQIGYALVPMIARGVMLGPDQPVILHMLDIAPAAESLNGVKMELVDAA 62
K+P+ +LVTGAAGQIGYALVPMI RG MLGP+QP+ILHMLDI PA ESL G+KMEL+DAA
Sbjct: 124 KEPITILVTGAAGQIGYALVPMIVRGAMLGPNQPMILHMLDIEPATESLKGLKMELIDAA 303
Query: 63 FPLLKGVVATTDVVEACTGVNIAVMVGGFPRKEGMERKDV 102
+PLL+GVVATTDVVEAC VNI VMVGGFPR EGMERKDV
Sbjct: 304 YPLLRGVVATTDVVEACKDVNIVVMVGGFPRXEGMERKDV 423
>BG238373 similar to GP|10334493|em cytosolic malate dehydrogenase {Cicer
arietinum}, partial (31%)
Length = 325
Score = 157 bits (397), Expect = 6e-39
Identities = 82/108 (75%), Positives = 89/108 (81%)
Frame = +2
Query: 17 IGYALVPMIARGVMLGPDQPVILHMLDIAPAAESLNGVKMELVDAAFPLLKGVVATTDVV 76
+G +LV M A+ + +GPDQP ILHMLDI PA + LNGVKME DAAFPLLKGV ATTDV
Sbjct: 2 VGCSLVLMNAKRITMGPDQPCILHMLDIPPAEKPLNGVKME*KDAAFPLLKGVAATTDVA 181
Query: 77 EACTGVNIAVMVGGFPRKEGMERKDVMSKNVSIYKSQASALEKHAAAN 124
EACT VNIA MVGGFP KE MERKDVM+KNV IYK QASALEKHAAAN
Sbjct: 182 EACTEVNIAEMVGGFPTKERMERKDVMTKNV*IYKCQASALEKHAAAN 325
>AW309941 similar to SP|O48905|MDHC_ Malate dehydrogenase cytoplasmic (EC
1.1.1.37). [Alfalfa] {Medicago sativa}, partial (17%)
Length = 202
Score = 99.0 bits (245), Expect = 3e-21
Identities = 49/66 (74%), Positives = 53/66 (80%)
Frame = -1
Query: 246 SAASAACDHIRDWVLGTPQGTFVSMGVYSDGSYNVPSGLIYSFPVTCANGEWKIVQGLSI 305
SAASA CDHIR + +P T VSMGVYSDGS NVP+ LIYSFPVTCAN EW IVQGLSI
Sbjct: 202 SAASAVCDHIRYSAVVSPHSTCVSMGVYSDGSSNVPAALIYSFPVTCANTEWAIVQGLSI 23
Query: 306 DEFSRK 311
+E SRK
Sbjct: 22 NELSRK 5
>AW348137 homologue to GP|10798652|emb malate dehydrogenase {Nicotiana
tabacum}, partial (13%)
Length = 409
Score = 67.4 bits (163), Expect(2) = 6e-15
Identities = 30/35 (85%), Positives = 31/35 (87%)
Frame = -3
Query: 270 MGVYSDGSYNVPSGLIYSFPVTCANGEWKIVQGLS 304
MGVYSDGSYNVP+GLIYSFPVTCANGEW IV S
Sbjct: 407 MGVYSDGSYNVPAGLIYSFPVTCANGEWTIVSRTS 303
Score = 30.8 bits (68), Expect(2) = 6e-15
Identities = 16/33 (48%), Positives = 19/33 (57%), Gaps = 1/33 (3%)
Frame = -1
Query: 301 QGLSIDEFSRKKLDLTAEE-LSEEKNLAYSCLS 332
QGL I EFSRKKLD ++ YSCL+
Sbjct: 313 QGLPIXEFSRKKLDFDRRRAFLRRRHXXYSCLN 215
>TC226417 similar to UP|MDHP_MEDSA (O48902) Malate dehydrogenase [NADP],
chloroplast precursor (NADP-MDH) , partial (34%)
Length = 777
Score = 77.0 bits (188), Expect = 1e-14
Identities = 42/118 (35%), Positives = 69/118 (57%), Gaps = 2/118 (1%)
Frame = +2
Query: 209 PVRELVSDDAWLNGEFISTVQQRGAAIIKARKLSSALSAASAACDHIRDWVLGTPQGTFV 268
PV E+V D W EF V +RG A+I+ SS S + + D IR V TP+G +
Sbjct: 20 PVXEVVKDHKWXXEEFTEKVXKRGGALIQKWGRSSXASTSVSIVDAIRSLVTPTPEGDWF 199
Query: 269 SMGVYSDGS-YNVPSGLIYSFPV-TCANGEWKIVQGLSIDEFSRKKLDLTAEELSEEK 324
S GVYS+G+ Y + G+++S P + +G++++V+ + D++ R+++ T EL EK
Sbjct: 200 SSGVYSNGNPYGIAEGIVFSMPCRSKGDGDYELVKDVIFDDYLRQRIAKTEAELLAEK 373
>BE059637 homologue to SP|O48902|MDHP Malate dehydrogenase [NADP]
chloroplast precursor (EC 1.1.1.82) (NADP-MDH).
