

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
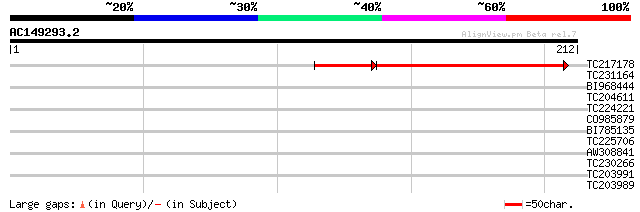
Query= AC149293.2 - phase: 2
(212 letters)
Database: GMGI
63,676 sequences; 37,918,896 total letters
Searching..................................................done
Score E
Sequences producing significant alignments: (bits) Value
TC217178 similar to UP|Q8LDE3 (Q8LDE3) Enhanced disease suscepti... 100 3e-30
TC231164 weakly similar to UP|UFO2_MANES (Q40285) Flavonol 3-O-g... 32 0.28
BI968444 weakly similar to GP|11994643|db UTP-glucose glucosyltr... 30 1.1
TC204611 similar to UP|Q9XEY5 (Q9XEY5) Nt-iaa28 deduced protein,... 28 3.1
TC224221 weakly similar to UP|Q8V7F2 (Q8V7F2) ORF1 (Fragment), p... 27 5.2
CO985879 27 5.2
BI785135 27 5.2
TC225706 27 6.9
AW308841 similar to PIR|JQ2128|JQ212 metallothionein - soybean, ... 27 6.9
TC230266 similar to UP|GRP1_PETHY (P09789) Glycine-rich cell wal... 27 6.9
TC203991 UP|Q7M213 (Q7M213) Metallothionein, complete 27 8.9
TC203989 homologue to UP|Q7M213 (Q7M213) Metallothionein, partia... 27 8.9
>TC217178 similar to UP|Q8LDE3 (Q8LDE3) Enhanced disease susceptibility 5,
partial (35%)
Length = 994
Score = 100 bits (248), Expect(2) = 3e-30
Identities = 51/72 (70%), Positives = 55/72 (75%)
Frame = +3
Query: 138 YGVNLSRKLLNHLCLNCYMELIGICQRPGCF*DLLQ*SELHLDCY*ELLEHQFLSYFPTF 197
YGVNLS K LNHLCLN YME IG+CQRP CF* L * EL+LDC * LLEH+FL YFP F
Sbjct: 186 YGVNLSPKPLNHLCLN*YME*IGVCQRPECF*GLS**LELYLDCC*GLLEHRFLGYFPIF 365
Query: 198 LHLIRWS*GRSV 209
LHLI WS R +
Sbjct: 366 LHLIGWSSRRCI 401
Score = 48.9 bits (115), Expect(2) = 3e-30
Identities = 22/23 (95%), Positives = 23/23 (99%)
Frame = +2
Query: 115 VAFYSLLIYFATSMGTHTMAAHQ 137
VAFY+LLIYFATSMGTHTMAAHQ
Sbjct: 86 VAFYALLIYFATSMGTHTMAAHQ 154
>TC231164 weakly similar to UP|UFO2_MANES (Q40285) Flavonol
3-O-glucosyltransferase 2 (UDP-glucose flavonoid
3-O-glucosyltransferase 2) (Fragment) , partial (47%)
Length = 901
Score = 31.6 bits (70), Expect = 0.28
Identities = 19/69 (27%), Positives = 30/69 (42%)
Frame = +2
Query: 13 LCGWVAQSASLGMKDSWGPLKALAAASVINGVGDIVLCTYLGYGIAGAAWATMASQVVAA 72
+CGWV Q+ L K A ++ G + L +G+ A W A Q + A
Sbjct: 254 VCGWVPQAVVLAHK---------AVGGFVSHCGWNSILESLWHGVPIATWPVYAEQQMNA 406
Query: 73 YMMMRTLNM 81
+ M+R L +
Sbjct: 407 FQMVRELGL 433
>BI968444 weakly similar to GP|11994643|db UTP-glucose glucosyltransferase
{Arabidopsis thaliana}, partial (25%)
Length = 586
Score = 29.6 bits (65), Expect = 1.1
Identities = 18/73 (24%), Positives = 33/73 (44%), Gaps = 6/73 (8%)
Frame = -3
Query: 15 GWVAQSASLGMKDSWGPLKALAAASVINGVGDIVLCTY------LGYGIAGAAWATMASQ 68
G++ ++A +G W P + A G + C + + +G+ A W A Q
Sbjct: 518 GFLDRTAGIGKVIGWAPQAQILAHPATGGF--VSHCGWNSTLESIYFGVPIATWPLYAEQ 345
Query: 69 VVAAYMMMRTLNM 81
A++++R LNM
Sbjct: 344 QTNAFLLVRELNM 306
>TC204611 similar to UP|Q9XEY5 (Q9XEY5) Nt-iaa28 deduced protein, partial
(50%)
Length = 715
Score = 28.1 bits (61), Expect = 3.1
Identities = 13/60 (21%), Positives = 30/60 (49%)
Frame = -3
Query: 108 FMTMMSKVAFYSLLIYFATSMGTHTMAAHQYGVNLSRKLLNHLCLNCYMELIGICQRPGC 167
+ + +K Y L+Y+ S+ + +H Y ++ RKL H+ + ++L+ + + C
Sbjct: 623 YFYLETKTGSYKHLLYYFLSLTNFQLFSHSYNTHI-RKLQRHIIIIIIIKLLFLKETRSC 447
>TC224221 weakly similar to UP|Q8V7F2 (Q8V7F2) ORF1 (Fragment), partial (38%)
