

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC149305.10 + phase: 0 /pseudo
(55 letters)
Database: GMGI
63,676 sequences; 37,918,896 total letters
Searching..................................................done
Score E
Sequences producing significant alignments: (bits) Value
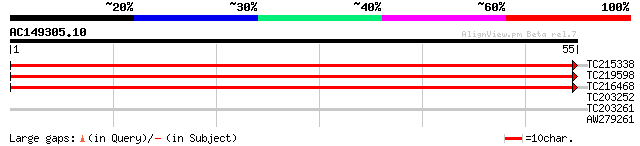
TC215338 UP|UGDH_SOYBN (Q96558) UDP-glucose 6-dehydrogenase (UD... 81 1e-16
TC219598 homologue to UP|UGDH_SOYBN (Q96558) UDP-glucose 6-dehyd... 81 1e-16
TC216468 homologue to UP|Q9FZE1 (Q9FZE1) T1K7.6 protein (At1g265... 69 4e-13
TC203252 similar to GB|BAA11677.1|1181599|CUSPSALG subunit of ph... 25 8.3
TC203261 similar to UP|Q84UU0 (Q84UU0) Photosystem I subunit XI,... 25 8.3
AW279261 similar to GP|1181599|dbj subunit of photosystem I {Cuc... 25 8.3
>TC215338 UP|UGDH_SOYBN (Q96558) UDP-glucose 6-dehydrogenase (UDP-Glc
dehydrogenase) (UDP-GlcDH) (UDPGDH) , complete
Length = 1780
Score = 81.3 bits (199), Expect = 1e-16
Identities = 38/55 (69%), Positives = 45/55 (81%)
Frame = +3
Query: 1 MAAIALKFTSTDVTVVDITKSRIVAWYNDQIPIYQPGFDGLVKQCRIKNLFFGTN 55
MA IALK S +V VVDI+KSRI AW +DQ+PIY+PG DG+VKQCR KNLFF T+
Sbjct: 117 MAVIALKCPSIEVAVVDISKSRIAAWNSDQLPIYEPGLDGVVKQCRGKNLFFSTD 281
>TC219598 homologue to UP|UGDH_SOYBN (Q96558) UDP-glucose 6-dehydrogenase
(UDP-Glc dehydrogenase) (UDP-GlcDH) (UDPGDH) , partial
(58%)
Length = 882
Score = 81.3 bits (199), Expect = 1e-16
Identities = 38/55 (69%), Positives = 45/55 (81%)
Frame = +3
Query: 1 MAAIALKFTSTDVTVVDITKSRIVAWYNDQIPIYQPGFDGLVKQCRIKNLFFGTN 55
MA IALK S +V VVDI+KSRI AW +DQ+PIY+PG DG+VKQCR KNLFF T+
Sbjct: 96 MAVIALKCPSIEVAVVDISKSRIAAWNSDQLPIYEPGLDGVVKQCRGKNLFFSTD 260
>TC216468 homologue to UP|Q9FZE1 (Q9FZE1) T1K7.6 protein (At1g26570/T1K7_6),
partial (25%)
Length = 574
Score = 69.3 bits (168), Expect = 4e-13
Identities = 33/55 (60%), Positives = 39/55 (70%)
Frame = +2
Query: 1 MAAIALKFTSTDVTVVDITKSRIVAWYNDQIPIYQPGFDGLVKQCRIKNLFFGTN 55
MA IALK +V VVDI RI AW +D +PIY+PG D +VKQCR KNLFF T+
Sbjct: 182 MAVIALKCPEIEVVVVDIAAPRINAWNSDHLPIYEPGLDDVVKQCRGKNLFFSTD 346
>TC203252 similar to GB|BAA11677.1|1181599|CUSPSALG subunit of photosystem I
{Cucumis sativus;} , partial (80%)
Length = 823
Score = 25.0 bits (53), Expect = 8.3
Identities = 8/26 (30%), Positives = 15/26 (56%)
Frame = +1
Query: 18 ITKSRIVAWYNDQIPIYQPGFDGLVK 43
+T S ++AWY +P Y+ L++
Sbjct: 223 VTSSPLIAWYLSNLPAYRTAVSPLLR 300
>TC203261 similar to UP|Q84UU0 (Q84UU0) Photosystem I subunit XI, partial
(88%)
Length = 997
Score = 25.0 bits (53), Expect = 8.3
Identities = 8/26 (30%), Positives = 15/26 (56%)
Frame = +1
Query: 18 ITKSRIVAWYNDQIPIYQPGFDGLVK 43
+T S ++AWY +P Y+ L++
Sbjct: 421 VTSSPLIAWYLSNLPAYRTAVSPLLR 498
>AW279261 similar to GP|1181599|dbj subunit of photosystem I {Cucumis
sativus}, partial (52%)
Length = 346
Score = 25.0 bits (53), Expect = 8.3
Identities = 8/26 (30%), Positives = 15/26 (56%)
Frame = +2
Query: 18 ITKSRIVAWYNDQIPIYQPGFDGLVK 43
+T S ++AWY +P Y+ L++
Sbjct: 44 VTSSPLIAWYLSNLPAYRTAVSPLLR 121
Database: GMGI
Posted date: Oct 22, 2004 4:58 PM
Number of letters in database: 37,918,896
Number of sequences in database: 63,676
Lambda K H
0.326 0.140 0.428
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 2,176,363
Number of Sequences: 63676
Number of extensions: 18026
Number of successful extensions: 103
Number of sequences better than 10.0: 13
Number of HSP's better than 10.0 without gapping: 103
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 103
length of query: 55
length of database: 12,639,632
effective HSP length: 31
effective length of query: 24
effective length of database: 10,665,676
effective search space: 255976224
effective search space used: 255976224
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 15 ( 7.1 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 40 (21.7 bits)
S2: 52 (24.6 bits)
Medicago: description of AC149305.10


