

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
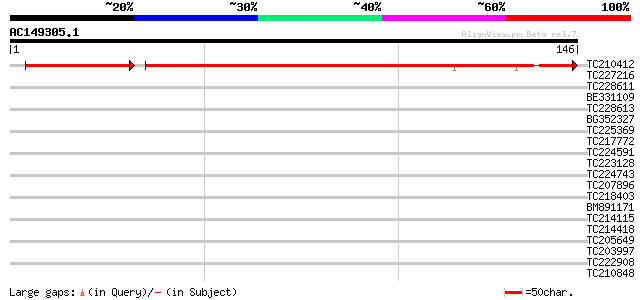
Query= AC149305.1 + phase: 0 /pseudo
(146 letters)
Database: GMGI
63,676 sequences; 37,918,896 total letters
Searching..................................................done
Score E
Sequences producing significant alignments: (bits) Value
TC210412 weakly similar to UP|Q7F266 (Q7F266) Nipped-B gene prod... 157 7e-46
TC227216 GB|AAA33994.1|170031|SOYNOD35G nodulin 35 {Glycine max;... 28 1.2
TC228611 weakly similar to GB|CAA72487.1|1890317|ATP26A peroxida... 28 1.6
BE331109 28 1.6
TC228613 weakly similar to UP|PE31_ARATH (Q9LHA7) Peroxidase 31 ... 28 1.6
BG352327 similar to PIR|D84745|D847 plastid protein [imported] -... 28 2.1
TC225369 similar to UP|Q9FNI7 (Q9FNI7) Glucosyltransferase-like ... 27 3.5
TC217772 similar to GB|AAM51588.1|21436021|AY116954 AT5g19430/F7... 27 3.5
TC224591 homologue to UP|Q6H8J2 (Q6H8J2) 40S ribosomal protein S... 27 4.6
TC223128 similar to UP|WR27_ARATH (Q9FLX8) Probable WRKY transcr... 27 4.6
TC224743 homologue to UP|Q6H8J2 (Q6H8J2) 40S ribosomal protein S... 27 4.6
TC207896 GB|BAA13184.1|1498170|D86929 uricase {Glycine max;} , c... 26 6.0
TC218403 UP|Q39895 (Q39895) TGACG-motif binding factor, complete 26 6.0
BM891171 26 7.8
TC214115 similar to UP|O22793 (O22793) Plastid protein (At2g3343... 26 7.8
TC214418 homologue to UP|Q8L949 (Q8L949) Plastid protein, partia... 26 7.8
TC205649 26 7.8
TC203997 similar to UP|Q9FFJ6 (Q9FFJ6) Phosphate/phosphoenolpyru... 26 7.8
TC222908 26 7.8
TC210848 homologue to GB|AAR28007.1|39545888|AY463605 TFIIE-alph... 26 7.8
>TC210412 weakly similar to UP|Q7F266 (Q7F266) Nipped-B gene product-like
protein, partial (6%)
Length = 817
Score = 157 bits (396), Expect(2) = 7e-46
Identities = 84/114 (73%), Positives = 96/114 (83%), Gaps = 3/114 (2%)
Frame = +1
Query: 36 SEPPKPGDVFSRQSIPFNIGESQFSLPTSPQELIQRYQEFKNALKEDTVDYSLYTANIKR 95
+E PKPG+V SRQ+I FNIG+SQFSLPTSPQELIQRYQEFK+AL+EDTVDYS YTANIKR
Sbjct: 256 TEQPKPGEVISRQNIAFNIGDSQFSLPTSPQELIQRYQEFKHALREDTVDYSHYTANIKR 435
Query: 96 KRPTQTPRKVRKSGPPMV-GGDNDEDDDEDWAGGS--RNISFSGGRRSSLRNSR 146
KRPT TPR+V+ P V GG ++ DDDED+ GGS R ISFS GRRSSL+NSR
Sbjct: 436 KRPTTTPRRVQVRKPVYVAGGYDNGDDDEDYTGGSATRKISFS-GRRSSLKNSR 594
Score = 43.1 bits (100), Expect(2) = 7e-46
Identities = 22/28 (78%), Positives = 24/28 (85%)
Frame = +2
Query: 5 RLTVYLRLRCNFF*S*RDT*KSCIVSMM 32
R TVY +L C+FF*S*RDT*KSCIV MM
Sbjct: 149 RPTVYQQLHCSFF*S*RDT*KSCIVLMM 232
>TC227216 GB|AAA33994.1|170031|SOYNOD35G nodulin 35 {Glycine max;} , complete
Length = 1267
Score = 28.5 bits (62), Expect = 1.2
Identities = 12/26 (46%), Positives = 16/26 (61%)
Frame = +2
Query: 56 ESQFSLPTSPQELIQRYQEFKNALKE 81
ESQ+SLP P ++YQE K L +
Sbjct: 677 ESQYSLPQKPFYFTEKYQEVKKVLAD 754
>TC228611 weakly similar to GB|CAA72487.1|1890317|ATP26A peroxidase ATP26a
{Arabidopsis thaliana;} , partial (49%)
Length = 731
Score = 28.1 bits (61), Expect = 1.6
Identities = 11/22 (50%), Positives = 14/22 (63%)
Frame = -3
Query: 90 TANIKRKRPTQTPRKVRKSGPP 111
TA + RP++ PRK K GPP
Sbjct: 492 TAEAREVRPSRRPRKTGKKGPP 427
>BE331109
Length = 475
Score = 28.1 bits (61), Expect = 1.6
