

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC149493.1 + phase: 0
(254 letters)
Database: GMGI
63,676 sequences; 37,918,896 total letters
Searching..................................................done
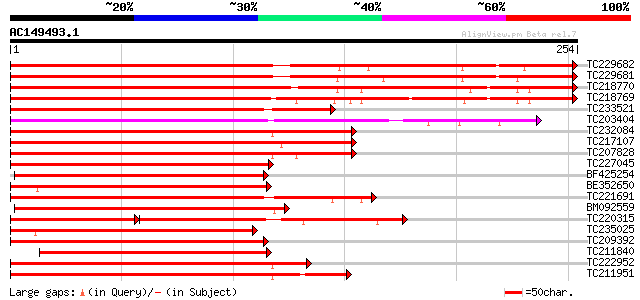
Score E
Sequences producing significant alignments: (bits) Value
TC229682 UP|Q9XIU9 (Q9XIU9) GmMYB29A2 protein, complete 332 1e-91
TC229681 UP|Q9S7E3 (Q9S7E3) GmMYB29A1 protein, complete 329 6e-91
TC218770 UP|Q9XIU5 (Q9XIU5) GmMYB29B2 protein, complete 317 3e-87
TC218769 UP|Q9XIU8 (Q9XIU8) GmMYB29A2 protein, complete 304 3e-83
TC233521 similar to UP|O04140 (O04140) Myb protein, partial (44%) 207 4e-54
TC203404 GmMYB16 194 2e-50
TC232084 UP|Q84PP2 (Q84PP2) Transcription factor MYB101 (Fragmen... 194 2e-50
TC217107 homologue to UP|Q7X9I1 (Q7X9I1) MYB transcription facto... 189 1e-48
TC207828 homologue to UP|Q6SL45 (Q6SL45) MYB6, partial (52%) 187 4e-48
TC227045 homologue to UP|Q43525 (Q43525) Transcription factor, p... 184 4e-47
BF425254 homologue to PIR|S26605|S26 myb-related protein 1 - gar... 182 2e-46
BE352650 181 4e-46
TC221691 GmMYB6 180 5e-46
BM092559 homologue to GP|13346186|gb GHMYB10 {Gossypium hirsutum... 179 1e-45
TC220315 similar to UP|Q8L5N8 (Q8L5N8) Myb-related transcription... 107 1e-44
TC235025 similar to UP|Q6R077 (Q6R077) MYB transcription factor,... 175 2e-44
TC209392 homologue to UP|Q8S3Y1 (Q8S3Y1) Typical P-type R2R3 Myb... 174 3e-44
TC211840 homologue to UP|Q8LBF0 (Q8LBF0) Myb-related protein M4,... 174 5e-44
TC222952 homologue to UP|Q40173 (Q40173) Myb-related transcripti... 171 2e-43
TC211951 similar to GB|AAS10022.1|41619094|AY519552 MYB transcri... 170 7e-43
>TC229682 UP|Q9XIU9 (Q9XIU9) GmMYB29A2 protein, complete
Length = 1249
Score = 332 bits (851), Expect = 1e-91
Identities = 179/272 (65%), Positives = 199/272 (72%), Gaps = 18/272 (6%)
Frame = +3
Query: 1 MVRAPCCEKKGLKKGPWTLEEDEILTSYINKHGHSNWRALPKDAGLLRCGKSCRLRWINY 60
MVRAPCCEK GLKKGPWT EED+IL SYI KHGH NWRALPK AGLLRCGKSCRLRWINY
Sbjct: 84 MVRAPCCEKMGLKKGPWTPEEDQILMSYIQKHGHGNWRALPKLAGLLRCGKSCRLRWINY 263
Query: 61 LRTDIKRGNFSNEEEQIIIKMHELLGNRWSAIAAKLPGRTDNEIKNMWHTHLKKRLLNTK 120
LR DIKRGNFS+EEE+IIIKMHELLGNRWSAIAAKLPGRTDNEIKN+WHTHLKKRLLN
Sbjct: 264 LRPDIKRGNFSSEEEEIIIKMHELLGNRWSAIAAKLPGRTDNEIKNVWHTHLKKRLLN-- 437
Query: 121 NNQPHSNTKKRVSKQKIKISDSNANS-NSMETTSSNCTFS----SDFSSQGKNLDNSIIC 175
