

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC149205.5 + phase: 0 /pseudo
(738 letters)
Database: GMGI
63,676 sequences; 37,918,896 total letters
Searching..................................................done
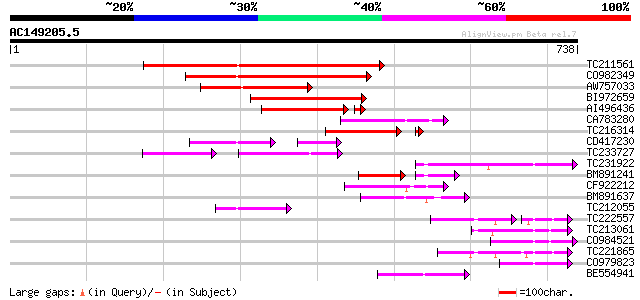
Score E
Sequences producing significant alignments: (bits) Value
TC211561 weakly similar to UP|O22675 (O22675) Reverse transcript... 302 4e-82
CO982349 209 4e-54
AW757033 136 4e-32
BI972659 129 6e-30
AI496436 100 2e-21
CA783280 weakly similar to PIR|T01871|T01 RNA-directed DNA polym... 100 3e-21
TC216314 similar to GB|AAP68269.1|31711826|BT008830 At1g70320 {A... 94 8e-20
CD417230 weakly similar to GP|2244915|emb| reverse transcriptase... 82 1e-19
TC233727 similar to UP|Q9FKH8 (Q9FKH8) Similarity to non-LTR ret... 67 4e-19
TC231922 88 2e-17
BM891241 71 6e-17
CF922212 83 5e-16
BM891637 77 2e-14
TC212055 73 5e-13
TC222557 45 3e-11
TC213061 67 4e-11
CO984521 65 8e-11
TC221865 65 1e-10
CO979823 64 2e-10
BE554941 60 3e-09
>TC211561 weakly similar to UP|O22675 (O22675) Reverse transcriptase
(Fragment) , partial (59%)
Length = 1077
Score = 302 bits (773), Expect = 4e-82
Identities = 150/313 (47%), Positives = 205/313 (64%)
Frame = +1
Query: 175 LDGILIANEVVDEAKKTKKELLLFKVDFEKAYDSVD*GYLDAVMQKMAFPLLWRKWIKEC 234
L LIANEV+DEAK++ K L+FKVD+EKAYDSV +L +M +M F W KWI+EC
Sbjct: 1 LHSALIANEVIDEAKRSNKSCLVFKVDYEKAYDSVSWNFLMYMMWRMDFSPKWIKWIEEC 180
Query: 235 ISTTTASILVNGSPTDEFSLHRGLRQGDPLSPFLFMIAAEGLNVLMKSVVENNLFKGYVV 294
+ + + S+LVNGSPT EF RGLRQGDPL+PFLF I AEGLN LM+ VE NL+K Y+V
Sbjct: 181 VKSASISVLVNGSPTAEFLPQRGLRQGDPLTPFLFNIVAEGLNGLMRRAVEENLYKPYLV 360
Query: 295 GASNSVVVSHLQFADDTLLLGEKSWANVRALRAVLTLFAEMSGLKVNFNKSLLVGVNIGE 354
GA N V +S LQ+ADDT+ GE + NV A++ +L F +S L++NF KS + +
Sbjct: 361 GA-NGVPISILQYADDTIFFGEAAKENVEAIKVILRSFELVSNLRINFAKSCFGVFGVTD 537
Query: 355 SWLLEAASVLNCKVGKILFLYLGLSIGGDPRKLAFWEPVVSTIKTRLSGWQSCFLSFGGR 414
W EAA+ L+C + F+YLG+ IG + R+ W+ ++ + +LS W+ +SFGGR
Sbjct: 538 QWKQEAANYLHCSLLAFPFIYLGIPIGANSRRCHMWDSIIRKCERKLSKWK*RHISFGGR 717
Query: 415 LVLLKSVLTSLPVYALSFFKSPSGIISLIESLFNKFFWRGSEDNRKISWIAWSSVCLKKE 474
