

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
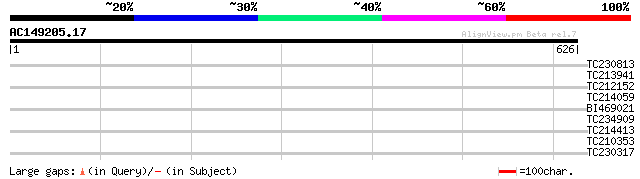
Query= AC149205.17 + phase: 0 /pseudo
(626 letters)
Database: GMGI
63,676 sequences; 37,918,896 total letters
Searching..................................................done
Score E
Sequences producing significant alignments: (bits) Value
TC230813 30 2.5
TC213941 similar to GB|AAO39924.1|28372884|BT003696 At4g04630 {A... 29 5.5
TC212152 29 5.5
TC214059 homologue to UP|Q93YG3 (Q93YG3) Chlorophyll a/b binding... 29 5.5
BI469021 similar to GP|9759176|dbj| contains similarity to patho... 29 5.5
TC234909 similar to UP|Q84ND9 (Q84ND9) PolI-like DNA polymerase,... 28 9.4
TC214413 similar to UP|Q93YG3 (Q93YG3) Chlorophyll a/b binding p... 28 9.4
TC210353 28 9.4
TC230317 25 9.6
>TC230813
Length = 1130
Score = 30.4 bits (67), Expect = 2.5
Identities = 20/72 (27%), Positives = 31/72 (42%)
Frame = +2
Query: 179 LKAKR*FQKVWPLPHQCGWIFSSIKVVKLNMKHFLLLGCQFVFPHRYNVVKSSVSYCCSS 238
L R ++ W L CG++ + + H LLL +FP Y + +SYCC
Sbjct: 440 LNPLRIMKQFWRLIRICGFLMPLTGLQFEDQLHILLLS--IIFPPLYFMPVPRISYCC-- 607
Query: 239 C*RDSYGFGTCC 250
++ G CC
Sbjct: 608 ---ETDLLGLCC 634
>TC213941 similar to GB|AAO39924.1|28372884|BT003696 At4g04630 {Arabidopsis
thaliana;} , partial (40%)
Length = 776
Score = 29.3 bits (64), Expect = 5.5
Identities = 16/57 (28%), Positives = 28/57 (49%), Gaps = 1/57 (1%)
Frame = +2
Query: 467 NFEPRKRSASSSKCEPSAKIPPEFPS-HKLVGCTVTIGKSSDDGSNTSQGDKIVDDD 522
N P ++ + + S+ P + P K+ G + G ++DDG++ GD DDD
Sbjct: 317 NANPLPAASDAPLVKGSSSAPMDIPDWSKIYGKSCKKGSTADDGASNKGGDDDDDDD 487
>TC212152
Length = 648
Score = 29.3 bits (64), Expect = 5.5
Identities = 24/84 (28%), Positives = 36/84 (42%)
Frame = -3
Query: 470 PRKRSASSSKCEPSAKIPPEFPSHKLVGCTVTIGKSSDDGSNTSQGDKIVDDDAPSGSIP 529
P + ASSSK + + P F S L I S+ GS TS+ V D+
Sbjct: 505 PVRTRASSSKIACNVLVSPRFSSRMLSRS*TAITGSTSQGSGTSRFMVRVLDNG------ 344
Query: 530 KLLKTMSFEKSVEDGLLAEKYVDG 553
+L +F +E+ L + +DG
Sbjct: 343 -MLAKNAFGPLIEESALLKSSIDG 275
>TC214059 homologue to UP|Q93YG3 (Q93YG3) Chlorophyll a/b binding protein
type II precursor, complete
Length = 1079
Score = 29.3 bits (64), Expect = 5.5
Identities = 19/60 (31%), Positives = 24/60 (39%), Gaps = 5/60 (8%)
Frame = +1
Query: 140 NNHFGRCHCFGGILSLWSSCFSPNLKIRRLERWNRSSSALKAKR*FQ-----KVWPLPHQ 194
NNH C+ I WS+C +R +RW R S A + VWP P Q
Sbjct: 94 NNHHHGHLCYSAISIHWSNCSEAAQ*VRPQDRWRRQRSH*HAPHRQECSSEHLVWP*PSQ 273
>BI469021 similar to GP|9759176|dbj| contains similarity to
pathogenesis-related genes transcriptional
activator~gene_id:K19E1.9, partial (11%)
Length = 417
Score = 29.3 bits (64), Expect = 5.5
Identities = 16/45 (35%), Positives = 24/45 (52%)