S+ +KRVSK +IK SDSN+++ +E TSS CT S S FS KN+DN I
Sbjct: 438 -----SDIQKRVSKPRIKRSDSNSSTLTQLEPTSSACTTSLSDFSSFSEGTKNMDNMIKS 602
Query: 176 EDPEDSFVTMPQIDESFWSETITDDE------TNSMTISHELPIQEYPYNNSLENFQNPF 229
ED E MP IDESFWSE D E +NS TIS+EL +Y + NS+E+FQ
Sbjct: 603 EDIESVETIMPPIDESFWSEATVDYESSTMMTSNSWTISNELAPPQYQF-NSVESFQQQS 779
Query: 230 -------DDDDGMGFWVDLLIRSEESTELPEF 254
DD DGM FW D+ I+S ES ELPEF
Sbjct: 780 VDYNGSNDDHDGMDFWYDIFIKSGESIELPEF 875
>TC229681 UP|Q9S7E3 (Q9S7E3) GmMYB29A1 protein, complete
Length = 795
Score = 329 bits (844), Expect = 6e-91
Identities = 178/272 (65%), Positives = 198/272 (72%), Gaps = 18/272 (6%)
Frame = +1
Query: 1 MVRAPCCEKKGLKKGPWTLEEDEILTSYINKHGHSNWRALPKDAGLLRCGKSCRLRWINY 60
MVRAPCCEK GLKKGPWT EED+IL SYI KHGH NWRALPK AGLLRCGKSCRLRWINY
Sbjct: 1 MVRAPCCEKMGLKKGPWTPEEDQILMSYIQKHGHGNWRALPKLAGLLRCGKSCRLRWINY 180
Query: 61 LRTDIKRGNFSNEEEQIIIKMHELLGNRWSAIAAKLPGRTDNEIKNMWHTHLKKRLLNTK 120
LR DIKRGNFS+EEE+IIIKMHELLGNRWSAIAAKLPGRTDNEIKN+WHTHLKKRL+N
Sbjct: 181 LRPDIKRGNFSSEEEEIIIKMHELLGNRWSAIAAKLPGRTDNEIKNVWHTHLKKRLMN-- 354
Query: 121 NNQPHSNTKKRVSKQKIKISDSNAN--SNSMETTSSNCTFSSDFSSQG---KNLDNSIIC 175
S+T KRVSK +IK SDSN++ + S T+SS CT SSDFSS KN+DN I
Sbjct: 355 -----SDTNKRVSKPRIKRSDSNSSTLTQSEPTSSSGCTTSSDFSSFSEGTKNMDNMIKR 519
Query: 176 EDPEDSFVTMPQIDESFWSETITDDE-------TNSMTISHELPIQEYPYNNSLENFQ-- 226
ED E P IDESFW + D E +NS TIS+EL +Y + NS+E FQ
Sbjct: 520 EDIESMETVKPPIDESFWPQETVDYESSTMMQSSNSWTISNELAPPQYQF-NSVETFQQQ 696
Query: 227 ----NPFDDDDGMGFWVDLLIRSEESTELPEF 254
N DDGM FW D+ I+S ES ELPEF
Sbjct: 697 SVGYNDSKFDDGMDFWYDIFIKSGESIELPEF 792
>TC218770 UP|Q9XIU5 (Q9XIU5) GmMYB29B2 protein, complete
Length = 1316
Score = 317 bits (812), Expect = 3e-87
Identities = 169/276 (61%), Positives = 198/276 (71%), Gaps = 22/276 (7%)
Frame = +1
Query: 1 MVRAPCCEKKGLKKGPWTLEEDEILTSYINKHGHSNWRALPKDAGLLRCGKSCRLRWINY 60
MVRAPCCEK GLKKGPW EED+ILTSYI KHGH NWRALPK AGLLRCGKSCRLRWINY
Sbjct: 112 MVRAPCCEKMGLKKGPWATEEDQILTSYIQKHGHGNWRALPKQAGLLRCGKSCRLRWINY 291
Query: 61 LRTDIKRGNFSNEEEQIIIKMHELLGNRWSAIAAKLPGRTDNEIKNMWHTHLKKRLLNTK 120
LR DIKRGNF+ EEE+ IIK+HE+LGNRWSAIAAKLPGRTDNEIKN+WHT+LKKRLL +
Sbjct: 292 LRPDIKRGNFTIEEEETIIKLHEMLGNRWSAIAAKLPGRTDNEIKNVWHTNLKKRLLKSD 471
Query: 121 NNQPHSNTKKRVSKQKIKISDSNAN-----------SNSMETTSSNC-TFSSDFSSQGKN 168
++P S KR +K KIK SDSN++ M+TTS+ C T SSDFSS