Length = 560
Score = 27.3 bits (59), Expect = 5.2
Identities = 12/27 (44%), Positives = 17/27 (62%)
Frame = +3
Query: 90 SIPSGREFITILGLAAPVFMTMMSKVA 116
++P G E + LA PV +TMM+K A
Sbjct: 54 TLPEGAEELRAHALAVPVMVTMMAKTA 134
>CO985879
Length = 695
Score = 27.3 bits (59), Expect = 5.2
Identities = 12/27 (44%), Positives = 17/27 (62%)
Frame = -3
Query: 90 SIPSGREFITILGLAAPVFMTMMSKVA 116
++P G E + LA PV +TMM+K A
Sbjct: 393 TLPEGAEELRAHALAVPVMVTMMAKTA 313
>BI785135
Length = 420
Score = 27.3 bits (59), Expect = 5.2
Identities = 14/47 (29%), Positives = 19/47 (39%)
Frame = -3
Query: 7 PAGLLYLCGWVAQSASLGMKDSWGPLKALAAASVINGVGDIVLCTYL 53
P +LYL W Q L SW PL + NG+ C ++
Sbjct: 283 PKFILYLVEWATQE*PLFS*GSWEPLITI*KIKKSNGIKHTFFCIFI 143
>TC225706
Length = 1302
Score = 26.9 bits (58), Expect = 6.9
Identities = 16/57 (28%), Positives = 23/57 (40%), Gaps = 6/57 (10%)
Frame = -3
Query: 6 IPAGLLYLCGWVAQSASLGMKDSWGPLKALAAASVINGVG------DIVLCTYLGYG 56
I A L L W+ M WGP + + +I+ +G V YLG+G
Sbjct: 340 IEASLPSLTAWIP*VTGKNMPPGWGPPETYSTRPLISHLGKAYLPKSFVFFRYLGFG 170
>AW308841 similar to PIR|JQ2128|JQ212 metallothionein - soybean, partial
(97%)
Length = 235
Score = 26.9 bits (58), Expect = 6.9
Identities = 11/26 (42%), Positives = 17/26 (65%)
Frame = -1
Query: 56 GIAGAAWATMASQVVAAYMMMRTLNM 81
GIAGA W+T A+ V++ + TL +
Sbjct: 223 GIAGAVWSTFAAIVLSGHTHFSTLEL 146
>TC230266 similar to UP|GRP1_PETHY (P09789) Glycine-rich cell wall structural
protein 1 precursor, partial (7%)
Length = 996
Score = 26.9 bits (58), Expect = 6.9
Identities = 19/47 (40%), Positives = 24/47 (50%), Gaps = 1/47 (2%)
Frame = -2
Query: 6 IPAGLLYLCGWVAQSASLGMKDSWGPLKALAAASVI-NGVGDIVLCT 51
IP G + + G+ SA L S K AAASV +GVG LC+
Sbjct: 167 IPGGRISIGGFFPASAKLVSLGSSREAKNAAAASVFCDGVGSTSLCS 27
>TC203991 UP|Q7M213 (Q7M213) Metallothionein, complete
Length = 697
Score = 26.6 bits (57), Expect = 8.9
Identities = 10/26 (38%), Positives = 17/26 (64%)
Frame = -3
Query: 56 GIAGAAWATMASQVVAAYMMMRTLNM 81
G+AGA W+T A+ V++ + TL +
Sbjct: 308 GVAGAVWSTFAAIVLSGHTHFSTLEL 231
>TC203989 homologue to UP|Q7M213 (Q7M213) Metallothionein, partial (72%)
Length = 725
Score = 26.6 bits (57), Expect = 8.9
Identities = 10/26 (38%), Positives = 17/26 (64%)
Frame = -2
Query: 56 GIAGAAWATMASQVVAAYMMMRTLNM 81
G+AGA W+T A+ V++ + TL +
Sbjct: 616 GVAGAVWSTFAAIVLSGHTHFSTLEL 539
Database: GMGI
Posted date: Oct 22, 2004 4:58 PM
Number of letters in database: 37,918,896
Number of sequences in database: 63,676
Lambda K H
0.338 0.146 0.479
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 10,129,569
Number of Sequences: 63676
Number of extensions: 147551
Number of successful extensions: 1046
Number of sequences better than 10.0: 24
Number of HSP's better than 10.0 without gapping: 1041
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 1045
length of query: 212
length of database: 12,639,632
effective HSP length: 93
effective length of query: 119
effective length of database: 6,717,764
effective search space: 799413916
effective search space used: 799413916
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 15 ( 7.3 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 39 (21.8 bits)
S2: 57 (26.6 bits)
Medicago: description of AC149293.2