Identities = 12/35 (34%), Positives = 21/35 (59%)
Frame = +3
Query: 90 TANIKRKRPTQTPRKVRKSGPPMVGGDNDEDDDED 124
+A I+ K T PRK + + P + G + D++DD +
Sbjct: 15 SA*IQAKGFTPMPRKCKSAEPQVAGANEDQEDDNN 119
>TC228613 weakly similar to UP|PE31_ARATH (Q9LHA7) Peroxidase 31 precursor
(Atperox P31) (ATP41) , partial (50%)
Length = 536
Score = 28.1 bits (61), Expect = 1.6
Identities = 11/22 (50%), Positives = 14/22 (63%)
Frame = -3
Query: 90 TANIKRKRPTQTPRKVRKSGPP 111
TA + RP++ PRK K GPP
Sbjct: 153 TAEAREVRPSRRPRKTGKKGPP 88
>BG352327 similar to PIR|D84745|D847 plastid protein [imported] - Arabidopsis
thaliana, partial (23%)
Length = 526
Score = 27.7 bits (60), Expect = 2.1
Identities = 13/23 (56%), Positives = 16/23 (69%), Gaps = 2/23 (8%)
Frame = -2
Query: 107 KSGPPMVGGD--NDEDDDEDWAG 127
K+GP VGG NDEDDD ++ G
Sbjct: 288 KTGPISVGGGSLNDEDDDPEFTG 220
>TC225369 similar to UP|Q9FNI7 (Q9FNI7) Glucosyltransferase-like protein
(AT5g22740/MDJ22_16), partial (90%)
Length = 2588
Score = 26.9 bits (58), Expect = 3.5
Identities = 16/62 (25%), Positives = 28/62 (44%)
Frame = +3
Query: 25 KSCIVSMMLDVSEPPKPGDVFSRQSIPFNIGESQFSLPTSPQELIQRYQEFKNALKEDTV 84
K C + D P+P F R+SIPF +G +L + + + ++E ++
Sbjct: 909 KHCEYVAIFDADFRPEPD--FLRRSIPFLVGNPDIALVQARWRFVNSDECLLTRMQEMSL 1082
Query: 85 DY 86
DY
Sbjct: 1083DY 1088
>TC217772 similar to GB|AAM51588.1|21436021|AY116954 AT5g19430/F7K24_180
{Arabidopsis thaliana;} , partial (61%)
Length = 1136
Score = 26.9 bits (58), Expect = 3.5
Identities = 11/24 (45%), Positives = 16/24 (65%)
Frame = +2
Query: 116 DNDEDDDEDWAGGSRNISFSGGRR 139
++D+D DE + GGS N+ G RR
Sbjct: 617 EDDDDMDEAYYGGSSNLRVIGNRR 688
>TC224591 homologue to UP|Q6H8J2 (Q6H8J2) 40S ribosomal protein S9, partial
(77%)
Length = 730
Score = 26.6 bits (57), Expect = 4.6
Identities = 15/41 (36%), Positives = 24/41 (57%), Gaps = 1/41 (2%)
Frame = +3
Query: 84 VDYSLYTANIKRKRPTQTPRK-VRKSGPPMVGGDNDEDDDE 123
+D+SL T+ + RP + RK R + GGD DE+D++
Sbjct: 342 IDFSL-TSPLGGGRPGRVKRKNQRAAAKKAAGGDGDEEDED 461
>TC223128 similar to UP|WR27_ARATH (Q9FLX8) Probable WRKY transcription
factor 27 (WRKY DNA-binding protein 27), partial (21%)
Length = 821
Score = 26.6 bits (57), Expect = 4.6
Identities = 15/39 (38%), Positives = 20/39 (50%), Gaps = 1/39 (2%)
Frame = +2
Query: 30 SMMLDVSEPPKPGDVFSRQSIPFNIGESQ-FSLPTSPQE 67
S L + PP PG + Q+ PF+ S PTSP+E
Sbjct: 203 SKTLVTNPPPSPGSLSCFQATPFSSSSSSPPHSPTSPEE 319
>TC224743 homologue to UP|Q6H8J2 (Q6H8J2) 40S ribosomal protein S9, complete
Length = 935
Score = 26.6 bits (57), Expect = 4.6
Identities = 15/41 (36%), Positives = 24/41 (57%), Gaps = 1/41 (2%)
Frame = +3
Query: 84 VDYSLYTANIKRKRPTQTPRK-VRKSGPPMVGGDNDEDDDE 123
+D+SL T+ + RP + RK R + GGD DE+D++
Sbjct: 522 IDFSL-TSPLGGGRPGRVKRKNQRAAAKKAAGGDGDEEDED 641
>TC207896 GB|BAA13184.1|1498170|D86929 uricase {Glycine max;} , complete
Length = 1348
Score = 26.2 bits (56), Expect = 6.0
Identities = 11/26 (42%), Positives = 15/26 (57%)
Frame = +3
Query: 56 ESQFSLPTSPQELIQRYQEFKNALKE 81
ES +SLP P ++YQE K L +
Sbjct: 642 ESLYSLPQKPLYFTEKYQEVKKVLAD 719
>TC218403 UP|Q39895 (Q39895) TGACG-motif binding factor, complete
Length = 1195
Score = 26.2 bits (56), Expect = 6.0