Sbjct: 472 QSKPSS---KRATKPKIKRSDSNSSIITQSEPAHLRFREMDTTSTACNTSSSDFSSVTVG 642
Query: 169 LDNSIICEDPEDSFVTMPQIDESFWSETITDDETNSM------TISHELPIQEYPYNNSL 222
+II + +S TMP IDESFWSE DDET +M TIS+++P+Q YP+ N
Sbjct: 643 DSKNIIKSEDIESMETMPVIDESFWSEAAIDDETPTMSSQSLITISNDMPLQ-YPFANYE 819
Query: 223 ENFQ--NPFDD--DDGMGFWVDLLIRSEESTELPEF 254
E FQ + +D DDGM FW D+ R+ +S EL EF
Sbjct: 820 ETFQQSHAYDSNFDDGMDFWYDIFTRTNDSIELSEF 927
>TC218769 UP|Q9XIU8 (Q9XIU8) GmMYB29A2 protein, complete
Length = 828
Score = 304 bits (778), Expect = 3e-83
Identities = 172/279 (61%), Positives = 198/279 (70%), Gaps = 25/279 (8%)
Frame = +1
Query: 1 MVRAPCCEKKGLKKGPWTLEEDEILTSYINKHGHSNWRALPKDAGLLRCGKSCRLRWINY 60
MVRAPCCEK GLKKGPW EED+ILTSYI+KHGH NWRALPK AGLLRCGKSCRLRWINY
Sbjct: 1 MVRAPCCEKMGLKKGPWAPEEDQILTSYIDKHGHGNWRALPKQAGLLRCGKSCRLRWINY 180
Query: 61 LRTDIKRGNFSNEEEQIIIKMHELLGNRWSAIAAKLPGRTDNEIKNMWHTHLKKRLLNTK 120
LR DIKRGNF+ EEE+ IIK+H++LGNRWSAIAAKLPGRTDNEIKN+WHT+LKKRLL K
Sbjct: 181 LRPDIKRGNFTIEEEETIIKLHDMLGNRWSAIAAKLPGRTDNEIKNVWHTNLKKRLL--K 354
Query: 121 NNQPHSN-TKKRVSKQKIKISDSNA-----------NSNSMET-TSSNC-TFSSDFSSQG 166
++Q S + KR K KI+ SDSN+ N M+T TSS C T SSDFSS
Sbjct: 355 SDQSKSKPSSKRAIKPKIERSDSNSSIITQSEPDNFNFREMDTITSSACTTSSSDFSSVT 534
Query: 167 KNLDNSIICEDPEDSFVTMPQIDESFWSETITDDET------NSMTISHELPIQEYPYNN 220
+I ED E S TMP IDESFWSE DDET S+TIS+E+ +Q YP+ N
Sbjct: 535 VGDSKNIKSEDTE-STETMPVIDESFWSEAAIDDETPTMSSSQSLTISNEMRLQ-YPFAN 708
Query: 221 SLENFQ---NPFDD--DDGMGFWVDLLIRSEESTELPEF 254
E FQ + +D DDGM FW D+ R+ +S EL EF
Sbjct: 709 YEETFQQGHHAYDSNFDDGMDFWYDIFTRTNDSIELLEF 825
>TC233521 similar to UP|O04140 (O04140) Myb protein, partial (44%)
Length = 886
Score = 207 bits (527), Expect = 4e-54
Identities = 96/146 (65%), Positives = 112/146 (75%)
Frame = +2
Query: 1 MVRAPCCEKKGLKKGPWTLEEDEILTSYINKHGHSNWRALPKDAGLLRCGKSCRLRWINY 60
M R PCCE+ GLKKGPWT EED+IL S+I ++GH NWRALPK AGLLRCGKSCRLRWINY
Sbjct: 20 MTRTPCCERMGLKKGPWTAEEDQILVSHIQRYGHGNWRALPKQAGLLRCGKSCRLRWINY 199
Query: 61 LRTDIKRGNFSNEEEQIIIKMHELLGNRWSAIAAKLPGRTDNEIKNMWHTHLKKRLLNTK 120
LR DIKRG FS EEE I+K+H +LGNRWSAIAA LPGRTDNEIKN WHTHLK+ K
Sbjct: 200 LRPDIKRGKFSKEEEDTILKLHGILGNRWSAIAASLPGRTDNEIKNFWHTHLKR---IQK 370
Query: 121 NNQPHSNTKKRVSKQKIKISDSNANS 146
+ + N R+ ++ + SNA+S
Sbjct: 371 SGVHNGNPSSRILQEAQANTSSNASS 448
>TC203404 GmMYB16
Length = 1138
Score = 194 bits (494), Expect = 2e-50