[Alfalfa], partial (31%)
Length = 419
Score = 74.7 bits (182), Expect = 5e-14
Identities = 47/140 (33%), Positives = 77/140 (54%), Gaps = 2/140 (1%)
Frame = +1
Query: 187 NHSSTQYPDVNHATVNTPAGEKPVRELVSDDAWLNGEFISTVQQRGAAIIKARKLSSALS 246
NHS+TQ PD +A ++ PV+E+V D WL EF VQ+RG A+I+ SSA S
Sbjct: 7 NHSTTQVPDFLNARIDG----LPVKEVVKDQKWLEEEFTEKVQKRGGALIQKWGRSSAAS 174
Query: 247 AASAACDHIRDWVLGTPQGTFVSMGVYSDGS-YNVPSGLIYSFPV-TCANGEWKIVQGLS 304
+ + D IR VYSDG+ Y + G+++S P + +G++++V+ +
Sbjct: 175 TSVSIVDAIRSL-------------VYSDGNPYGIAEGIVFSMPCRSKGDGDYELVKDVI 315
Query: 305 IDEFSRKKLDLTAEELSEEK 324
D++ ++++ T EL EK
Sbjct: 316 FDDYLQQRIAKTEAELLAEK 375
>TC206043 homologue to UP|O81278 (O81278) Nodule-enhanced malate
dehydrogenase , complete
Length = 1658
Score = 67.0 bits (162), Expect = 1e-11
Identities = 64/252 (25%), Positives = 117/252 (46%), Gaps = 7/252 (2%)
Frame = +2
Query: 7 RVLVTGAAGQIGYALVPMIARGVMLGPDQPVILHMLDIAPAAESLNGVKMELVDAAFPL- 65
+V V GAAG IG L +I ++ LH+ DIA ++ GV ++ P
Sbjct: 389 KVAVLGAAGGIGQPLALLIKMSPLISD-----LHLYDIA----NVKGVAADISHCNTPSQ 541
Query: 66 LKGVVATTDVVEACTGVNIAVMVGGFPRKEGMERKDVMSKNVSIYKSQASALEKHAAANC 125
++ +++ VN+ V+ G PRK GM R D+ + N I + SA+ + + +
Sbjct: 542 VRDFTGASELANCLKNVNVVVIPAGVPRKPGMTRDDLFNINAGIVRDLVSAVADN-SPDA 718
Query: 126 KVLVVANPANTN----ALILKEFAPSIPERNISCLTRLDHNRALGQISERLNVQVSDVKN 181
+ +++NP N+ A +LK+ P++ + +T LD RA +++R N+++ DV
Sbjct: 719 FIQIISNPVNSTVPIAAEVLKQKGVYDPKK-LFGVTTLDVVRANTFVAQRKNLKLIDVDV 895
Query: 182 VIIWGNHSSTQYPDVNHATVNTPAGEKPVRELVSDDAWLNGEFISTVQQRGAAII--KAR 239
++ G+ T P ++ + ++ + EL +Q G ++ KA
Sbjct: 896 PVVGGHAGITILPLLSKTRPSASFTDEEIDELT-----------VRIQNAGTEVVEAKAG 1042
Query: 240 KLSSALSAASAA 251
S+ LS A AA
Sbjct: 1043TGSATLSMAYAA 1078
>BE608244 similar to GP|10798652|emb malate dehydrogenase {Nicotiana
tabacum}, partial (10%)
Length = 112
Score = 62.8 bits (151), Expect = 2e-10
Identities = 28/37 (75%), Positives = 31/37 (83%)
Frame = +2
Query: 190 STQYPDVNHATVNTPAGEKPVRELVSDDAWLNGEFIS 226
ST YPDVNHATV T GEKP REL++DDAW+ GEFIS
Sbjct: 2 STHYPDVNHATVATSFGEKPFRELIADDAWVYGEFIS 112
>AW310037 similar to GP|27462764|gb| malate dehydrogenase {Lupinus albus},
partial (12%)
Length = 130
Score = 60.8 bits (146), Expect = 8e-10
Identities = 31/42 (73%), Positives = 36/42 (84%)
Frame = -1
Query: 233 AAIIKARKLSSALSAASAACDHIRDWVLGTPQGTFVSMGVYS 274
AAII+A LSSA+SAASAA HI DWV+GTPQGT+VS+ VYS
Sbjct: 127 AAIIRAIMLSSAISAASAASLHITDWVIGTPQGTWVSIRVYS 2
>BE611174
Length = 424
Score = 58.9 bits (141), Expect = 3e-09
Identities = 29/31 (93%), Positives = 30/31 (96%)
Frame = +3
Query: 302 GLSIDEFSRKKLDLTAEELSEEKNLAYSCLS 332
GLSIDEFSRKKLDLTAEELSEEK LAYSCL+
Sbjct: 174 GLSIDEFSRKKLDLTAEELSEEKALAYSCLN 266
>TC206042 homologue to UP|O81278 (O81278) Nodule-enhanced malate
dehydrogenase , partial (53%)
Length = 751
Score = 52.4 bits (124), Expect = 3e-07
Identities = 39/135 (28%), Positives = 64/135 (46%), Gaps = 1/135 (0%)
Frame = +2
