

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC149491.1 + phase: 0 /pseudo
(1441 letters)
Database: GMGI
63,676 sequences; 37,918,896 total letters
Searching..................................................done
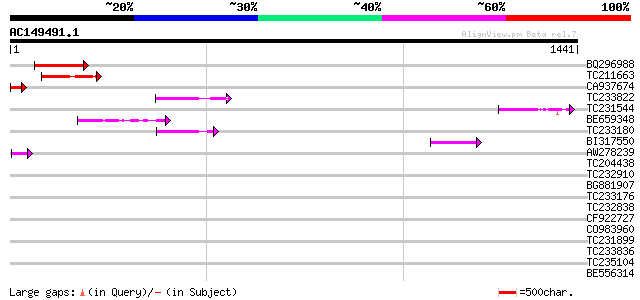
Score E
Sequences producing significant alignments: (bits) Value
BQ296988 similar to GP|21740616|em OSJNBb0089K24.12 {Oryza sativ... 150 4e-36
TC211663 121 3e-27
CA937674 similar to GP|28172880|emb xylosyltransferase I {Mus mu... 61 4e-09
TC233822 58 3e-08
TC231544 weakly similar to UP|Q9FLR2 (Q9FLR2) Polyprotein-like, ... 55 3e-07
BE659348 weakly similar to PIR|JC7809|JC78 sulfakinin receptor p... 51 4e-06
TC233180 similar to UP|Q944K0 (Q944K0) At1g18030/T10F20_3, parti... 50 5e-06
BI317550 weakly similar to GP|9759590|dbj| polyprotein-like {Ara... 50 7e-06
AW278239 49 2e-05
TC204438 homologue to UP|Q84VH6 (Q84VH6) Gag-pol polyprotein, co... 42 0.002
TC232910 similar to UP|Q6I8N0 (Q6I8N0) Pol polypeptide, partial ... 33 0.003
BG881907 homologue to GP|21432|emb|CA ORF2 {Solanum tuberosum}, ... 41 0.004
TC233176 weakly similar to GB|AAL07033.1|15810291|AY056184 auxin... 40 0.006
TC232838 similar to GB|AAP21377.1|30102918|BT006569 At1g47970 {A... 40 0.007
CF922727 40 0.010
CO983960 39 0.017
TC231899 similar to UP|Q850H7 (Q850H7) Gag-pol polyprotein (Frag... 39 0.017
TC233836 39 0.017
TC235104 weakly similar to UP|Q850H8 (Q850H8) Gag-pol polyprotei... 39 0.022
BE556314 similar to GP|29150404|gb gag-pol polyprotein {Oryza sa... 39 0.022
>BQ296988 similar to GP|21740616|em OSJNBb0089K24.12 {Oryza sativa (japonica
cultivar-group)}, partial (1%)
Length = 408
Score = 150 bits (379), Expect = 4e-36
Identities = 70/136 (51%), Positives = 99/136 (72%)
Frame = -1
Query: 64 RCNNLILSWIINSVSPQIAQTIVFHEYAIDVWIELQERFSKVDRIRVASLRSSINNLKQG 123
RCN LI SWI+NSV P I+++IVF + A DVW++L+ERFS+ D +RV+ ++ I L QG
Sbjct: 408 RCNMLIHSWILNSVEPSISRSIVFMDNASDVWLDLKERFSQGDLVRVSEIQQEIYALTQG 229
Query: 124 DKSVLDYFTSIKSLWEELNSHRPMPMCTCPYPCRCESMRAARDFRMEDQVIQFLTGLNDS 183
+SV +++ K+LWEEL + P+P CTC + C C++MR AR V++FLTGLND
Sbjct: 228 TRSVTTFYSDKKALWEELEIYMPIPNCTCHHRCSCDAMRLARRHHHTLHVMRFLTGLNDE 49
Query: 184 FSVVKTQVLLIDPLPS 199
F+ VK+Q+LLI+PLPS
Sbjct: 48 FNAVKSQILLIEPLPS 1
>TC211663
Length = 426
Score = 121 bits (303), Expect = 3e-27
Identities = 62/153 (40%), Positives = 99/153 (64%)
Frame = -3
