

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC149204.7 + phase: 0
(600 letters)
Database: GMGI
63,676 sequences; 37,918,896 total letters
Searching..................................................done
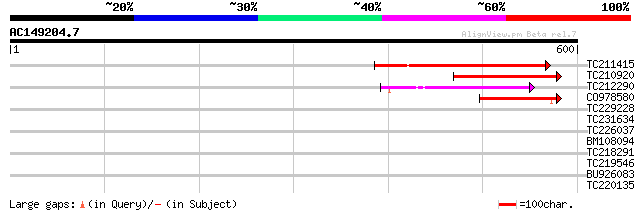
Score E
Sequences producing significant alignments: (bits) Value
TC211415 207 8e-54
TC210920 weakly similar to UP|Q75GS6 (Q75GS6) Expressed protein,... 158 6e-39
TC212290 weakly similar to UP|Q9FZE2 (Q9FZE2) T1K7.5 protein (At... 117 1e-26
CO978580 112 5e-25
TC229228 weakly similar to UP|Q9LN90 (Q9LN90) T12C24.7, partial ... 33 0.36
TC231634 similar to UP|Q8RY18 (Q8RY18) AT5g43560/K9D7_6, partial... 32 1.1
TC226037 homologue to UP|Q9AVG7 (Q9AVG7) Isopentenyl diphosphate... 30 2.3
BM108094 30 3.1
TC218291 similar to GB|AAM91387.1|22137084|AY133557 At3g13350/MD... 30 3.1
TC219546 UP|HS70_SOYBN (P26413) Heat shock 70 kDa protein, complete 30 4.0
BU926083 29 6.8
TC220135 similar to UP|Q9FH92 (Q9FH92) Emb|CAB62301.1 (At5g67210... 28 8.9
>TC211415
Length = 1065
Score = 207 bits (528), Expect = 8e-54
Identities = 99/186 (53%), Positives = 129/186 (69%)
Frame = +1
Query: 387 IPLGPNHQAEVPEWTGTTHKSDSKWLGTQIWPLQIVKSKLLEGEPVGKGRQDSCRCQVQG 446
+ +GP QAEVPEWTG +SDSKWLGT +W L+ ++ VG+GRQ+ C C+ G
Sbjct: 73 VSVGPRFQAEVPEWTGVFSESDSKWLGTHVWSLKN-DTEPATATDVGRGRQEMCSCEFHG 249
Query: 447 SVECVRFHIAEKSAKLKLEIGVAFYQWNLDKAGEEVRRCWTAEEEKKFKDVVKSNPASLD 506
SVECVR HIAE KLKLE+G FY+ D+ GEEV WT EEE++FKD++KSN +S +
Sbjct: 250 SVECVRLHIAENRMKLKLELGSEFYRLGFDRIGEEVSLQWTTEEEQRFKDIMKSNISSKN 429
Query: 507 RCFWDDIFTTFPKKSRESLVSYYFNVFLLQRRGYQNRHTPDNIDSDDEESEFTPLKGSFG 566
+ FW++ FPKK+R +LV+YYFNVFL+Q R YQNR TP+++DSDD+E EF FG
Sbjct: 430 KYFWNNPSKYFPKKTRRNLVNYYFNVFLIQLRTYQNRVTPESVDSDDDEVEFGSFSDGFG 609
Query: 567 HQTDKS 572
KS
Sbjct: 610 MDAVKS 627
>TC210920 weakly similar to UP|Q75GS6 (Q75GS6) Expressed protein, partial
(8%)
Length = 900
Score = 158 bits (400), Expect = 6e-39
Identities = 72/115 (62%), Positives = 85/115 (73%)
Frame = +1
Query: 470 FYQWNLDKAGEEVRRCWTAEEEKKFKDVVKSNPASLDRCFWDDIFTTFPKKSRESLVSYY 529
FY WN D+ GE+V+ W EEEKKFKDV++SNP S + FWD IF FP KSR LVSYY
Sbjct: 13 FYDWNFDQVGEDVKLLWMEEEEKKFKDVIRSNPPSPETYFWDHIFRAFPTKSRADLVSYY 192
Query: 530 FNVFLLQRRGYQNRHTPDNIDSDDEESEFTPLKGSFGHQTDKSNSFTLLSPKKPQ 584
FNVF+L+RRGYQNRHTP+NI+SDDE+ E PL+ FG QT S LL+PKK Q
Sbjct: 193 FNVFILERRGYQNRHTPNNINSDDEDDEAGPLRNVFGQQTQNSRGSILLTPKKSQ 357
>TC212290 weakly similar to UP|Q9FZE2 (Q9FZE2) T1K7.5 protein (At1g26580),
partial (28%)
Length = 791
Score = 117 bits (293), Expect = 1e-26
Identities = 69/185 (37%), Positives = 102/185 (54%), Gaps = 22/185 (11%)
Frame = +3
Query: 393 HQAEVPEW---------------------TGTTHKSDSKWLGTQIWPLQIVKSKLLEGEP 431
HQA+VP W G +++ + +GT + P+ ++ + E