Identities = 17/56 (30%), Positives = 26/56 (46%), Gaps = 1/56 (1%)
Frame = +1
Query: 86 YSLYTANIKRKRPT-QTPRKVRKSGPPMVGGDNDEDDDEDWAGGSRNISFSGGRRS 140
Y N+ + P +T R+ G + G D+DEDD +D G I++ G S
Sbjct: 253 YEYELKNVSQSCPQCKTTFTSRQEGAEVEGDDDDEDDADDLDNG---INYGQGNNS 411
>BM891171
Length = 424
Score = 25.8 bits (55), Expect = 7.8
Identities = 9/16 (56%), Positives = 11/16 (68%)
Frame = +2
Query: 113 VGGDNDEDDDEDWAGG 128
V D+D+DDD DW G
Sbjct: 266 VAADDDDDDDVDWEEG 313
>TC214115 similar to UP|O22793 (O22793) Plastid protein (At2g33430/F4P9.20),
partial (46%)
Length = 557
Score = 25.8 bits (55), Expect = 7.8
Identities = 13/23 (56%), Positives = 16/23 (69%), Gaps = 2/23 (8%)
Frame = -2
Query: 107 KSGPPMVGGD--NDEDDDEDWAG 127
KSG VGG NDEDDD +++G
Sbjct: 325 KSGAISVGGRSLNDEDDDPEFSG 257
>TC214418 homologue to UP|Q8L949 (Q8L949) Plastid protein, partial (74%)
Length = 1130
Score = 25.8 bits (55), Expect = 7.8
Identities = 13/23 (56%), Positives = 16/23 (69%), Gaps = 2/23 (8%)
Frame = -2
Query: 107 KSGPPMVGGD--NDEDDDEDWAG 127
KSG VGG NDEDDD +++G
Sbjct: 136 KSGAISVGGRSLNDEDDDPEFSG 68
>TC205649
Length = 1747
Score = 25.8 bits (55), Expect = 7.8
Identities = 11/43 (25%), Positives = 21/43 (48%)
Frame = -1
Query: 89 YTANIKRKRPTQTPRKVRKSGPPMVGGDNDEDDDEDWAGGSRN 131
+T N + + + ++ + M+G D+D D D D+ RN
Sbjct: 277 HTPNKMKPKHNRA*LEIARPNMKMIGSDSDSDSDSDFTLTHRN 149
>TC203997 similar to UP|Q9FFJ6 (Q9FFJ6) Phosphate/phosphoenolpyruvate
translocator protein-like, partial (97%)
Length = 1271
Score = 25.8 bits (55), Expect = 7.8
Identities = 10/28 (35%), Positives = 14/28 (49%)
Frame = -3
Query: 98 PTQTPRKVRKSGPPMVGGDNDEDDDEDW 125
P QTPR++ PP G +N+ W
Sbjct: 102 PQQTPRRLHFPSPPQSGSENNNIAHH*W 19
>TC222908
Length = 535
Score = 25.8 bits (55), Expect = 7.8
Identities = 12/31 (38%), Positives = 17/31 (54%), Gaps = 4/31 (12%)
Frame = +2
Query: 100 QTPRKVRKSGPPMVGGDNDEDDDE----DWA 126
++PR R GPP +G D +D E +WA
Sbjct: 332 RSPRWWRSGGPPRIGVSGDAEDAEVVEGNWA 424
>TC210848 homologue to GB|AAR28007.1|39545888|AY463605 TFIIE-alpha 1
{Arabidopsis thaliana;} , partial (4%)
Length = 916
Score = 25.8 bits (55), Expect = 7.8
Identities = 9/16 (56%), Positives = 11/16 (68%)
Frame = +3
Query: 113 VGGDNDEDDDEDWAGG 128
V D+D+DDD DW G
Sbjct: 459 VAADDDDDDDVDWEEG 506
Database: GMGI
Posted date: Oct 22, 2004 4:58 PM
Number of letters in database: 37,918,896
Number of sequences in database: 63,676
Lambda K H
0.329 0.143 0.449
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 6,501,767
Number of Sequences: 63676
Number of extensions: 82463
Number of successful extensions: 893
Number of sequences better than 10.0: 41
Number of HSP's better than 10.0 without gapping: 759
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 860
length of query: 146
length of database: 12,639,632
effective HSP length: 88
effective length of query: 58
effective length of database: 7,036,144
effective search space: 408096352
effective search space used: 408096352
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 15 ( 7.1 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 40 (21.8 bits)
S2: 54 (25.4 bits)
Medicago: description of AC149305.1