Query: 81 IAQTIVFHEYAIDVWIELQERFSKVDRIRVASLRSSINNLKQGDKSVLDYFTSIKSLWEE 140
IAQ +++ ++A ++W +L+E FS+ D +++A L+ I LKQG +VLD+FT +K +WEE
Sbjct: 424 IAQIVIYFDHATNIWNDLKEGFSQGDLLQIAELQEEIYRLKQGSHTVLDFFTKLKFVWEE 245
Query: 141 LNSHRPMPMCTCPYPCRCESMRAARDFRMEDQVIQFLTGLNDSFSVVKTQVLLIDPLPSI 200
L+++ M +CTCP +R + +D VI FL GL++ FSVV ++VLL+D LPS
Sbjct: 244 LDNYGLMNLCTCP----------SRTYHQQDFVIHFLKGLDERFSVVCSEVLLMDHLPST 95
Query: 201 NKVYSMVIQEESNIIPPTSLASNEDSSILVNAS 233
+++SMVIQ E+ S S ED + +N +
Sbjct: 94 KRIFSMVIQHETQ---HASHTSAEDQNRFINVA 5
>CA937674 similar to GP|28172880|emb xylosyltransferase I {Mus musculus},
partial (2%)
Length = 420
Score = 60.8 bits (146), Expect = 4e-09
Identities = 27/42 (64%), Positives = 34/42 (80%)
Frame = -1
Query: 1 SIYYVHPSEGPNSVTVTPLLTGPNYLAWNRSMKRALGTKNKF 42
S ++VHPS+GP+SV VTPLL G NY +W RS++RALG K KF
Sbjct: 129 SPFFVHPSDGPSSVKVTPLLDGSNYHSWARSLRRALGAKLKF 4
>TC233822
Length = 632
Score = 58.2 bits (139), Expect = 3e-08
Identities = 49/193 (25%), Positives = 86/193 (44%), Gaps = 1/193 (0%)
Frame = +2
Query: 371 DHWLIDSGANEHICSSLHLFHSYYRIK-PICVNLPNGSSVIVQYAGTVVFSPHFHITHVL 429
D +IDS A++HI + LF S K P + L NG+ V + G P ++ +
Sbjct: 29 DPRVIDSTASDHIFGNSSLFTSQSPPKIPHLITLANGTKVTSKGFGKFSIFPSLNLDPIH 208
Query: 430 YSPSFKVYLISVSKICQSLPYHVHFLLNTYVIQDVKTQKMIGLGNLCDGLYRLHPFAPAS 489
+ P+ L+S+S++ ++L + + F +++VIQ+ T +IG+ + GLY L
Sbjct: 209 FVPNCPFNLVSLSQLTKALNFSITFDADSFVIQERDTSWLIGVEHESRGLYYL------- 367
Query: 490 PQAHFISSAVSPTNKMSCNSVSSNNNVSSIPSNAIWHFRLGHLSNQRLSMMHSLYSSITI 549
+SS+ + + PS + H LGH +L M+ SL +
Sbjct: 368 -------------------EISSSMSCFATPSPKLLHNHLGHPHLLKLKMVPSLNKLQVL 490
Query: 550 DNKAVCDICHFAK 562
+ C+ C K
Sbjct: 491 E----CESCQLGK 517
>TC231544 weakly similar to UP|Q9FLR2 (Q9FLR2) Polyprotein-like, partial (16%)
Length = 662
Score = 54.7 bits (130), Expect = 3e-07
Identities = 63/204 (30%), Positives = 105/204 (50%), Gaps = 11/204 (5%)
Frame = +2
Query: 1243 NSNSLSCYS*SSKIPKRFTRQRNFLPTRFRITHSGIH*CRLGRL*RYKAFYLRTLFLSWT 1302
++++L CYS S+IP+R +++R R ++S I *CRLG + +K L LSW
Sbjct: 2 HASALCCYS-CSQIPQRMSKKRALF**RISHSNSWIQ*CRLGNMY*FKQVNHMVLLLSW* 178
Query: 1303 ISYMLENQETTYSFQ--ILF*S*V*SNGICYL*NAMVIILAQRFTSSMCSTSCALL**SK 1360
S ++E+ ET + F ++ *S*V*S+ + ++ + + ++R S C +L* SK
Sbjct: 179 FSNLMES*ETKHCFPLILIL*S*V*SSHLHHMRITVAYLPSKR---SPCYFD--IL*QSK 343
Query: 1361 CYAHCIKSCFS*KN*TS*N*LSHCKGKATSR---------HLQTPSSYHS*SNW*FFYKG 1411
C+A I + S + T +*L HC + ++R LQ+ S + *S+