Sbjct: 6 HQADVPAWDILGATNRPNASDAVSVSDFTVGHIDETEKRLMGTCVIPMPQMELSSNDDE- 182
Query: 432 VGKGRQDSCRCQVQGSVECVRFHIAEKSAKLKLEIGVA-FYQWNLDKAGEEVRRCWTAEE 490
VGK D C C+ QGS+ CVR HIAE+ K GV F + GE+V W+AE+
Sbjct: 183 VGKASTD-CSCEDQGSMRCVRQHIAEEREKHIKTFGVEKFTELGFTNMGEQVAENWSAED 359
Query: 491 EKKFKDVVKSNPASLDRCFWDDIFTTFPKKSRESLVSYYFNVFLLQRRGYQNRHTPDNID 550
E+ F +VV +NP SLD+ FW+ + FP +++ +VSYYFNVF+L+RR QNR+ +ID
Sbjct: 360 EQLFHEVVFNNPVSLDKNFWNYLSIVFPSLTKKEIVSYYFNVFMLRRRAEQNRNDLLSID 539
Query: 551 SDDEE 555
SD +E
Sbjct: 540 SDSDE 554
>CO978580
Length = 670
Score = 112 bits (280), Expect = 5e-25
Identities = 54/94 (57%), Positives = 65/94 (68%), Gaps = 7/94 (7%)
Frame = -1
Query: 498 VKSNPASLDRCFWDDIFTTFPKKSRESLVSYYFNVFLLQRRGYQNRHTPDNIDSDDEESE 557
++SNP S + FWD IF FP KSR LVSYYFNVF+L+RRGYQNRHTPD+I+SDD++ E
Sbjct: 670 IRSNPPSXETYFWDHIFRAFPTKSRADLVSYYFNVFILERRGYQNRHTPDDINSDDDDDE 491
Query: 558 FTPLKGSFGHQTDKS-------NSFTLLSPKKPQ 584
PL+ FGHQT S LL+PKK Q
Sbjct: 490 AGPLRNVFGHQTPNSGYQAPNPRGSILLTPKKSQ 389
>TC229228 weakly similar to UP|Q9LN90 (Q9LN90) T12C24.7, partial (10%)
Length = 885
Score = 33.1 bits (74), Expect = 0.36
Identities = 20/53 (37%), Positives = 32/53 (59%)
Frame = +2
Query: 31 LWDLVGEEYGLGVKVGSSVKLVYSKYLSGLETPLKNVVDGEFPKRDLVGDRVK 83
+WDL+ + GLG ++ +V+ V+ + LSG E PL + +GE P+ D D K
Sbjct: 407 VWDLILDNNGLGKEISETVERVFCR-LSGQEPPLFPLPNGE-PQPDKETDNRK 559
>TC231634 similar to UP|Q8RY18 (Q8RY18) AT5g43560/K9D7_6, partial (11%)
Length = 936
Score = 31.6 bits (70), Expect = 1.1
Identities = 46/193 (23%), Positives = 75/193 (38%), Gaps = 16/193 (8%)
Frame = +3
Query: 135 SELEKVYEYIDGRKSCGTNKMKDTNLVSNMAKKVESEGLVDVLMQDCKTNETSLRKLVNQ 194
SE EK + ++ K KD A V + QD +E + K+
Sbjct: 84 SEREKKSKKKQAKQKRNNRKGKDKEREERTAASVPDKN------QDNAVDEKNDSKMEEA 245
Query: 195 NAAME---IMDDVSDVANSMPGLSDGSKSCAND-DANDSGDDVLMLDPSSVNRESFGRKR 250
A E M+DVSD+++S+ G+++ + + D DA+ D D S VN + R
Sbjct: 246 QAVSEKPDAMEDVSDMSDSVDGVAETLQLDSEDRDASPVNWDT---DASEVNPPTKARNN 416
Query: 251 KRDD-------LMSEMQSWVIRTAKNPCDPVLGSMP-----EKSKWKSYGNQEIWKKVLL 298
DD + + S VI + + C S+P + K S+ N ++ K
Sbjct: 417 GIDDVSTMQNGISEKRSSSVIDDSSSTCS--TDSLPSVVMNDPHKGNSFSNYKVQKSPSR 590
Query: 299 FREAAFLKKDFGS 311
+ D GS
Sbjct: 591 GKNRGKTSSDVGS 629
>TC226037 homologue to UP|Q9AVG7 (Q9AVG7) Isopentenyl diphosphate isomerase
2, partial (98%)
Length = 1229
Score = 30.4 bits (67), Expect = 2.3
Identities = 21/66 (31%), Positives = 30/66 (44%), Gaps = 5/66 (7%)
Frame = +3
Query: 361 PDSEAKEKLLDSCVPESILDAPPTVNIPLGPNHQAEVPEWTGTTHKSDSK-----WLGTQ 415
P +E+LL P S L PP +PL P +P WT ++ S SK W+
Sbjct: 177 PSPSLREQLLSLSPPPSPL-TPPWARLPLPP----PMPAWTPSSAVSCSKTNAFWWMRMT 341
Query: 416 IWPLQI 421
+W + I
Sbjct: 342 VWLVMI 359
>BM108094
Length = 635
Score = 30.0 bits (66), Expect = 3.1