+ L+KSVLTS+P+Y SFF+ P ++ + + F W G D +KISWI W +CL KE
Sbjct: 718 VTLIKSVLTSIPIYFFSFFRVPQSVVDKLVKI*RTFLWGGGPDQKKISWIRWEKLCLPKE 897
Query: 475 FGGLGVRLLREFN 487
GG+ RL N
Sbjct: 898 RGGI*KRLPNNHN 936
>CO982349
Length = 795
Score = 209 bits (532), Expect = 4e-54
Identities = 100/242 (41%), Positives = 151/242 (62%)
Frame = +3
Query: 230 WIKECISTTTASILVNGSPTDEFSLHRGLRQGDPLSPFLFMIAAEGLNVLMKSVVENNLF 289
WI+ C+ + + SILVN SP +EFS RGLRQGDPL+P LF I AEGL LM+ V+ F
Sbjct: 69 WIRGCLHSASVSILVNESPINEFSPQRGLRQGDPLAPLLFNIVAEGLTGLMREAVDRKRF 248
Query: 290 KGYVVGASNSVVVSHLQFADDTLLLGEKSWANVRALRAVLTLFAEMSGLKVNFNKSLLVG 349
++VG N VS LQ+ADDT+ E + N++ ++ +L F SGLK+NF +S
Sbjct: 249 NSFLVG-KNKEPVSILQYADDTIFFEEATMENMKVIKTILRCFELASGLKINFAESRFGA 425
Query: 350 VNIGESWLLEAASVLNCKVGKILFLYLGLSIGGDPRKLAFWEPVVSTIKTRLSGWQSCFL 409
+ + W EAA LNC + + F YLG+ I +PR +P++ +T+L+ W+ +
Sbjct: 426 IWKPDHWCKEAAEFLNCSMLSMPFSYLGIPIEANPRCREI*DPIIRKCETKLARWKQRHI 605
Query: 410 SFGGRLVLLKSVLTSLPVYALSFFKSPSGIISLIESLFNKFFWRGSEDNRKISWIAWSSV 469
S GGR+ L+ ++LT+LP+Y SFF++PS ++ + ++ KF W G R+I+W+ W +V
Sbjct: 606 SLGGRVTLINAILTALPIYFFSFFRAPSKVVQKLVNIQRKFLWGGGSXQRRIAWVKWETV 785
Query: 470 CL 471
CL
Sbjct: 786 CL 791
>AW757033
Length = 441
Score = 136 bits (342), Expect = 4e-32
Identities = 69/146 (47%), Positives = 93/146 (63%)
Frame = -2
Query: 249 TDEFSLHRGLRQGDPLSPFLFMIAAEGLNVLMKSVVENNLFKGYVVGASNSVVVSHLQFA 308
T EFS HRGLRQGDPL+P LF IAAEGL LM+ V N +K + VG V S LQ+A
Sbjct: 440 TREFSPHRGLRQGDPLAPLLFNIAAEGLTGLMREAVARNHYKSFAVGKYKKPV-SILQYA 264
Query: 309 DDTLLLGEKSWANVRALRAVLTLFAEMSGLKVNFNKSLLVGVNIGESWLLEAASVLNCKV 368
DDT+ GE + NVR ++ +L F SGLK+NF KS +++ + W EAA +NC +
Sbjct: 263 DDTIFFGEATMENVRVIKTILRGFELASGLKINFAKSRFGAISVPD*WRKEAAEFMNCSL 84
Query: 369 GKILFLYLGLSIGGDPRKLAFWEPVV 394
+ F YLG+ IG +PR+ W+P++
Sbjct: 83 LSLPFSYLGIPIGANPRRRETWDPII 6
>BI972659
Length = 453
Score = 129 bits (323), Expect = 6e-30
Identities = 62/151 (41%), Positives = 92/151 (60%)
Frame = +1
Query: 314 LGEKSWANVRALRAVLTLFAEMSGLKVNFNKSLLVGVNIGESWLLEAASVLNCKVGKILF 373
+GE SW N+ L+++L F +SGL++N+ KS V +W +AA +LNC+ I F
Sbjct: 1 VGEASWDNIIVLKSMLRGFEMVSGLRINYAKSQFGIVGFQPNWAHDAAQLLNCRQLDIPF 180