Sbjct: 344 CFA--IAANQSNISWTIGD*LPHCSREDSTRIDALFTPCFLLQSVSRHLH*SS------- 496
Query: 1412 FISSTF*PLAFQVGDVRHLPPSNL 1435
++ TF Q + HLP S+L
Sbjct: 497 -LAQTFQQQLIQAWAI*HLPTSSL 565
>BE659348 weakly similar to PIR|JC7809|JC78 sulfakinin receptor protein
DSK-R1 - fruit fly (Drosophila melanogaster), partial
(6%)
Length = 770
Score = 50.8 bits (120), Expect = 4e-06
Identities = 57/238 (23%), Positives = 100/238 (41%), Gaps = 1/238 (0%)
Frame = -1
Query: 173 VIQFLTGLNDSFSVVKTQVLLIDPLPSINKVYSMVIQEESNIIPPTSLASNEDSSILVNA 232
++ L G++ F ++ QVL +PS+ + + +++ S I S+ N ++S++V+
Sbjct: 623 MVLVLRGMHPDFDHIRDQVLTSQEVPSLENLITRLLRVPSPKIGGNSV-DNIETSVMVSN 447
Query: 233 SDARKPFLRGKSSGTSQSKNNSRYCTFCRRNNHTVEYCYLKHDFPNANKPTASSNAVTSE 292
+ RG G + + C++C+R HT + CY H FP S N SE
Sbjct: 446 RGGQGG--RGNQGGRA-GRGGRP*CSYCKRVGHTQDTCYSIHGFPG-----KSVNISKSE 291
Query: 293 HAVDSHTSSEGTSSSSQTGLTQEQYVHLVSLLQQSSLVPSATPPNPASTNHVATSFPSSI 352
TS ++Y+ L + + + + + H +T+ S +
Sbjct: 290 -----------TSEIKFLEADYQEYLQLKATKESQT--------SSVISGHNSTACISQV 168
Query: 353 DFTSGINTIFSCSLHVPSDHWLIDSGANEHICSSLHLFHSYYRIK-PICVNLPNGSSV 409
G N W+IDSGA++HI S+ LF K P + L +GS V
Sbjct: 167 ----GNN----------QSPWIIDSGASDHIASNSSLF*FLSPPKIPHFITLADGSRV 36
>TC233180 similar to UP|Q944K0 (Q944K0) At1g18030/T10F20_3, partial (11%)
Length = 916
Score = 50.4 bits (119), Expect = 5e-06
Identities = 47/166 (28%), Positives = 70/166 (41%), Gaps = 8/166 (4%)
Frame = +2
Query: 374 LIDSGANEHICSSLHLFHSY-YRIKPICVNLPNGSSVIVQYAGTVVFSPHFHITHVLYSP 432
+IDSG +H+ F SY + I + + NGS + + G + H+ +VLY P
Sbjct: 455 IIDSGVTDHMTPHSSYFSSYTFLIGNQHIIVANGSHIPIIGCGNIQLQSSLHLNNVLYVP 634
Query: 433 SFKVYLISVSKICQSLPYHVHFLLNTYVIQDVKTQKMIGLGNLCDGLYRLHPFAPASPQA 492
L+S+ KI L V F + V D+ +MIG+ GLY L
Sbjct: 635 KLSNNLLSIHKIT*DLNCVVTFFHSHCVF*DLAMGRMIGIAKE*GGLYYL---------- 784
Query: 493 HFISSAVSPTNKMSC---NSVSSNNNVSSIP--SNAIW--HFRLGH 531
+ C +++SN+ SS P S+ IW H LGH
Sbjct: 785 -------QHEDNKECTR*KALTSNHQTSSEPWSSSQIWLQHKCLGH 901
>BI317550 weakly similar to GP|9759590|dbj| polyprotein-like {Arabidopsis
thaliana}, partial (18%)
Length = 421
Score = 50.1 bits (118), Expect = 7e-06
Identities = 44/130 (33%), Positives = 64/130 (48%)
Frame = -1
Query: 1069 LIWSQTSQ*KVV*KADHSPYFQ*L*TSCI*CFFVYQINF*DFHYYFSLC**YNFSWQFSH 1128
L+W + S+*+VV*KAD + L T +Y FH +C *++ F+
Sbjct: 415 LVWVEASK*EVV*KAD*FASQRRLHTINFRLLPIYSHQRKYFHCTVGICR*HHSCR*FN* 236