Identities = 105/247 (42%), Positives = 145/247 (58%), Gaps = 9/247 (3%)
Frame = +1
Query: 1 MVRAPCCEKKGLKKGPWTLEEDEILTSYINKHGHSNWRALPKDAGLLRCGKSCRLRWINY 60
M RAPCC K GL KGPWT +ED +LT YI HG W++LPK AGLLRCGKSCRLRW+NY
Sbjct: 106 MGRAPCCSKVGLHKGPWTPKEDALLTKYIQAHGEGQWKSLPKKAGLLRCGKSCRLRWMNY 285
Query: 61 LRTDIKRGNFSNEEEQIIIKMHELLGNRWSAIAAKLPGRTDNEIKNMWHTHLKKRLLNTK 120
LR DIKRGN + EE+ +II+MH LLGNRWS IA +LPGRTDNEIKN W+THL K+L
Sbjct: 286 LRPDIKRGNIAPEEDDLIIRMHSLLGNRWSLIAGRLPGRTDNEIKNYWNTHLSKKL--KI 459
Query: 121 NNQPHSNTKKRVSKQKIKISDSNANSNSMETTSSNCTFSSDFSSQGKNLDNSIICEDPED 180
++T K + + + + N+N + N + ++GK D +P
Sbjct: 460 QGTEDTDTHKMLENPQEEAASDGGNNNKKKKKKKNGGKKNKQKNKGKEND------EPPK 621
Query: 181 SFVTMP---QIDESFWSETITDD---ETNSMTISHELPIQEYPY---NNSLENFQNPFDD 231
+ V +P ++ + T ++ ++NS + S +E P +N + N ++
Sbjct: 622 TQVYLPKPIRVKAMYLQRTDSNTFTFDSNSASGSTSQEKEESPVTKESNVVSEVGNVGEE 801
Query: 232 DDGMGFW 238
DG GF+
Sbjct: 802 SDGFGFF 822
>TC232084 UP|Q84PP2 (Q84PP2) Transcription factor MYB101 (Fragment), complete
Length = 1383
Score = 194 bits (494), Expect = 2e-50
Identities = 94/159 (59%), Positives = 113/159 (70%), Gaps = 4/159 (2%)
Frame = +2
Query: 1 MVRAPCCEKKGLKKGPWTLEEDEILTSYINKHGHSNWRALPKDAGLLRCGKSCRLRWINY 60
M R PC ++ GLKKGPWT EED IL YI KHGH +WRALPK A L RCGKSCRLRW NY
Sbjct: 53 MGRTPCSDENGLKKGPWTPEEDRILVDYIQKHGHGSWRALPKHARLNRCGKSCRLRWTNY 232
Query: 61 LRTDIKRGNFSNEEEQIIIKMHELLGNRWSAIAAKLPGRTDNEIKNMWHTHLKKRL---- 116
LR DIKRG FS EEEQ+II +H +LGN+WSAIA LPGRTDNEIKN W+THLKK+L
Sbjct: 233 LRPDIKRGKFSEEEEQLIINLHAVLGNKWSAIAGHLPGRTDNEIKNFWNTHLKKKLLQMG 412
Query: 117 LNTKNNQPHSNTKKRVSKQKIKISDSNANSNSMETTSSN 155
L+ ++P S+ +S + ++ +N SN T +N
Sbjct: 413 LDPVTHRPRSDHLNLLSNLQQLLAATNIVSNLTNTWDTN 529
>TC217107 homologue to UP|Q7X9I1 (Q7X9I1) MYB transcription factor R2R3 type,
partial (68%)
Length = 1058
Score = 189 bits (479), Expect = 1e-48
Identities = 88/156 (56%), Positives = 110/156 (70%), Gaps = 1/156 (0%)
Frame = +2
Query: 1 MVRAPCCEKKGLKKGPWTLEEDEILTSYINKHGHSNWRALPKDAGLLRCGKSCRLRWINY 60
M R+PCCEK KG WT EED+ L SYI HG WR+LPK AGLLRCGKSCRLRWINY
Sbjct: 266 MGRSPCCEKAHTNKGAWTKEEDDRLISYIRAHGEGCWRSLPKAAGLLRCGKSCRLRWINY 445
Query: 61 LRTDIKRGNFSNEEEQIIIKMHELLGNRWSAIAAKLPGRTDNEIKNMWHTHLKKRLLNTK 120
LR D+KRGNF+ EE+++IIK+H LLGN+WS IA +LPGRTDNEIKN W+TH++++LLN
Sbjct: 446 LRPDLKRGNFTEEEDELIIKLHSLLGNKWSLIAGRLPGRTDNEIKNYWNTHIRRKLLNRG 625