Query: 374 LYLGLSIGGDPRKLAFWEPVVSTIKTRLSGWQSCFLSFGGRLVLLKSVLTSLPVYALSFF 433
YLG+ I WEP+++ K +LS W +LS GGR+ L+KSVL +LP+Y LSFF
Sbjct: 181 HYLGMPIAVKASSRMVWEPLINKFKAKLSKWNQKYLSMGGRVTLIKSVLNALPIYLLSFF 360
Query: 434 KSPSGIISLIESLFNKFFWRGSEDNRKISWI 464
K P I+ + SL F W G++ + +ISW+
Sbjct: 361 KIPQRIVDKLVSLQRTFMWGGNQHHNRISWV 453
>AI496436
Length = 414
Score = 100 bits (248), Expect(2) = 2e-21
Identities = 49/113 (43%), Positives = 70/113 (61%)
Frame = +1
Query: 328 VLTLFAEMSGLKVNFNKSLLVGVNIGESWLLEAASVLNCKVGKILFLYLGLSIGGDPRKL 387
+L F SGLK+NF KS + + W EAA LN V + F+YLG+ IG + R
Sbjct: 4 ILRCFELSSGLKINFAKSQFGTICKSDIWRKEAADYLNSSVLSMPFVYLGIPIGANSRHS 183
Query: 388 AFWEPVVSTIKTRLSGWQSCFLSFGGRLVLLKSVLTSLPVYALSFFKSPSGII 440
WEPVV + +L+ W+ ++SFGGR+ L+ SVLT+LP+Y LSFF+ P ++
Sbjct: 184 DVWEPVVRKCERKLASWKQKYVSFGGRVTLINSVLTALPIYLLSFFRIPDKVV 342
Score = 21.6 bits (44), Expect(2) = 2e-21
Identities = 7/15 (46%), Positives = 10/15 (66%)
Frame = +2
Query: 449 KFFWRGSEDNRKISW 463
+F W G + RKI+W
Sbjct: 368 RFLWGGDLE*RKIAW 412
>CA783280 weakly similar to PIR|T01871|T01 RNA-directed DNA polymerase
homolog T24M8.8 - Arabidopsis thaliana, partial (4%)
Length = 456
Score = 100 bits (248), Expect = 3e-21
Identities = 57/143 (39%), Positives = 76/143 (52%), Gaps = 2/143 (1%)
Frame = +2
Query: 431 SFFKSPSGIISLIESLFNKFFWRGSEDNRKISWIAWSSVCLKKEFGGLGVRLLREFNLAL 490
SFF P II+ +ESL +F W G D+RKI+W+ W +VCL K GGLG++ LR FN L
Sbjct: 5 SFFSLP*KIIAKLESLQRRFLWGGEADSRKIAWVNWKTVCLPKAKGGLGIKDLRTFNTTL 184
Query: 491 MSKWCWRLLIDRGGFWYRVLVARYGEEARRLEVG--GRSVSSWWKEVAKIKDGVGEDDEG 548
+ KW W L + W +VL ++YG R LE G G S+WWK++ K +
Sbjct: 185 LGKWRWDLFYIQQEPWAKVLQSKYG-GWRALEEGSSGSKDSAWWKDLIKTQQLQRNIP-- 355
Query: 549 WFVGKVSKLVGDGSNTLFWFDRW 571
+ VG G FW D W
Sbjct: 356 -LKRETIWKVGGGDRIKFWEDLW 421
>TC216314 similar to GB|AAP68269.1|31711826|BT008830 At1g70320 {Arabidopsis
thaliana;} , partial (18%)
Length = 759
Score = 94.4 bits (233), Expect(2) = 8e-20
Identities = 40/98 (40%), Positives = 61/98 (61%)
Frame = +2
Query: 412 GGRLVLLKSVLTSLPVYALSFFKSPSGIISLIESLFNKFFWRGSEDNRKISWIAWSSVCL 471
GGR+ L+KSVL++LP+ LSFFK P I+ + +L +F W G++ + +I W+ W+ +C
