

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
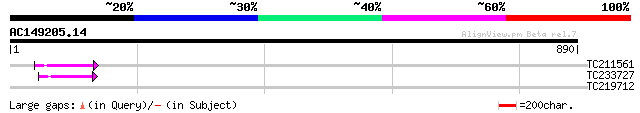
Query= AC149205.14 - phase: 0 /pseudo
(890 letters)
Database: GMGI
63,676 sequences; 37,918,896 total letters
Searching..................................................done
Score E
Sequences producing significant alignments: (bits) Value
TC211561 weakly similar to UP|O22675 (O22675) Reverse transcript... 71 2e-12
TC233727 similar to UP|Q9FKH8 (Q9FKH8) Similarity to non-LTR ret... 66 8e-11
TC219712 similar to UP|G6PI_SPIOL (O82059) Glucose-6-phosphate i... 30 4.7
>TC211561 weakly similar to UP|O22675 (O22675) Reverse transcriptase
(Fragment) , partial (59%)
Length = 1077
Score = 70.9 bits (172), Expect = 2e-12
Identities = 37/100 (37%), Positives = 57/100 (57%)
Frame = +1
Query: 40 LNNAMVAIEVIHFMKTKTRGQDKYVALKLDISKAYDRMDWDYLKEIMIKMGFSQKWIQWM 99
L++A++A EVI K R + K+D KAYD + W++L +M +M FS KWI+W+
Sbjct: 1 LHSALIANEVIDEAK---RSNKSCLVFKVDYEKAYDSVSWNFLMYMMWRMDFSPKWIKWI 171
Query: 100 VVSTESVDYNVLVNAEHVGPVILGRGLRQGGPLSLSFYHL 139
+S +VLVN + RGLRQG PL+ +++
Sbjct: 172 EECVKSASISVLVNGSPTAEFLPQRGLRQGDPLTPFLFNI 291
>TC233727 similar to UP|Q9FKH8 (Q9FKH8) Similarity to non-LTR retroelement
reverse transcriptase, partial (7%)
Length = 956
Score = 65.9 bits (159), Expect = 8e-11
Identities = 38/93 (40%), Positives = 54/93 (57%)
Frame = -3
Query: 45 VAIEVIHFMKTKTRGQDKYVALKLDISKAYDRMDWDYLKEIMIKMGFSQKWIQWMVVSTE 104
V EVI+ + K G + +ALK+DI KA+D +D D+L ++ K G+S + W+ V
Sbjct: 816 VTSEVINMLDKKVFGGN--LALKIDIVKAFDTLDRDFLISVLNKFGYSHTFCNWIRVILL 643
Query: 105 SVDYNVLVNAEHVGPVILGRGLRQGGPLSLSFY 137
S ++ VN E VG RG+RQG PLS S Y
Sbjct: 642 SAKLSISVNGEPVGFFSCQRGVRQGDPLSPSLY 544
>TC219712 similar to UP|G6PI_SPIOL (O82059) Glucose-6-phosphate isomerase,
cytosolic (GPI) (Phosphoglucose isomerase) (PGI)
(Phosphohexose isomerase) (PHI) , partial (56%)
Length = 1208
Score = 30.0 bits (66), Expect = 4.7
Identities = 16/40 (40%), Positives = 22/40 (55%), Gaps = 6/40 (15%)
Frame = +2
Query: 683 VWHSSLEC------GWFMV*CSTCCYSQ*YSYCCYFLSS* 716
VWH S C GW+ +*C CC+S + +FLS+*
Sbjct: 29 VWH*SQ*CFCILGLGWWPI*CLQCCWSIAFIPTIWFLSN* 148
Database: GMGI
Posted date: Oct 22, 2004 4:58 PM
Number of letters in database: 37,918,896
Number of sequences in database: 63,676
Lambda K H
0.368 0.164 0.641
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 45,081,859
Number of Sequences: 63676
Number of extensions: 708976
Number of successful extensions: 8419
Number of sequences better than 10.0: 6
Number of HSP's better than 10.0 without gapping: 3296
Number of HSP's successfully gapped in prelim test: 330
Number of HSP's that attempted gapping in prelim test: 5011
Number of HSP's gapped (non-prelim): 3876
length of query: 890
length of database: 12,639,632
effective HSP length: 106
effective length of query: 784
effective length of database: 5,889,976
effective search space: 4617741184
effective search space used: 4617741184
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 14 ( 7.4 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 36 (21.7 bits)
S2: 63 (28.9 bits)
Medicago: description of AC149205.14