Query: 121 -NNQPHSNTKKRVSKQKIKISDSNANSNSMETTSSN 155
+ H + + + + S ET+SSN
Sbjct: 626 IDPATHRPLNEAATAATVTTNISFGKQEQQETSSSN 733
>TC207828 homologue to UP|Q6SL45 (Q6SL45) MYB6, partial (52%)
Length = 999
Score = 187 bits (475), Expect = 4e-48
Identities = 91/170 (53%), Positives = 116/170 (67%), Gaps = 15/170 (8%)
Frame = +2
Query: 1 MVRAPCCEKKGLKKGPWTLEEDEILTSYINKHGHSNWRALPKDAGLLRCGKSCRLRWINY 60
M R+PCCEK+ KG WT EEDE L +YI HG WR+LPK AGLLRCGKSCRLRWINY
Sbjct: 41 MGRSPCCEKEHTNKGAWTKEEDERLINYIKLHGEGCWRSLPKAAGLLRCGKSCRLRWINY 220
Query: 61 LRTDIKRGNFSNEEEQIIIKMHELLGNRWSAIAAKLPGRTDNEIKNMWHTHLKKRL---- 116
LR D+KRGNF+ EE+++II +H LLGN+WS IAA+LPGRTDNEIKN W+TH+K++L
Sbjct: 221 LRPDLKRGNFTEEEDELIINLHSLLGNKWSLIAARLPGRTDNEIKNYWNTHIKRKLYSRG 400
Query: 117 --------LNTKNNQPHSN---TKKRVSKQKIKISDSNANSNSMETTSSN 155
LN + P + T S K KI ++N N+N + +++
Sbjct: 401 IDPQTHRPLNAASATPATTVTATAVPSSNSKKKIKNNNNNNNGFQLVNNS 550
>TC227045 homologue to UP|Q43525 (Q43525) Transcription factor, partial (55%)
Length = 560
Score = 184 bits (466), Expect = 4e-47
Identities = 82/118 (69%), Positives = 96/118 (80%)
Frame = +3
Query: 1 MVRAPCCEKKGLKKGPWTLEEDEILTSYINKHGHSNWRALPKDAGLLRCGKSCRLRWINY 60
M R+PCCEK KG WT EED L SYI HG WR+LPK AGLLRCGKSCRLRWINY
Sbjct: 15 MGRSPCCEKAHTNKGAWTKEEDHRLISYIRAHGEGCWRSLPKAAGLLRCGKSCRLRWINY 194
Query: 61 LRTDIKRGNFSNEEEQIIIKMHELLGNRWSAIAAKLPGRTDNEIKNMWHTHLKKRLLN 118
LR D+KRGNFS EE+Q+IIK+H LLGN+WS IA +LPGRTDNEIKN W+TH++++LL+
Sbjct: 195 LRPDLKRGNFSLEEDQLIIKLHSLLGNKWSLIAGRLPGRTDNEIKNYWNTHIRRKLLS 368
>BF425254 homologue to PIR|S26605|S26 myb-related protein 1 - garden petunia,
partial (29%)
Length = 375
Score = 182 bits (461), Expect = 2e-46
Identities = 81/114 (71%), Positives = 93/114 (81%)
Frame = +3
Query: 3 RAPCCEKKGLKKGPWTLEEDEILTSYINKHGHSNWRALPKDAGLLRCGKSCRLRWINYLR 62
R+PCC+K GLKKGPWT EED+ L +YI +HGH +WRALP AGL RCGKSCRLRW NYLR
Sbjct: 3 RSPCCDKVGLKKGPWTPEEDQKLLAYIEEHGHGSWRALPAKAGLQRCGKSCRLRWTNYLR 182
Query: 63 TDIKRGNFSNEEEQIIIKMHELLGNRWSAIAAKLPGRTDNEIKNMWHTHLKKRL 116
DIKRG F +EEQ II++H LLGNRWS+I+ LP RTDNEIKN W+THLKK L
Sbjct: 183 PDIKRGKFXMQEEQTIIQLHALLGNRWSSISTHLPKRTDNEIKNYWNTHLKKXL 344
>BE352650
Length = 528
Score = 181 bits (458), Expect = 4e-46
Identities = 82/118 (69%), Positives = 95/118 (80%), Gaps = 1/118 (0%)
Frame = +1
Query: 1 MVRAPCCEKKG-LKKGPWTLEEDEILTSYINKHGHSNWRALPKDAGLLRCGKSCRLRWIN 59
