

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC149197.6 + phase: 0 /pseudo
(174 letters)
Database: GMGI
63,676 sequences; 37,918,896 total letters
Searching..................................................done
Score E
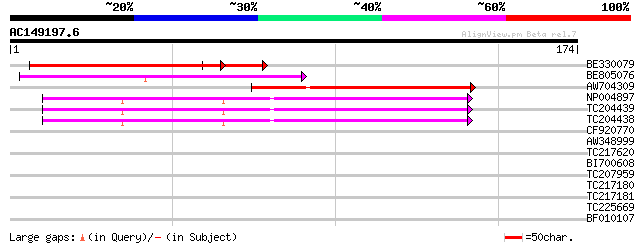
Sequences producing significant alignments: (bits) Value
BE330079 57 4e-10
BE805076 weakly similar to GP|9294121|dbj copia-like retrotransp... 54 3e-08
AW704309 weakly similar to GP|6642775|gb| gag-pol polyprotein {V... 50 7e-07
NP004897 gag-protease polyprotein 49 1e-06
TC204439 UP|Q84VI4 (Q84VI4) Gag-pol polyprotein, complete 47 3e-06
TC204438 homologue to UP|Q84VH6 (Q84VH6) Gag-pol polyprotein, co... 46 8e-06
CF920770 36 0.008
AW348999 similar to GP|9294602|dbj hydrolase-like protein {Arabi... 28 2.2
TC217620 similar to GB|AAO39914.1|28372864|BT003686 At1g71430 {A... 28 2.9
BI700608 homologue to GP|19578319|em homeodomain-leucine zipper ... 27 4.9
TC207959 UP|C972_SOYBN (O48921) Cytochrome P450 97B2 , complete 27 4.9
TC217180 similar to UP|Q9LQA1 (Q9LQA1) F4N2.17, partial (18%) 27 4.9
TC217181 similar to UP|Q9LQA1 (Q9LQA1) F4N2.17, partial (13%) 27 4.9
TC225669 UP|Q42792 (Q42792) Asparagine synthetase , complete 27 6.4
BF010107 26 8.3
>BE330079
Length = 447
Score = 57.4 bits (137), Expect(2) = 4e-10
Identities = 23/60 (38%), Positives = 43/60 (71%)
Frame = -1
Query: 7 PTHLPVFEGKNYDQWIVKMKVICRFQDVLEVVNDSVPALASNATDMQ*TANRDAKKKDGK 66
P +LP+ +GKN+DQW+V+M+++ RF DVLE+V + + +T+ Q +++A +++GK
Sbjct: 219 PRNLPIQDGKNWDQWVVQMRILFRF*DVLEIVETEIKEPGACSTESQRAEHKEAIRREGK 40
Score = 23.1 bits (48), Expect(2) = 4e-10
Identities = 9/20 (45%), Positives = 13/20 (65%)
Frame = -3
Query: 60 AKKKDGKAMFFMHQSVSNEI 79
+ +K GKA+F +HQ V I
Sbjct: 61 SNQKGGKALFLIHQCVDPNI 2
>BE805076 weakly similar to GP|9294121|dbj copia-like retrotransposable
element {Arabidopsis thaliana}, partial (2%)
Length = 388
Score = 54.3 bits (129), Expect = 3e-08
Identities = 29/90 (32%), Positives = 52/90 (57%), Gaps = 2/90 (2%)
Frame = +1
Query: 4 ESFPTHLPVFEGKNYDQWIVKMKVICRFQDVLEVVND--SVPALASNATDMQ*TANRDAK 61
+S +P+F G+NYD W VKM+ QD+ ++V + ++PA S Q + K
Sbjct: 112 QSSAISVPIFNGENYDFWRVKMETYLSSQDLWDIVEEGFTIPADTSALNASQEKELKKNK 291
Query: 62 KKDGKAMFFMHQSVSNEIFEKIVHYESAKE 91
+K+ K +F + Q+ ++ IF +I+ ++AKE
Sbjct: 292 QKNSKTLFTLQQAETDPIFPRIMGAKTAKE 381
>AW704309 weakly similar to GP|6642775|gb| gag-pol polyprotein {Vitis
vinifera}, partial (23%)
Length = 423
Score = 49.7 bits (117), Expect = 7e-07
Identities = 24/69 (34%), Positives = 46/69 (65%)
Frame = -1
Query: 75 VSNEIFEKIVHYESAKEETWEALEKLYSGDGKLKKVRLQVLRRQYEVLTMEEEETISQYF 134
VS IF +I+ +S K W+ L++ Y+GD +++ +++ LRR++E+ M+E ETI +Y
Sbjct: 399 VSQMIFIRIMTLKSPKA-IWDCLKEEYAGDDRIRSMQVLNLRREFEL*RMKESETIKEYS 223
Query: 135 DKFVNLSNQ 143
+K + + N+
Sbjct: 222 NKLLGIVNK 196
>NP004897 gag-protease polyprotein
Length = 1923
Score = 48.9 bits (115), Expect = 1e-06
Identities = 34/145 (23%), Positives = 65/145 (44%), Gaps = 13/145 (8%)
Frame = +1