Query: 1129 *ISYHQECFTSGIQNQGSWHFEIFSWP*SSSFTQWDFSMPKKILFGSVE*FRPSWF*ACF 1188
*I QEC QN S EI SW *S SF + + ++IL GS FR +W C
Sbjct: 235 *I*QDQECS*FSFQN*KSG*VEILSWA*SCSF*TGYYHLTEEILSGSA*GFRITWVQTC* 56
Query: 1189 YTL*SLYQTS 1198
+T * +++ +
Sbjct: 55 HTS*HIHKVA 26
>AW278239
Length = 417
Score = 48.9 bits (115), Expect = 2e-05
Identities = 21/53 (39%), Positives = 31/53 (57%)
Frame = +3
Query: 4 YVHPSEGPNSVTVTPLLTGPNYLAWNRSMKRALGTKNKFVFIDGSVPIPPMDD 56
++H S+GP + + L NY W+ +M A KNK FIDGS+P+P + D
Sbjct: 252 FLHHSDGPGLILTSQPLDHKNYTTWSLAMMVAFSVKNKVAFIDGSLPMPTIAD 410
>TC204438 homologue to UP|Q84VH6 (Q84VH6) Gag-pol polyprotein, complete
Length = 4734
Score = 42.0 bits (97), Expect = 0.002
Identities = 67/346 (19%), Positives = 113/346 (32%), Gaps = 8/346 (2%)
Frame = +1
Query: 231 NASDARKPFLRGKSSGTSQSKNNSRY--CTFCRRNNHTVEYCYLKHDFPNANKPTASSNA 288
N++ A R + GT Q K+ + C +C + H +CY H P+
Sbjct: 1429 NSTGATMSQHRSRHHGTQQKKSKRKKWRCHYCGKYGHIKPFCYHLHGHPH---------- 1578
Query: 289 VTSEHAVDSHTSSEGTSSSSQTGLTQEQYVH-LVSLLQQSSLVPSATPPNPASTNHVATS 347
GT SSS H +VSL+ +SL SA
Sbjct: 1579 -------------HGTQSSSSGRKMMWVPKHKIVSLVVHTSLRASA-------------- 1677
Query: 348 FPSSIDFTSGINTIFSCSLHVPSDHWLIDSGANEHICSSLHLFHSYYRIKPICVNLPNGS 407
+ W +DSG + H+ + V +GS
Sbjct: 1678 ----------------------KEDWYLDSGCSRHMTGVKEFLVNIEPCSTSYVTFGDGS 1791
Query: 408 SVIVQYAGTVVFSPHFHITHVLYSPSFKVYLISVSKICQSLPYHVHFLLNTYVIQDVKTQ 467
+ G +V + VL LIS+S++C ++V+F + ++ + K++
Sbjct: 1792 KGKITGMGKLVHDGLPSLNKVLLVKGLTANLISISQLCDE-GFNVNFTKSECLVTNEKSE 1968
Query: 468 KMIGLGNLCDGLYRLHPFAPASPQAHFISSAVSPTNKMSCNSVSSNNNVSSIPSNAIWHF 527
++ D Y P T+ S S + V IWH
Sbjct: 1969 VLMKGSRSKDNCYLWTP---------------QETSYSSTCLFSKEDEVK------IWHQ 2085
Query: 528 RLGHLSNQRLSMMHSL-----YSSITIDNKAVCDICHFAKQRKLPY 568
R GHL + + + ++ I+ +C C KQ K+ +
Sbjct: 2086 RFGHLHLRGMKKIIDKGAVRGIPNLKIEEGRICGECQIGKQVKMSH 2223
Score = 35.8 bits (81), Expect = 0.14
Identities = 48/184 (26%), Positives = 77/184 (41%), Gaps = 11/184 (5%)
Frame = +3
Query: 1210 SL*KTHWQIDLLEYNQT*HYFHYPAIESVS*QTNSNSLSCYS*SSKIPKRFTRQRNFLPT 1269
S+ K W++ + QT*H+ + +S Q+ SL +S+I K +++ +
Sbjct: 4065 SVQKHDWELTIFNS*QT*HHLCSRCLCKISSQS*DKSLESSKENSEICKWHQ*LWDYVLS 4244
Query: 1270 RFRITHSGIH*CRLGRL*RYKAFYLRTLFLSWTISYMLENQETTYSFQILF*S*V*SNGI 1329
FR + *C LG R + + +FL SY + QE I
Sbjct: 4245 LFRFNAGWVL*C*LGWKCR*QKKHFWWMFLFGNQSYFMVQQEAELCVPI----------- 4391