M R+PCCE+ +KKGPWT EEDE L YI+KHGH +WR LPK AGL RCGKSCRLRW N
Sbjct: 103 MGRSPCCEESSSVKKGPWTPEEDEKLIDYISKHGHGSWRTLPKRAGLNRCGKSCRLRWTN 282
Query: 60 YLRTDIKRGNFSNEEEQIIIKMHELLGNRWSAIAAKLPGRTDNEIKNMWHTHLKKRLL 117
YLR DIKRG FS ++E+III H +LGN+WS IAA LPGRTDNEIKN W TH++K+LL
Sbjct: 283 YLRPDIKRGKFSEDDERIIINFHSVLGNKWSKIAAHLPGRTDNEIKNYWTTHIRKKLL 456
>TC221691 GmMYB6
Length = 590
Score = 180 bits (457), Expect = 5e-46
Identities = 91/173 (52%), Positives = 117/173 (67%), Gaps = 9/173 (5%)
Frame = +3
Query: 1 MVRAPCCEKKGLKKGPWTLEEDEILTSYINKHGHSNWRALPKDAGLLRCGKSCRLRWINY 60
MVR P C+K G +KG WT EED L +Y+ ++G NWR LP+ AGL RCGKSCRLRW+NY
Sbjct: 78 MVRTPSCDKSGTRKGTWTPEEDRKLIAYVTRYGSWNWRQLPRFAGLARCGKSCRLRWMNY 257
Query: 61 LRTDIKRGNFSNEEEQIIIKMHELLGNRWSAIAAKLPGRTDNEIKNMWHTHLKKRLLNTK 120
LR ++KRGNF+ +E++ II+MH+ LGN+WSAIAA+LPGRTDNEIKN WHT LKK
Sbjct: 258 LRPNVKRGNFTQQEDECIIRMHKKLGNKWSAIAAELPGRTDNEIKNHWHTTLKK----WS 425
Query: 121 NNQPHSNTKKRVSKQKIKISDSN------ANSNSMETTSSNC---TFSSDFSS 164
+N + R SK K K+ + ANS+ M SS+ + S+FSS
Sbjct: 426 QQNAITNEEARTSKSKDKVPNKGVTVTLPANSSLMSDNSSSSPVXSTCSEFSS 584
>BM092559 homologue to GP|13346186|gb GHMYB10 {Gossypium hirsutum}, partial
(38%)
Length = 421
Score = 179 bits (453), Expect = 1e-45
Identities = 81/131 (61%), Positives = 98/131 (73%), Gaps = 8/131 (6%)
Frame = +2
Query: 3 RAPCCEKKGLKKGPWTLEEDEILTSYINKHGHSNWRALPKDAGLLRCGKSCRLRWINYLR 62
R+PCC K+GL +G WT ED+IL YI HG WR LPK AGL RCGKSCRLRW+NYLR
Sbjct: 5 RSPCCSKEGLNRGAWTAHEDKILREYIRVHGEGRWRNLPKRAGLKRCGKSCRLRWLNYLR 184
Query: 63 TDIKRGNFSNEEEQIIIKMHELLGNRWSAIAAKLPGRTDNEIKNMWHTHLKKRLL----- 117
DIKRGN S +EE++II++H+LLGNRWS IA +LPGRTDNEIKN W+T+L K++
Sbjct: 185 PDIKRGNISPDEEELIIRLHKLLGNRWSLIAGRLPGRTDNEIKNYWNTNLGKKVKDGHQT 364
Query: 118 ---NTKNNQPH 125
NT+N PH
Sbjct: 365 TANNTQNPMPH 397
>TC220315 similar to UP|Q8L5N8 (Q8L5N8) Myb-related transcription factor
VlMYBB1-1, partial (45%)
Length = 654
Score = 107 bits (268), Expect(2) = 1e-44
Identities = 60/123 (48%), Positives = 81/123 (65%), Gaps = 3/123 (2%)
Frame = +2
Query: 59 NYLRTDIKRGNFSNEEEQIIIKMHELLGNRWSAIAAKLPGRTDNEIKNMWHTHLKKRLLN 118
NYLR D+KRGNF+ +EE+ II+MH+ LGNRWS IAA+LPGRTDNEIKN WHT LKKR
Sbjct: 254 NYLRPDVKRGNFTQQEEEFIIRMHKKLGNRWSTIAAELPGRTDNEIKNHWHTTLKKR--- 424
Query: 119 TKNNQPHSNTKK--RVSKQKIKISDSNANSNSMETTSSNCTFSSDFS-SQGKNLDNSIIC 175