Sbjct: 5 GGRVTLIKSVLSALPIXXLSFFKIPQRIVDKLVTLQRQFLWGGTQHHNRIPWVKWADICN 184
Query: 472 KKEFGGLGVRLLREFNLALMSKWCWRLLIDRGGFWYRV 509
K GGLG++ L FN AL +W W L + W R+
Sbjct: 185 PKIDGGLGIKDLSNFNAALRGRWIWGLASNHNQLWARL 298
Score = 21.6 bits (44), Expect(2) = 8e-20
Identities = 6/10 (60%), Positives = 9/10 (90%)
Frame = +3
Query: 529 SSWWKEVAKI 538
SSWWK++ K+
Sbjct: 327 SSWWKDLKKL 356
>CD417230 weakly similar to GP|2244915|emb| reverse transcriptase like
protein {Arabidopsis thaliana}, partial (7%)
Length = 688
Score = 82.0 bits (201), Expect(2) = 1e-19
Identities = 44/112 (39%), Positives = 67/112 (59%)
Frame = -3
Query: 234 CISTTTASILVNGSPTDEFSLHRGLRQGDPLSPFLFMIAAEGLNVLMKSVVENNLFKGYV 293
C+STTT S+L NG D FS + G+RQ DP++P+LF++ E L+ L+ V+ L +
Sbjct: 668 CMSTTTMSLLWNGEVLDHFSPNIGVRQRDPIAPYLFVLCLERLSHLINLVMNKKL*RSIQ 489
Query: 294 VGASNSVVVSHLQFADDTLLLGEKSWANVRALRAVLTLFAEMSGLKVNFNKS 345
V +++SHL F DD +L E + V + L LF++ SG KVN +K+
Sbjct: 488 VN-RGGLLISHLAFVDDLILFVEANMDQVDVINLTLELFSDSSGAKVNVDKT 336
Score = 33.1 bits (74), Expect(2) = 1e-19
Identities = 19/58 (32%), Positives = 30/58 (50%)
Frame = -2
Query: 375 YLGLSIGGDPRKLAFWEPVVSTIKTRLSGWQSCFLSFGGRLVLLKSVLTSLPVYALSF 432
YLG+ I ++ ++ + R S W+S LS GRL L K V+ +LP + + F
Sbjct: 243 YLGVPIFHKNPTSHTFDFIIDKVNKRFSRWKSNLLSMVGRLTLTK*VV*ALPSHIMQF 70
>TC233727 similar to UP|Q9FKH8 (Q9FKH8) Similarity to non-LTR retroelement
reverse transcriptase, partial (7%)
Length = 956
Score = 66.6 bits (161), Expect(2) = 4e-19
Identities = 37/97 (38%), Positives = 55/97 (56%), Gaps = 1/97 (1%)
Frame = -3
Query: 174 ILDGILIANEVVDEA-KKTKKELLLFKVDFEKAYDSVD*GYLDAVMQKMAFPLLWRKWIK 232
I D + +EV++ KK L K+D KA+D++D +L +V+ K + + WI+
Sbjct: 834 IKDCTCVTSEVINMLDKKVFGGNLALKIDIVKAFDTLDRDFLISVLNKFGYSHTFCNWIR 655
Query: 233 ECISTTTASILVNGSPTDEFSLHRGLRQGDPLSPFLF 269
+ + SI VNG P FS RG+RQGDPLSP L+
Sbjct: 654 VILLSAKLSISVNGEPVGFFSCQRGVRQGDPLSPSLY 544
Score = 47.0 bits (110), Expect(2) = 4e-19
Identities = 37/136 (27%), Positives = 65/136 (47%), Gaps = 1/136 (0%)
Frame = -2
Query: 299 SVVVSHLQFADDTLLLGEKSWANVRALRAVLTLFAEMSGLKVNFNKSLLVGVNIGESWLL 358