Query: 1330 CYL*NAMVIILAQRFTSSM---------CSTSC--ALL**SKCYAHCIKSCFS*KN*TS* 1378
Y *+ + Q FT+S+ C T C +L* +CY + KSC + +N *
Sbjct: 4392 -YC*SRVYCSRKQLFTTSLDEADAEGVQCRTRCHDIVL*QHECY*YF*KSCSTQQNQAH* 4568
Query: 1379 N*LS 1382
+* S
Sbjct: 4569 H*TS 4580
>TC232910 similar to UP|Q6I8N0 (Q6I8N0) Pol polypeptide, partial (3%)
Length = 690
Score = 33.1 bits (74), Expect(2) = 0.003
Identities = 32/136 (23%), Positives = 57/136 (41%), Gaps = 1/136 (0%)
Frame = +2
Query: 428 VLYSPSFKVYLISVSKICQSLPYHVHFLLNTYVIQDVKTQKMIGLGNLCDGLYRLHPFAP 487
VL+ P + L+S I + + + Y++ + +G G C+ +++L+ P
Sbjct: 332 VLFVPGIRKNLLS-GMILNNCGFKQVLESDKYILS--RHGSFVGFGYRCNEMFKLNIDVP 502
Query: 488 ASPQAHFISSAVSPTNKMSCNSVSSNNNVSSIPSNAIWHFRLGHLSNQRLSMMHSLYSSI 547
F+ +V SC+S+++ + + IWH RLGH+ +RL M
Sbjct: 503 ------FVHESVCMA---SCSSITN------MTKSEIWHARLGHVHYKRLKDMSKTCMIP 637
Query: 548 TID-NKAVCDICHFAK 562
D N C C K
Sbjct: 638 PFDMNIEKCKTCMLTK 685
Score = 27.3 bits (59), Expect(2) = 0.003
Identities = 9/19 (47%), Positives = 11/19 (57%)
Frame = +1
Query: 373 WLIDSGANEHICSSLHLFH 391
W DSGA H+C LF+
Sbjct: 154 WWFDSGATSHVCKDRRLFY 210
>BG881907 homologue to GP|21432|emb|CA ORF2 {Solanum tuberosum}, partial (3%)
Length = 429
Score = 40.8 bits (94), Expect = 0.004
Identities = 14/65 (21%), Positives = 41/65 (62%), Gaps = 1/65 (1%)
Frame = -1
Query: 81 IAQTIVFHEYAIDVWIELQERFSKVDRIR-VASLRSSINNLKQGDKSVLDYFTSIKSLWE 139
I + ++++ ++W ++E +S +D + ++S +++L+QG+KSV ++ ++ W+
Sbjct: 429 IGENFMYYDPTKEIWDAVKETYSNIDNTSAIFEIKSLLHDLQQGEKSVTEFSNTLSRYWQ 250
Query: 140 ELNSH 144
L+++
Sbjct: 249 RLDTY 235
>TC233176 weakly similar to GB|AAL07033.1|15810291|AY056184 auxin response
factor ARF17 {Arabidopsis thaliana;} , partial (17%)
Length = 958
Score = 40.4 bits (93), Expect = 0.006
Identities = 23/72 (31%), Positives = 39/72 (53%), Gaps = 1/72 (1%)
Frame = +3
Query: 467 QKMIGLGNLCDGLYRLHPFAPASP-QAHFISSAVSPTNKMSCNSVSSNNNVSSIPSNAIW 525
Q+ IG + +GLYRL + S ++ F++S++ + SCN + + +W
Sbjct: 768 QQKIGTIEMVEGLYRLKMISFKSKYKSDFVNSSIFNVSAFSCNKIPID----------LW 917
Query: 526 HFRLGHLSNQRL 537
H+RLGH S +RL
Sbjct: 918 HYRLGHPSLERL 953
>TC232838 similar to GB|AAP21377.1|30102918|BT006569 At1g47970 {Arabidopsis
thaliana;} , partial (46%)
Length = 902
Score = 40.0 bits (92), Expect = 0.007
Identities = 37/154 (24%), Positives = 70/154 (45%)
Frame = -3
Query: 212 SNIIPPTSLASNEDSSILVNASDARKPFLRGKSSGTSQSKNNSRYCTFCRRNNHTVEYCY 271
S+ PP+S + + S + + + +S +S S +S + ++ V