+Q ++ TK+ R SK K K+ + + ++ TSS + +S S + + + S I
Sbjct: 425 ---SQQNTLTKEEARASKSKDKVLPNKGVTVTLPATSSQISDNSSLSPASSTSSEFSNIT 595
Query: 176 EDP 178
DP
Sbjct: 596 SDP 604
Score = 89.4 bits (220), Expect(2) = 1e-44
Identities = 37/58 (63%), Positives = 45/58 (76%)
Frame = +1
Query: 1 MVRAPCCEKKGLKKGPWTLEEDEILTSYINKHGHSNWRALPKDAGLLRCGKSCRLRWI 58
MVR P C+K G++KG WTLEED L +Y+ ++G NWR LP+ AGL RCGKSCRLRWI
Sbjct: 79 MVRTPSCDKSGMRKGTWTLEEDRKLIAYVTRYGSWNWRQLPRFAGLARCGKSCRLRWI 252
>TC235025 similar to UP|Q6R077 (Q6R077) MYB transcription factor, partial
(35%)
Length = 434
Score = 175 bits (443), Expect = 2e-44
Identities = 79/112 (70%), Positives = 90/112 (79%), Gaps = 1/112 (0%)
Frame = +1
Query: 1 MVRAPCCEKK-GLKKGPWTLEEDEILTSYINKHGHSNWRALPKDAGLLRCGKSCRLRWIN 59
M R+PCCE+ G+KKGPWT EEDE L YI+KHGH +WR LPK AGL RCGKSCRLRW N
Sbjct: 97 MGRSPCCEENVGVKKGPWTPEEDEKLIDYISKHGHGSWRTLPKRAGLNRCGKSCRLRWTN 276
Query: 60 YLRTDIKRGNFSNEEEQIIIKMHELLGNRWSAIAAKLPGRTDNEIKNMWHTH 111
YL DIKRG FS E+E+III +H +LGN+WS IA LPGRTDNEIKN W+TH
Sbjct: 277 YLTPDIKRGKFSEEDERIIINLHSVLGNKWSKIATHLPGRTDNEIKNYWNTH 432
>TC209392 homologue to UP|Q8S3Y1 (Q8S3Y1) Typical P-type R2R3 Myb protein
(Fragment), partial (59%)
Length = 866
Score = 174 bits (442), Expect = 3e-44
Identities = 77/116 (66%), Positives = 94/116 (80%)
Frame = +3
Query: 1 MVRAPCCEKKGLKKGPWTLEEDEILTSYINKHGHSNWRALPKDAGLLRCGKSCRLRWINY 60
M R PCC+K+G+KKGPWT EED IL SYI +HG NWRA+P GL RC KSCRLRW NY
Sbjct: 429 MGRPPCCDKEGVKKGPWTPEEDIILVSYIQEHGPGNWRAVPAKTGLSRCSKSCRLRWTNY 608
Query: 61 LRTDIKRGNFSNEEEQIIIKMHELLGNRWSAIAAKLPGRTDNEIKNMWHTHLKKRL 116
LR IKRGNF+ +EE++II + +LLGNRW+AIA+ LP RTDN+IKN W+THL+K+L
Sbjct: 609 LRPGIKRGNFTEQEEKMIIHLQDLLGNRWAAIASYLPQRTDNDIKNYWNTHLRKKL 776
>TC211840 homologue to UP|Q8LBF0 (Q8LBF0) Myb-related protein M4, partial
(38%)
Length = 516
Score = 174 bits (440), Expect = 5e-44
Identities = 76/104 (73%), Positives = 89/104 (85%)
Frame = +2
Query: 14 KGPWTLEEDEILTSYINKHGHSNWRALPKDAGLLRCGKSCRLRWINYLRTDIKRGNFSNE 73
KGPWT EED+ L YI KHG+ NWR LPK+AGL RCGKSCRLRW NYLR DIKRG FS E
Sbjct: 2 KGPWTTEEDQKLIDYIQKHGYGNWRTLPKNAGLQRCGKSCRLRWTNYLRPDIKRGRFSFE 181
Query: 74 EEQIIIKMHELLGNRWSAIAAKLPGRTDNEIKNMWHTHLKKRLL 117
EE+ II++H +LGN+WSAIA++LPGRTDNEIKN W+TH++KRLL
Sbjct: 182 EEETIIQLHSILGNKWSAIASRLPGRTDNEIKNYWNTHIRKRLL 313
>TC222952 homologue to UP|Q40173 (Q40173) Myb-related transcription factor,