Query: 7 RVLVTGAAGQIGYALVPMIARGVMLGPDQPVILHMLDIAPAAESLNGVKMELVDAAFPL- 65
+V V GAAG IG L +I ++ LH+ DIA ++ GV ++ P
Sbjct: 371 KVAVLGAAGGIGQPLSLLIKMSPLVSN-----LHLYDIA----NVKGVAADISHCNTPSQ 523
Query: 66 LKGVVATTDVVEACTGVNIAVMVGGFPRKEGMERKDVMSKNVSIYKSQASALEKHAAANC 125
++ +++ VN+ V+ G PRK GM R D+ + N I K SA+ + +
Sbjct: 524 VRDFTGASELPNCLKDVNVVVIPAGVPRKPGMTRDDLFNINAGIVKDLVSAVADY-CPDA 700
Query: 126 KVLVVANPANTNALI 140
V +++NP N+ I
Sbjct: 701 FVQIISNPVNSTVPI 745
>TC217284 similar to UP|Q71G33 (Q71G33) COP8-like protein, partial (66%)
Length = 1093
Score = 29.3 bits (64), Expect = 2.6
Identities = 17/52 (32%), Positives = 24/52 (45%)
Frame = -2
Query: 132 NPANTNALILKEFAPSIPERNISCLTRLDHNRALGQISERLNVQVSDVKNVI 183
N + LIL P + R +C + L H R + R +QVS V+ VI
Sbjct: 246 NDGSKYLLILLYLLPLVCNRRSACKSTLHHCRRSASVERRRQLQVSPVEVVI 91
>TC211626 homologue to UP|Q9LH76 (Q9LH76) Arabidopsis thaliana genomic DNA,
chromosome 3, BAC clone: T21E2 (AT3g14790/T21E2_4),
partial (17%)
Length = 440
Score = 28.1 bits (61), Expect = 5.8
Identities = 21/72 (29%), Positives = 32/72 (44%)
Frame = +1
Query: 5 PVRVLVTGAAGQIGYALVPMIARGVMLGPDQPVILHMLDIAPAAESLNGVKMELVDAAFP 64
P +L+TGAAG I + I R PD +I +LD +L + F
Sbjct: 118 PKNILITGAAGFIASHVCNRIVRNY---PDYKII--VLDKLDYCSNLKNLIPSRSSPNFK 282
Query: 65 LLKGVVATTDVV 76
+KG + + D+V
Sbjct: 283 FIKGDIGSADLV 318
>TC226909 similar to PIR|H96764|H96764 protein RING zinc finger protein
F25P22.18 [imported] - Arabidopsis thaliana {Arabidopsis
thaliana;} , partial (32%)
Length = 1522
Score = 28.1 bits (61), Expect = 5.8
Identities = 12/32 (37%), Positives = 21/32 (65%)
Frame = +1
Query: 184 IWGNHSSTQYPDVNHATVNTPAGEKPVRELVS 215
+W N S+ + P+ N T++TP +KP++ L S
Sbjct: 70 LWQNTSNGEDPETNSTTISTPF-QKPIQTLKS 162
>TC209057 similar to UP|Q94AC4 (Q94AC4) At2g25730/F3N11.18, partial (33%)
Length = 932
Score = 27.7 bits (60), Expect = 7.5
Identities = 14/37 (37%), Positives = 19/37 (50%)
Frame = +2
Query: 74 DVVEACTGVNIAVMVGGFPRKEGMERKDVMSKNVSIY 110
D + A + +NI M FPR E R V + N S+Y
Sbjct: 233 DSLSADSYLNILYMPSTFPRSERSRRSQVSANNNSVY 343
Database: GMGI
Posted date: Oct 22, 2004 4:58 PM
Number of letters in database: 37,918,896
Number of sequences in database: 63,676
Lambda K H
0.316 0.132 0.380
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 12,707,492
Number of Sequences: 63676
Number of extensions: 161297
Number of successful extensions: 661
Number of sequences better than 10.0: 41
Number of HSP's better than 10.0 without gapping: 656
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 657
length of query: 332
length of database: 12,639,632
effective HSP length: 98
effective length of query: 234
effective length of database: 6,399,384
effective search space: 1497455856
effective search space used: 1497455856
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 16 ( 7.3 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.6 bits)
S2: 59 (27.3 bits)
Medicago: description of AC149493.2