Sbjct: 768 SSSSPPSSSSESSLSDLFLFGGASAFSQSSSSTSSSSSSPFSSGSKSLDASSSSAVSKGR 589
Query: 272 LKHDFPNANKPTASSNAVTSEHAVDSHTSSEGTSSSSQTGLTQEQYVHLVSLLQQSSLVP 331
+++ P ++ SS++ +S + S +SSE SSSS + S SS
Sbjct: 588 IRYSVPKSSSSPCSSSSSSSSSSSSSESSSESLSSSSGSP------PRWTSPSSSSSSSS 427
Query: 332 SATPPNPASTNHVATSFPSSIDFTSGINTIFSCS 365
S++PP+ AS++ + S PSS +S ++ S S
Sbjct: 426 SSSPPSGASSSSSSLSSPSSSSSSSSSSSSSSAS 325
>CF922727
Length = 791
Score = 39.7 bits (91), Expect = 0.010
Identities = 32/144 (22%), Positives = 64/144 (44%), Gaps = 8/144 (5%)
Frame = -2
Query: 8 SEGPNSVTVTPLLTGPNYLAWNRSMKRALGTKN-KFVFIDGSVPI------PPMDDLNRT 60
S+ N P+L G NY W + LG + + P+ P + DL
Sbjct: 460 SQSMNISCDLPILKGDNYKVWKERVLLHLGWMDIDYAIRKDEPPVITETSEPDVVDLYEK 281
Query: 61 AWERCNNLILSWIINSVSPQIAQTIVFHEYAIDVWIELQERFSKVDRIRVASLRSSINNL 120
WER N L + +I ++S I ++ H+ D+ + ERF+ ++ ++L +++
Sbjct: 280 -WERSNRLSVMFIKTNISTSIWGSVDQHDKVRDLLKAIDERFTTSEKSLASTLIMQFSSI 104
Query: 121 K-QGDKSVLDYFTSIKSLWEELNS 143
K G + V ++ ++ + +L +
Sbjct: 103 KLTGTRGVREHIMRLRDIVAQLKT 32
>CO983960
Length = 875
Score = 38.9 bits (89), Expect = 0.017
Identities = 21/66 (31%), Positives = 37/66 (55%)
Frame = +1
Query: 44 FIDGSVPIPPMDDLNRTAWERCNNLILSWIINSVSPQIAQTIVFHEYAIDVWIELQERFS 103
F+ ++ P D AW++C L+LSW+ + VS +IA I++ DV + + RFS
Sbjct: 160 FVGNTISKPRDKDPLYFAWKQCKTLVLSWLNHRVSHEIAANIIWINDDKDV*QDHKYRFS 339
Query: 104 KVDRIR 109
+ D ++
Sbjct: 340 QDDFVQ 357
>TC231899 similar to UP|Q850H7 (Q850H7) Gag-pol polyprotein (Fragment), partial
(30%)
Length = 687
Score = 38.9 bits (89), Expect = 0.017
Identities = 38/105 (36%), Positives = 52/105 (49%)
Frame = +1
Query: 1278 IH*CRLGRL*RYKAFYLRTLFLSWTISYMLENQETTYSFQILF*S*V*SNGICYL*NAMV 1337
I *C LG L Y+R L L W S LE QET + S*+ +G YL* +
Sbjct: 25 IL*C*LGWLSHG*EVYIRLLCLHWRKSCFLEKQETDGCSSV*CRS*ISIDGYGYL*THVD 204
Query: 1338 IILAQRFTSSMCSTSCALL**SKCYAHCIKSCFS*KN*TS*N*LS 1382
++ R + +L**S C ++C+K S*K+* +*LS
Sbjct: 205 *TISARIEVL*RVANEVVL**SGCSSYCLKPSLS*KD*AYRD*LS 339
>TC233836
Length = 739
Score = 38.9 bits (89), Expect = 0.017
Identities = 30/102 (29%), Positives = 44/102 (42%), Gaps = 4/102 (3%)
Frame = -3
Query: 60 TAWERCNNLILSWIINSVSPQIAQTIVFHEYAIDVWIE-LQERFSKVDRIRVASLRSSIN 118
+AW++CN L+LSW VSP Q F + + + F KV + + +
Sbjct: 716 SAWKQCNTLVLSWSSCPVSPTRLQPTSFGSMLTKISGKIINTIFHKVTLFKSSKFKVIFF 537
Query: 119 NLKQGDKSVLDYFTSIKSLWEELNSHRPMP---MCTCPYPCR 157
LKQ D S+ Y TS ++ PMP CT C+
