

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC149305.4 - phase: 0 /partial
(542 letters)
Database: GMGI
63,676 sequences; 37,918,896 total letters
Searching..................................................done
Score E
Sequences producing significant alignments: (bits) Value
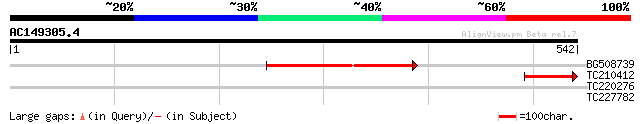
BG508739 244 9e-65
TC210412 weakly similar to UP|Q7F266 (Q7F266) Nipped-B gene prod... 69 5e-12
TC220276 weakly similar to UP|O04238 (O04238) Transcription fact... 29 4.7
TC227782 29 6.1
>BG508739
Length = 437
Score = 244 bits (622), Expect = 9e-65
Identities = 123/145 (84%), Positives = 134/145 (91%)
Frame = +1
Query: 246 FNTQVRQSALKIVEVVLRQGLVHPITCVPYLIALETDPLESNSKMAHHLLMNMHEKYPAF 305
F+ VRQSALKIVEVVLRQGLVHPITCVPYLIALETDPLESNSK+AHHLLMNM+EKYPAF
Sbjct: 4 FSEHVRQSALKIVEVVLRQGLVHPITCVPYLIALETDPLESNSKLAHHLLMNMNEKYPAF 183
Query: 306 FESCLGDGLQMSFMFMQSIFVSPDENVNHKSQSKIAVSGKGKPEADSLAQSRVGVSRIYK 365
FES LGDGLQMSFMFMQSI S ENV+HK QSK + SGKGKPEA SLAQ+++GVSRIYK
Sbjct: 184 FESRLGDGLQMSFMFMQSICGS-SENVDHKIQSKTSTSGKGKPEAGSLAQAKLGVSRIYK 360
Query: 366 LIRGNRISRNKFMSSIVRKFDNPKW 390
LIRGNR+SRNKF+SSIVRKFDNP+W
Sbjct: 361 LIRGNRVSRNKFLSSIVRKFDNPRW 435
>TC210412 weakly similar to UP|Q7F266 (Q7F266) Nipped-B gene product-like
protein, partial (6%)
Length = 817
Score = 68.9 bits (167), Expect = 5e-12
Identities = 30/50 (60%), Positives = 40/50 (80%)
Frame = +1
Query: 493 DDMTSVDLNGTNHQLPDYPLSHNGRLKVKPQAAGFADSFTFSKDDLEKVQ 542
+DM ++DLNG+NHQL DYPLS+ G + K +AG+ D F+F+KDDLEKVQ
Sbjct: 1 NDMRTLDLNGSNHQLTDYPLSYKGSSEAKLHSAGYTDPFSFAKDDLEKVQ 150
>TC220276 weakly similar to UP|O04238 (O04238) Transcription factor, partial
(6%)
Length = 538
Score = 29.3 bits (64), Expect = 4.7
Identities = 18/57 (31%), Positives = 27/57 (46%)
Frame = +3
Query: 433 PLEANFKAWSSSLLQSEGQGTPHGNGMYQRATDETIHSTQGQSMDLNGPFQQNVDVQ 489
P NF+A + + GQG+ H + + T HS Q + M N P + +V VQ
Sbjct: 3 PQSGNFQAAARGASVAAGQGSSHARNV--PTSGATTHSHQARGMVANQPARPSVLVQ 167
>TC227782
Length = 460
Score = 28.9 bits (63), Expect = 6.1
Identities = 20/73 (27%), Positives = 35/73 (47%), Gaps = 7/73 (9%)
Frame = +1
Query: 286 SNSKMAHHLLMNMHEKYPAFFESCLGDGLQMSFMFMQSIFVSPD---ENVNHKSQS---- 338
+NS M HHLLM+ + + + MS + ++++ +SP ++ S+S
Sbjct: 226 NNSLMKHHLLMSQTRRLAVLIRAPRSPSVTMSLLSLKTLALSPPW*WTSLKLGSRSLRR* 405
Query: 339 KIAVSGKGKPEAD 351
+ KGKPE D
Sbjct: 406 TLLWRRKGKPEMD 444
Database: GMGI
Posted date: Oct 22, 2004 4:58 PM
Number of letters in database: 37,918,896
Number of sequences in database: 63,676
Lambda K H
0.321 0.137 0.397
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 22,028,727
Number of Sequences: 63676
Number of extensions: 297160
Number of successful extensions: 1393
Number of sequences better than 10.0: 8
Number of HSP's better than 10.0 without gapping: 1386
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 1392
length of query: 542
length of database: 12,639,632
effective HSP length: 102
effective length of query: 440
effective length of database: 6,144,680
effective search space: 2703659200
effective search space used: 2703659200
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 16 ( 7.4 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.8 bits)
S2: 61 (28.1 bits)
Medicago: description of AC149305.4