Identities = 15/31 (48%), Positives = 19/31 (60%)
Frame = -3
Query: 77 LVGDRVKFGERLTELQAELVLDDYGEGDVGD 107
L +R+ + L EL AELV D G+GD GD
Sbjct: 243 LSSNRMAYPNFLPELNAELVRDSSGDGDGGD 151
>TC218291 similar to GB|AAM91387.1|22137084|AY133557 At3g13350/MDC11_14
{Arabidopsis thaliana;} , partial (22%)
Length = 697
Score = 30.0 bits (66), Expect = 3.1
Identities = 17/54 (31%), Positives = 28/54 (51%)
Frame = +3
Query: 7 TLDLYKLFMVVKDKGGYDVVCKNELWDLVGEEYGLGVKVGSSVKLVYSKYLSGL 60
+LDL++LF+ V +GG + V + W V + + S+ +V YLS L
Sbjct: 324 SLDLHRLFVEVTSRGGIEKVIVDRKWKEVILTFNFKDTITSASXVVRKSYLSML 485
>TC219546 UP|HS70_SOYBN (P26413) Heat shock 70 kDa protein, complete
Length = 2099
Score = 29.6 bits (65), Expect = 4.0
Identities = 16/58 (27%), Positives = 29/58 (49%), Gaps = 1/58 (1%)
Frame = +1
Query: 247 GRKRKRDDLMSEMQSWVIRTAKNPCDPVLGSMPEKSKWKSYGNQEIWKKVLL-FREAA 303
GR+ + ++M+ W + +PCD + + K + K + +EI VL+ RE A
Sbjct: 229 GRRFSDSSVQNDMKLWPFKVGGSPCDKPMIVVNYKGEEKKFSAEEISSMVLVKMREVA 402
>BU926083
Length = 450
Score = 28.9 bits (63), Expect = 6.8
Identities = 20/51 (39%), Positives = 30/51 (58%), Gaps = 3/51 (5%)
Frame = -3
Query: 10 LYKLFMVVKDKGGYDVVCKNELWD---LVGEEYGLGVKVGSSVKLVYSKYL 57
L+ + VV GG +VC+N LWD G E L +K+G+ +L++SK L
Sbjct: 196 LHPVLHVVPSFGGSTLVCENLLWDRDLKHGFEPILTIKIGT--QLLHSKGL 50
>TC220135 similar to UP|Q9FH92 (Q9FH92) Emb|CAB62301.1 (At5g67210), partial
(44%)
Length = 670
Score = 28.5 bits (62), Expect = 8.9
Identities = 31/102 (30%), Positives = 45/102 (43%), Gaps = 1/102 (0%)
Frame = -3
Query: 500 SNPASLDRCFWDDIFTTFPKKSRESLVSYYFNVFLLQRRGYQNRHTPD-NIDSDDEESEF 558
SN SL R F+T P+KS + F FLL R TP NI +
Sbjct: 353 SNKFSLHRNSLPQTFSTSPEKSCKKT*VLGFPPFLLLAR------TPTVNIADILPGASG 192
Query: 559 TPLKGSFGHQTDKSNSFTLLSPKKPQPTPHTKGRQLVNGLKS 600
+G + T +S S+T L + +P+ H++ R+ GL S
Sbjct: 191 QSPRGPSTNMTSQSTSYTWLG-RSLRPSLHSEKRRFCTGLHS 69
Database: GMGI
Posted date: Oct 22, 2004 4:58 PM
Number of letters in database: 37,918,896
Number of sequences in database: 63,676
Lambda K H
0.314 0.133 0.395
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 26,959,568
Number of Sequences: 63676
Number of extensions: 382665
Number of successful extensions: 1440
Number of sequences better than 10.0: 26
Number of HSP's better than 10.0 without gapping: 1430
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 1436
length of query: 600
length of database: 12,639,632
effective HSP length: 103
effective length of query: 497
effective length of database: 6,081,004
effective search space: 3022258988
effective search space used: 3022258988
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 16 ( 7.3 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 42 (21.9 bits)
S2: 62 (28.5 bits)
Medicago: description of AC149204.7