Sbjct: 536 ILKQADMSITSYCTSSYAV--------PMP*FSKCTMLLDCQ 435
>TC235104 weakly similar to UP|Q850H8 (Q850H8) Gag-pol polyprotein
(Fragment), partial (28%)
Length = 865
Score = 38.5 bits (88), Expect = 0.022
Identities = 40/165 (24%), Positives = 64/165 (38%)
Frame = +2
Query: 398 PICVNLPNGSSVIVQYAGTVVFSPHFHITHVLYSPSFKVYLISVSKICQSLPYHVHFLLN 457
P + L +GS V+ G V + + V++ + S+S++ + V F N
Sbjct: 11 PYFITLADGSRVVATGIGHVSPTSSLSLNSVVFILGCPFNITSLSQLTRFRNCSVTFDAN 190
Query: 458 TYVIQDVKTQKMIGLGNLCDGLYRLHPFAPASPQAHFISSAVSPTNKMSCNSVSSNNNVS 517
++VIQ+ T IG+G GLY L P ++ SAV+
Sbjct: 191 SFVIQECGTGWTIGVGIESHGLYYL------KPNLSWVCSAVT----------------- 301
Query: 518 SIPSNAIWHFRLGHLSNQRLSMMHSLYSSITIDNKAVCDICHFAK 562
S + H RLGH LS + + S+ C+ C K
Sbjct: 302 ---SPKLLHERLGH---PHLSKLKIMVPSLEKIKDLFCESCQLGK 418
>BE556314 similar to GP|29150404|gb gag-pol polyprotein {Oryza sativa
(japonica cultivar-group)}, partial (1%)
Length = 430
Score = 38.5 bits (88), Expect = 0.022
Identities = 31/103 (30%), Positives = 47/103 (45%), Gaps = 2/103 (1%)
Frame = -2
Query: 493 HFISSAVSPTNKMSCNS--VSSNNNVSSIPSNAIWHFRLGHLSNQRLSMMHSLYSSITID 550
H I + S T ++ NS V SN SS+ +WH RLGH ++ +
Sbjct: 294 HVIGDSPSSTLAINNNSSTVMSNFVSSSLSIANLWHARLGHPNDHVM------------- 154
Query: 551 NKAVCDICHFAKQRKLPYNLSTLLLHLNLNYCILIFGVLFLLL 593
K V C+ K L L++ +NL Y + +F +LF+LL
Sbjct: 153 -KLVLTHCNIPSSNKNSTGLVLLVVWVNLIYYLFMFLLLFILL 28
Database: GMGI
Posted date: Oct 22, 2004 4:58 PM
Number of letters in database: 37,918,896
Number of sequences in database: 63,676
Lambda K H
0.356 0.156 0.555
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 76,378,679
Number of Sequences: 63676
Number of extensions: 1313751
Number of successful extensions: 17826
Number of sequences better than 10.0: 70
Number of HSP's better than 10.0 without gapping: 10131
Number of HSP's successfully gapped in prelim test: 910
Number of HSP's that attempted gapping in prelim test: 6989
Number of HSP's gapped (non-prelim): 12240
length of query: 1441
length of database: 12,639,632
effective HSP length: 109
effective length of query: 1332
effective length of database: 5,698,948
effective search space: 7590998736
effective search space used: 7590998736
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 14 ( 7.2 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 37 (21.7 bits)
S2: 65 (29.6 bits)
Medicago: description of AC149491.1