S+ SH+ ADD ++ + + NV L L+ + G ++ +KS + +I L
Sbjct: 463 SLTRSHVMHADDIMIFCKGTSRNVHNLINFFQLYKDSFGQVISKHKSKIFIGSIPGQRLR 284
Query: 359 EAASVLNCKVGKILFLYLGLSI-GGDPRKLAFWEPVVSTIKTRLSGWQSCFLSFGGRLVL 417
+ +L I F Y G+ I G P + + V IK +L+ W+ LS GR+ L
Sbjct: 283 VISDLLQYFHSDIPFNYFGVPIFKGKPTRRHLY-CVADKIKCKLASWKGSILSIMGRIQL 107
Query: 418 LKSVLTSLPVYALSFF 433
+ SV+ + +Y+ S +
Sbjct: 106 VNSVIHGMLLYSFSVY 59
>TC231922
Length = 803
Score = 87.8 bits (216), Expect = 2e-17
Identities = 56/216 (25%), Positives = 103/216 (46%), Gaps = 6/216 (2%)
Frame = -3
Query: 529 SSWWKEVAKIKDGVGEDDEGWFVGKVSKLVGDGSNTLFWFDRWLGD-VPFNRRFARLFDL 587
S WWK++ ++ + + D + VG G FW D WL + +++ +L+ +
Sbjct: 777 SHWWKDLRRLYN---QPDFHSIHQNMVWRVGCGDKIKFWQDSWLSEGCNLQQKYNQLYTI 607
Query: 588 TTNKMCTVADMYRLGWEEGGDAWVWRRRLWAWEE----ECRSLLDNVRLRLNVLDRWQWD 643
+ T++ M + + WRRRL+ +E + + + + ++ D W
Sbjct: 606 SRQ*NLTISKMGKFSQNARSWEFKWRRRLFDYEYAMAVDFMNEISGISIQNQAHDSMFWK 427
Query: 644 PDTIDVYTVRGVYQILTTSVSNTIVKTD-ELVWHKQVPLN*VSIVAWRLLKDRLPTKTNL 702
D+ VY+ + Y++L S+S + +++WH ++P ++ +WRL DRLPT+ NL
Sbjct: 426 ADSSGVYSTKSAYRLLMPSISPAPSRRIFQILWHLKIPPR-AAVFSWRLFLDRLPTRGNL 250
Query: 703 QRRDCLQATVNTCVSGCGIAESASHLFLHCEVFSSL 738
RR + + GC E A HLF HC++ L
Sbjct: 249 SRRSIPIQDIMCPLCGCQ-HEEAGHLFFHCKMTKGL 145
>BM891241
Length = 407
Score = 70.9 bits (172), Expect(2) = 6e-17
Identities = 27/62 (43%), Positives = 41/62 (65%)
Frame = -2
Query: 454 GSEDNRKISWIAWSSVCLKKEFGGLGVRLLREFNLALMSKWCWRLLIDRGGFWYRVLVAR 513
G D++KI W+ W VCL K GGLG++ + +FN+AL+ KW W L D+ W R++ ++
Sbjct: 403 GDHDHKKIPWVKWEVVCLPKTKGGLGIKDISKFNIALLGKWIWALASDQQQLWARIINSK 224
Query: 514 YG 515
YG
Sbjct: 223 YG 218
Score = 35.4 bits (80), Expect(2) = 6e-17
Identities = 19/58 (32%), Positives = 29/58 (49%), Gaps = 1/58 (1%)
Frame = -3
Query: 529 SSWWKEVAKIKDGVGEDDEGWFVGKVSKLVGDGSNTLFWFDRWLG-DVPFNRRFARLF 585
S WW+++ K + D F +S VG G FW D+WLG D +++ +LF
Sbjct: 174 SYWWRDLRKFYH---QSDHSIFHQYMSWKVGCGDKINFWTDKWLGEDYTLEQKYNQLF 10
>CF922212
Length = 445
Score = 82.8 bits (203), Expect = 5e-16
Identities = 50/142 (35%), Positives = 73/142 (51%), Gaps = 6/142 (4%)
Frame = -2