partial (47%)
Length = 612
Score = 171 bits (434), Expect = 2e-43
Identities = 81/139 (58%), Positives = 101/139 (72%), Gaps = 4/139 (2%)
Frame = +2
Query: 1 MVRAPCCEKKGLKKGPWTLEEDEILTSYINKHGHSNWRALPKDAGLLRCGKSCRLRWINY 60
M R PCC+K GLKKGPWT EED+ L S+I +G WRA+PK AGLLRCGKSCRLRW NY
Sbjct: 125 MGRQPCCDKVGLKKGPWTAEEDKKLISFILTNGQCCWRAVPKLAGLLRCGKSCRLRWTNY 304
Query: 61 LRTDIKRGNFSNEEEQIIIKMHELLGNRWSAIAAKLPGRTDNEIKNMWHTHLKKRL---- 116
LR D+KRG S EE+++I +H LGNRWS IA+ LPGRTDNEIKN W+TH+KK+L
Sbjct: 305 LRPDLKRGLLSEYEEKMVIDLHAQLGNRWSKIASHLPGRTDNEIKNHWNTHIKKKLKKMG 484
Query: 117 LNTKNNQPHSNTKKRVSKQ 135
++ ++P N ++ Q
Sbjct: 485 IDPVTHKPLPNATEQTKNQ 541
>TC211951 similar to GB|AAS10022.1|41619094|AY519552 MYB transcription factor
{Arabidopsis thaliana;} , partial (38%)
Length = 776
Score = 170 bits (430), Expect = 7e-43
Identities = 86/157 (54%), Positives = 108/157 (68%), Gaps = 4/157 (2%)
Frame = +2
Query: 1 MVRAPCCEKKGLKKGPWTLEEDEILTSYINKHGHSNWRALPKDAGLLRCGKSCRLRWINY 60
M R CC K+ L+KG W+ EEDE L +YI KHGH W ++PK AGL RCGKSCRLRWINY
Sbjct: 92 MGRHSCCYKQKLRKGLWSPEEDEKLLNYITKHGHGCWSSVPKLAGLQRCGKSCRLRWINY 271
Query: 61 LRTDIKRGNFSNEEEQIIIKMHELLGNRWSAIAAKLPGRTDNEIKNMWHTHLKKRL---- 116
LR D+KRG FS +EE II++H +LGNRWS IAA+LPGRTDNEIKN+W++ LKK+L
Sbjct: 272 LRPDLKRGAFSQQEENSIIELHAVLGNRWSQIAAQLPGRTDNEIKNLWNSCLKKKLRQRG 451
Query: 117 LNTKNNQPHSNTKKRVSKQKIKISDSNANSNSMETTS 153
++ +QP S + K K +D + S E S
Sbjct: 452 IDPNTHQPLSEIEN--DKDKPLTADKSNQKASNEVIS 556
Database: GMGI
Posted date: Oct 22, 2004 4:58 PM
Number of letters in database: 37,918,896
Number of sequences in database: 63,676
Lambda K H
0.313 0.130 0.396
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 12,988,188
Number of Sequences: 63676
Number of extensions: 182815
Number of successful extensions: 1053
Number of sequences better than 10.0: 197
Number of HSP's better than 10.0 without gapping: 985
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 1005
length of query: 254
length of database: 12,639,632
effective HSP length: 95
effective length of query: 159
effective length of database: 6,590,412
effective search space: 1047875508
effective search space used: 1047875508
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 16 ( 7.2 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 42 (21.9 bits)
S2: 58 (26.9 bits)
Medicago: description of AC149493.1