Frame = +3
Query: 470 PRKRSASSSKCEPSAKIPPEFPSHKLVGCTVTIGKSSDDGSNTSQ 514
P K + S PS + PP+ P +LV TVT +++D S+ Q
Sbjct: 78 PSKYTHHSKLLIPSEEDPPKHPYPRLVRITVTDTEATDSSSDEEQ 212
>TC234909 similar to UP|Q84ND9 (Q84ND9) PolI-like DNA polymerase, partial
(4%)
Length = 658
Score = 28.5 bits (62), Expect = 9.4
Identities = 16/58 (27%), Positives = 29/58 (49%), Gaps = 7/58 (12%)
Frame = +2
Query: 185 FQKVWPLPHQCGWIFSSIKVV-----KLNMKHFLLLGC--QFVFPHRYNVVKSSVSYC 235
F + +P C W+ +I ++ K+N+ FLL+ C FV H + +S+ + C
Sbjct: 470 FHALHHVPVCCFWLLQTISILYINYYKMNVSAFLLICCNISFVETHAMTIGRSNRALC 643
>TC214413 similar to UP|Q93YG3 (Q93YG3) Chlorophyll a/b binding protein type
II precursor, partial (49%)
Length = 800
Score = 28.5 bits (62), Expect = 9.4
Identities = 19/60 (31%), Positives = 25/60 (41%), Gaps = 5/60 (8%)
Frame = +2
Query: 140 NNHFGRCHCFGGILSLWSSCFSPNLKIRRLERWNRSSSALKAKR*FQ-----KVWPLPHQ 194
+NH G C+ I WS+C +R +RW R S A + VWP P Q
Sbjct: 377 HNHHGHL-CYSAISIHWSNCSEAAQ*VRPQDRWRRQRSH*HAPHRQECSSEHVVWP*PSQ 553
>TC210353
Length = 438
Score = 28.5 bits (62), Expect = 9.4
Identities = 12/33 (36%), Positives = 19/33 (57%)
Frame = -1
Query: 189 WPLPHQCGWIFSSIKVVKLNMKHFLLLGCQFVF 221
WPL C W F ++ VV++ + FLLL ++
Sbjct: 153 WPLL*ACKW*FYNVVVVRVELWKFLLLVASAIY 55
>TC230317
Length = 1368
Score = 24.6 bits (52), Expect(2) = 9.6
Identities = 11/26 (42%), Positives = 14/26 (53%)
Frame = +3
Query: 196 GWIFSSIKVVKLNMKHFLLLGCQFVF 221
GW SS+ + FLLL C F+F
Sbjct: 885 GWRVSSLDKTAYGVLCFLLLSCLFLF 962
Score = 21.9 bits (45), Expect(2) = 9.6
Identities = 6/9 (66%), Positives = 8/9 (88%)
Frame = +2
Query: 145 RCHCFGGIL 153
RCHC GG++
Sbjct: 728 RCHCVGGLV 754
Database: GMGI
Posted date: Oct 22, 2004 4:58 PM
Number of letters in database: 37,918,896
Number of sequences in database: 63,676
Lambda K H
0.350 0.153 0.557
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 32,860,883
Number of Sequences: 63676
Number of extensions: 552194
Number of successful extensions: 5056
Number of sequences better than 10.0: 18
Number of HSP's better than 10.0 without gapping: 3711
Number of HSP's successfully gapped in prelim test: 158
Number of HSP's that attempted gapping in prelim test: 1300
Number of HSP's gapped (non-prelim): 3957
length of query: 626
length of database: 12,639,632
effective HSP length: 103
effective length of query: 523
effective length of database: 6,081,004
effective search space: 3180365092
effective search space used: 3180365092
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 14 ( 7.1 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 38 (21.9 bits)
S2: 62 (28.5 bits)
Medicago: description of AC149205.17