Query: 436 PSGIISLIESLFNKFFWRGSEDNRKISWIAWSSVCLKKEFGGLGVRLLREFNLALMSKWC 495
P+ ++ + L F W G +KI+W+ W SVCL KE GLG R LR+FN AL+ K
Sbjct: 438 PNKVLDKLVRLQQWFLWGGGLGQKKIAWVNWDSVCLPKEKEGLGNRDLRKFNYALLGKRR 259
Query: 496 WRLLIDRGGFWYRVLVARYG-----EEARRLEVGGRSVSSWWKEVAKIKDGVGEDDEGWF 550
W L +G RVL ++Y +E RR+ +S S WW+E++ I +D WF
Sbjct: 258 WNLFHHQGELGARVLDSKYKRWRNLDEERRV----KSESFWWQEISFITHST--EDGSWF 97
Query: 551 VGKV-SKLVGDGSNTLFWFDRW 571
+ + +G + FW D W
Sbjct: 96 EKRTKTGRLGCRAKVRFWEDGW 31
>BM891637
Length = 427
Score = 77.4 bits (189), Expect = 2e-14
Identities = 44/149 (29%), Positives = 73/149 (48%), Gaps = 7/149 (4%)
Frame = -1
Query: 457 DNRKISWIAWSSVCLKKEFGGLGVRLLREFNLALMSKWCWRLLIDRGGFWYRVLVARYGE 516
+ +KI+WI+W C + GGLG++ ++ N AL+ KW W + W R+L+++Y +
Sbjct: 424 EGKKIAWISWRQCCASGDVGGLGIKDIKILNNALLIKWKWLMFHQPHQLWNRILISKY-K 248
Query: 517 EARRLEVGGRS--VSSWWKEVAKIKDG----VGEDDEGWFVGKVSKLVGDGSNTLFWFDR 570
R L+ G + S WW ++ I + W VG+ G LFW D
Sbjct: 247 GWRGLDQGPQKYYFSPWWADLRAINQHQSMIAASNQFCWKVGR-------GDQILFWEDS 89
Query: 571 WLGD-VPFNRRFARLFDLTTNKMCTVADM 598
W+ D P +F L+ L++ + +ADM
Sbjct: 88 WVDDGTPLKDQFPELYRLSSQRNFIMADM 2
>TC212055
Length = 776
Score = 72.8 bits (177), Expect = 5e-13
Identities = 39/101 (38%), Positives = 61/101 (59%), Gaps = 1/101 (0%)
Frame = +1
Query: 268 LFMIAAEGLNVLMKSVVENNLFKGYVVGASNSVVVSHLQFADDTLLLGEKSWANVRALRA 327
LF I AEGL LM+ ++ +L+ +VG N + V+ LQ+ADDT+ LGE + NV +++
Sbjct: 1 LFNIVAEGLTGLMREALDKSLYSSLMVG-KNKIPVNILQYADDTIFLGEATMQNVMTIKS 177
Query: 328 VLTLFAEMSGLKVNFNKSLLVGVNIGESWLLEAA-SVLNCK 367
+L +F SGLK++F KS + + W EA + NC+
Sbjct: 178 MLRVFELASGLKIHFAKSSFGAIGKPDHWQKEAGLNTFNCR 300
>TC222557
Length = 1002
Score = 45.1 bits (105), Expect(2) = 3e-11
Identities = 27/69 (39%), Positives = 39/69 (56%), Gaps = 3/69 (4%)
Frame = +1
Query: 667 IVKTDELV---WHKQVPLN*VSIVAWRLLKDRLPTKTNLQRRDCLQATVNTCVSGCGIAE 723
+++ DE + W ++PL + AWRL+K+R+PTK NL RR +Q C E
Sbjct: 409 LIRLDEALEDLWQLKIPLK-ATTFAWRLIKERIPTKGNLWRRR-VQLNNLMCPFCNRQEE 582
Query: 724 SASHLFLHC 732
ASHLF +C
Sbjct: 583 EASHLFFNC 609
Score = 41.6 bits (96), Expect(2) = 3e-11
Identities = 34/122 (27%), Positives = 54/122 (43%), Gaps = 10/122 (8%)