Query: 11 PVFEGKNYDQWIVKMKVICRFQD-------VLEVVNDSVPALASNATDMQ*TANRDAKKK 63
P+ +G NY+ W +M + D + + + + TD K++
Sbjct: 40 PILDGTNYEYWKARMVAFLKSLDSRTWKAVIKDWEHPKMLDTEGKPTDGLKPEEDWTKEE 219
Query: 64 D------GKAMFFMHQSVSNEIFEKIVHYESAKEETWEALEKLYSGDGKLKKVRLQVLRR 117
D KA+ + V IF ++++ + ++ WE L+ + G K+K RLQ+L
Sbjct: 220 DELALGNSKALNALFNGVDKNIF-RLINTCTVAKDAWEILKTTHEGTSKVKMSRLQLLAT 396
Query: 118 QYEVLTMEEEETISQYFDKFVNLSN 142
++E L M+EEE I + + ++N
Sbjct: 397 KFENLKMKEEECIHDFHMNILEIAN 471
>TC204439 UP|Q84VI4 (Q84VI4) Gag-pol polyprotein, complete
Length = 4731
Score = 47.4 bits (111), Expect = 3e-06
Identities = 34/145 (23%), Positives = 64/145 (43%), Gaps = 13/145 (8%)
Frame = +1
Query: 11 PVFEGKNYDQWIVKMKVICRFQD-------VLEVVNDSVPALASNATDMQ*TANRDAKKK 63
P+ +G NY+ W +M + D + + + TD K++
Sbjct: 40 PILDGSNYEYWKARMVAFLKSLDSRTWKAVIKGWEHPKMLDTEGKPTDELKPEEDWTKEE 219
Query: 64 D------GKAMFFMHQSVSNEIFEKIVHYESAKEETWEALEKLYSGDGKLKKVRLQVLRR 117
D KA+ + V IF ++++ + ++ WE L+ + G K+K RLQ+L
Sbjct: 220 DELALGNSKALNALFNGVDKNIF-RLINTCTVAKDAWEILKITHEGTSKVKMSRLQLLAT 396
Query: 118 QYEVLTMEEEETISQYFDKFVNLSN 142
++E L M+EEE I + + ++N
Sbjct: 397 KFENLKMKEEECIHDFHMNILEIAN 471
>TC204438 homologue to UP|Q84VH6 (Q84VH6) Gag-pol polyprotein, complete
Length = 4734
Score = 46.2 bits (108), Expect = 8e-06
Identities = 33/145 (22%), Positives = 63/145 (42%), Gaps = 13/145 (8%)
Frame = +1
Query: 11 PVFEGKNYDQWIVKMKVICRFQD------VLEVVNDSVPALASNATDMQ*TANRDAKKKD 64
P+ +G NY+ W +M + D V++ + D K++
Sbjct: 40 PILDGTNYEYWKARMVAFLKSLDSRTWKAVIKGWEHPKMLDTEGKPTNELKPEEDWTKEE 219
Query: 65 -------GKAMFFMHQSVSNEIFEKIVHYESAKEETWEALEKLYSGDGKLKKVRLQVLRR 117
KA+ + V IF ++++ + ++ WE L+ + G K+K RLQ+L
Sbjct: 220 DELALGNSKALNALFNGVDKNIF-RLINTCTVAKDAWEILKTTHEGTSKVKMSRLQLLAT 396
Query: 118 QYEVLTMEEEETISQYFDKFVNLSN 142
++E L M+EEE I + + ++N
Sbjct: 397 KFENLKMKEEECIHDFHMNILEIAN 471
>CF920770
Length = 581
Score = 36.2 bits (82), Expect = 0.008
Identities = 19/77 (24%), Positives = 38/77 (48%)
Frame = -2
Query: 66 KAMFFMHQSVSNEIFEKIVHYESAKEETWEALEKLYSGDGKLKKVRLQVLRRQYEVLTME 125
KA + ++ + + ++ + +SAKE W+ L + G +K+ R+ L +YE+ M
Sbjct: 415 KAKNIITSALGMDEYFRVSNCKSAKE-MWDTLRLTHEGTTDVKRSRINALTHEYELFRMN 239
Query: 126 EEETISQYFDKFVNLSN 142
E I +F ++ N
Sbjct: 238 TNENIQSMQKRFTHIVN 188
>AW348999 similar to GP|9294602|dbj hydrolase-like protein {Arabidopsis
thaliana}, partial (46%)
Length = 642
Score = 28.1 bits (61), Expect = 2.2
Identities = 13/38 (34%), Positives = 20/38 (52%)
Frame = -3
Query: 8 THLPVFEGKNYDQWIVKMKVICRFQDVLEVVNDSVPAL 45
T LPV+ G+NY + IC DV ++ ++ P L
Sbjct: 439 TPLPVWHGQNYRSLGFRFGNICYISDVSDIPEETYPLL 326
>TC217620 similar to GB|AAO39914.1|28372864|BT003686 At1g71430 {Arabidopsis
thaliana;} , partial (40%)
Length = 1060
Score = 27.7 bits (60), Expect = 2.9
Identities = 15/56 (26%), Positives = 28/56 (49%)
Frame = -2
Query: 10 LPVFEGKNYDQWIVKMKVICRFQDVLEVVNDSVPALASNATDMQ*TANRDAKKKDG 65
L +F ++WI++ K +CR+ E + S+P L Q + N + ++DG
Sbjct: 303 LILFYYHKQEEWILRKKGVCRYAQT*EALCSSLPYL---VVSCQLSQNLETFQRDG 145