Frame = +3
Query: 548 GWFVGKVSKLVGDGSNTLFWFDRWLGDV-PFNRRFARLFDLTTN--KMCTVADMYRLG-W 603
G F + VG G FW D WL + P ++ RL++++ K+ V + W
Sbjct: 45 GLFQSAIDWKVGCGDKIRFWEDCWLMEQEPLRAKYPRLYNISCQQQKLILVMGFHSANVW 224
Query: 604 EEGGDAWV--WRRRLWAWE-EECRSLLDNV---RLRLNVLDRWQWDPDTIDVYTVRGVYQ 657
E W WRR L+ E + S LD++ + LD W W P+ ++ R Y+
Sbjct: 225 E-----WKLEWRRHLFDNEVQAAASFLDDISWGHIDRWTLDCWVWKPEPNGQFSTRSAYR 389
Query: 658 IL 659
+L
Sbjct: 390 ML 395
>TC213061
Length = 823
Score = 66.6 bits (161), Expect = 4e-11
Identities = 44/136 (32%), Positives = 70/136 (51%), Gaps = 5/136 (3%)
Frame = +2
Query: 602 GWEEGGDAWVWRRRLWAWEEECRSL----LDNVRLRLNVLDRWQWDPDTIDVYTVRGVYQ 657
GWE + WRR L+ E E ++ ++R ++ D+W W ++ Y R Y+
Sbjct: 17 GWEW---QFQWRRPLFDSEIEMLVAFIQEVEGHKIRSDIEDQWVWAAESSGSYLARSAYR 187
Query: 658 ILTTSVSNTIVKTD-ELVWHKQVPLN*VSIVAWRLLKDRLPTKTNLQRRDCLQATVNTCV 716
++ + + + +W +VP+ V++ AWRLLKD LPT+ NL RR ++ C
Sbjct: 188 VIREGIPEEEQDREFKELWKLKVPMK-VTMFAWRLLKDILPTRDNL-RRKRVELHEYVCP 361
Query: 717 SGCGIAESASHLFLHC 732
+ ESASHLF HC
Sbjct: 362 LCRSMDESASHLFFHC 409
>CO984521
Length = 716
Score = 65.5 bits (158), Expect = 8e-11
Identities = 41/114 (35%), Positives = 60/114 (51%), Gaps = 2/114 (1%)
Frame = -2
Query: 627 LDNVRLRLNVLDRWQWDPDTIDVYTVRGVYQIL--TTSVSNTIVKTDELVWHKQVPLN*V 684
L+ +R + D+W+W + Y+ + Y+ L T K EL W +VPL V
Sbjct: 679 LEGFTIRPELSDQWKWAAEPSGCYSTKSAYKALHHVTVGEEQDGKFKEL-WKLRVPLK-V 506
Query: 685 SIVAWRLLKDRLPTKTNLQRRDCLQATVNTCVSGCGIAESASHLFLHCEVFSSL 738
+I AWRL++D+LPTK NL R+ ++ C + E+ASHLF HC S L
Sbjct: 505 AIFAWRLIQDKLPTKANL-RKKRVELQEYLCPLCRSVEETASHLFFHCSKVSPL 347
>TC221865
Length = 1045
Score = 65.1 bits (157), Expect = 1e-10
Identities = 52/186 (27%), Positives = 86/186 (45%), Gaps = 11/186 (5%)
Frame = -3
Query: 558 VGDGSNTLFWFDRWLGD-VPFNRRFARLFDLTTNKMCTVADM---YRLGWEEGGDAWVWR 613
VG G FW D+ +G+ R++ R++ ++ + + + WE + WR
Sbjct: 923 VGCGDKFRFWEDKLVGEGETLMRKYPRMYQISCQQQQLIQQVGSHTETTWEW---KFQWR 753
Query: 614 RRLWAWEEECR-SLLDNVR---LRLNVLDRWQWDPDTIDVYTVRGVYQILTTSVSNTIVK 669
R L+ E + L+++ + ++ D W W + Y+ R Y ++ +
Sbjct: 752 RPLFDNEVDTTIGFLEDISRFPIHRHLTDCWVWKSEPNGHYSTRSAYHLMQHEAAKA--N 579
Query: 670 TD---ELVWHKQVPLN*VSIVAWRLLKDRLPTKTNLQRRDCLQATVNTCVSGCGIAESAS 726
TD E +W ++P +I A RL+KDRLPTK NL+RR +Q C+ E AS
Sbjct: 578 TDPAFEEIWKLKIPAK-AAIFA*RLVKDRLPTKNNLRRRQ-VQLNGTLCLFCRNYEEEAS 405
Query: 727 HLFLHC 732
HLF C
Sbjct: 404 HLFFSC 387
>CO979823
Length = 853
Score = 64.3 bits (155), Expect = 2e-10
Identities = 33/96 (34%), Positives = 53/96 (54%), Gaps = 1/96 (1%)
Frame = -2
Query: 638 DRWQWDPDTIDVYTVRGVYQILTTSVSNTIVKTDEL-VWHKQVPLN*VSIVAWRLLKDRL 696
D+W W P+ Y+ + Y +L ++ I D +W ++P ++ AWRL++DRL
Sbjct: 702 DKWLWKPEPGGHYSTKSGYHVLWGELTEEIQDADFAEIWKLKIPTK-AAVFAWRLVRDRL 526
Query: 697 PTKTNLQRRDCLQATVNTCVSGCGIAESASHLFLHC 732
PTK+NL+RR + + C I E A+HLF +C
Sbjct: 525 PTKSNLRRRQVMVQDM-VCPLCNNIEEGAAHLFFNC 421
>BE554941
Length = 383
Score = 60.5 bits (145), Expect = 3e-09
Identities = 34/120 (28%), Positives = 54/120 (44%)
Frame = +1
Query: 479 GVRLLREFNLALMSKWCWRLLIDRGGFWYRVLVARYGEEARRLEVGGRSVSSWWKEVAKI 538
G++ L FN++L+ K L++ W +L YG S WW+++ +I
Sbjct: 7 GIKNLSIFNISLLVKRR*HFLVETKAIWAPILKFHYGSLDLSNTGSASRTSQWWRDILRI 186
Query: 539 KDGVGEDDEGWFVGKVSKLVGDGSNTLFWFDRWLGDVPFNRRFARLFDLTTNKMCTVADM 598
G + GW + +GD +TLFW R L + P + ARL L +K +V M
Sbjct: 187 --GNCQYQPGWLTSTFCRKIGDRKDTLFWRHRCLNETPLMQSHARLXSLAMDKDISVQGM 360
Database: GMGI
Posted date: Oct 22, 2004 4:58 PM
Number of letters in database: 37,918,896
Number of sequences in database: 63,676
Lambda K H
0.335 0.146 0.498
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 39,594,451
Number of Sequences: 63676
Number of extensions: 630517
Number of successful extensions: 5610
Number of sequences better than 10.0: 106
Number of HSP's better than 10.0 without gapping: 5407
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 5561
length of query: 738
length of database: 12,639,632
effective HSP length: 104
effective length of query: 634
effective length of database: 6,017,328
effective search space: 3814985952
effective search space used: 3814985952
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 15 ( 7.2 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 39 (21.6 bits)
S2: 63 (28.9 bits)
Medicago: description of AC149205.5