>BI700608 homologue to GP|19578319|em homeodomain-leucine zipper protein
{Arabidopsis thaliana}, partial (13%)
Length = 421
Score = 26.9 bits (58), Expect = 4.9
Identities = 16/43 (37%), Positives = 23/43 (53%)
Frame = +3
Query: 92 ETWEALEKLYSGDGKLKKVRLQVLRRQYEVLTMEEEETISQYF 134
E EALE+LY K +R Q L R+ +L+ E + I +F
Sbjct: 117 EQVEALERLYHDCPKPSSIRRQQLIRECPILSNIEPKQIKVWF 245
>TC207959 UP|C972_SOYBN (O48921) Cytochrome P450 97B2 , complete
Length = 1863
Score = 26.9 bits (58), Expect = 4.9
Identities = 29/113 (25%), Positives = 51/113 (44%), Gaps = 18/113 (15%)
Frame = +1
Query: 20 QWIVKMKVICRFQDVLEVVNDSVPALASNA------TDMQ*TANRD-AKKKDGKAMFFM- 71
+WIV + +FQD L+V+N + L NA TD++ RD KD + F+
Sbjct: 871 RWIVPRQR--KFQDDLKVINTCLDGLIRNAKESRQETDVEKLQQRDYLNLKDASLLRFLV 1044
Query: 72 --------HQSVSNEIFEKIV--HYESAKEETWEALEKLYSGDGKLKKVRLQV 114
+ + +++ ++ H +A TW A+ L K+KK + +V
Sbjct: 1045DMRGADVDDRQLRDDLMTMLIAGHETTAAVLTW-AVFLLAQNPSKMKKAQAEV 1200
>TC217180 similar to UP|Q9LQA1 (Q9LQA1) F4N2.17, partial (18%)
Length = 1298
Score = 26.9 bits (58), Expect = 4.9
Identities = 12/19 (63%), Positives = 16/19 (84%)
Frame = +1
Query: 93 TWEALEKLYSGDGKLKKVR 111
T EAL++L+SGDG+ KK R
Sbjct: 841 TVEALQELFSGDGQSKKGR 897
>TC217181 similar to UP|Q9LQA1 (Q9LQA1) F4N2.17, partial (13%)
Length = 820
Score = 26.9 bits (58), Expect = 4.9
Identities = 12/19 (63%), Positives = 16/19 (84%)
Frame = +2
Query: 93 TWEALEKLYSGDGKLKKVR 111
T EAL++L+SGDG+ KK R
Sbjct: 233 TVEALQELFSGDGQSKKGR 289
>TC225669 UP|Q42792 (Q42792) Asparagine synthetase , complete
Length = 2231
Score = 26.6 bits (57), Expect = 6.4
Identities = 11/41 (26%), Positives = 22/41 (52%)
Frame = -3
Query: 69 FFMHQSVSNEIFEKIVHYESAKEETWEALEKLYSGDGKLKK 109
FF+H +++ VHY+SA+ E +K++ +L +
Sbjct: 2214 FFLHSNLTEHFLRNPVHYDSAQMT*IENEDKMWMAQRRLDR 2092
>BF010107
Length = 380
Score = 26.2 bits (56), Expect = 8.3
Identities = 16/62 (25%), Positives = 33/62 (52%)
Frame = -2
Query: 72 HQSVSNEIFEKIVHYESAKEETWEALEKLYSGDGKLKKVRLQVLRRQYEVLTMEEEETIS 131
HQ++ + E +VH S +++ L LYSG L+ ++ +L R+ ++E+E +
Sbjct: 220 HQNIYY-VKEGLVHCMSLRQKL*VKLCLLYSGQQSLEHMKEDMLSRETGSRILDEDELVE 44
Query: 132 QY 133
+
Sbjct: 43 GF 38
Database: GMGI
Posted date: Oct 22, 2004 4:58 PM
Number of letters in database: 37,918,896
Number of sequences in database: 63,676
Lambda K H
0.332 0.140 0.436
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 6,434,703
Number of Sequences: 63676
Number of extensions: 61637
Number of successful extensions: 559
Number of sequences better than 10.0: 30
Number of HSP's better than 10.0 without gapping: 558
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 559
length of query: 174
length of database: 12,639,632
effective HSP length: 91
effective length of query: 83
effective length of database: 6,845,116
effective search space: 568144628
effective search space used: 568144628
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 15 ( 7.2 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 39 (21.5 bits)
S2: 55 (25.8 bits)
Medicago: description of AC149197.6


