

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC149135.13 - phase: 0
(316 letters)
Database: GMGI
63,676 sequences; 37,918,896 total letters
Searching..................................................done
Score E
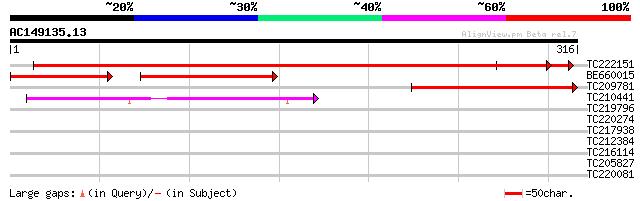
Sequences producing significant alignments: (bits) Value
TC222151 homologue to UP|Q8LAA2 (Q8LAA2) Cell division-like prot... 502 e-142
BE660015 138 2e-62
TC209781 similar to UP|Q8LAA2 (Q8LAA2) Cell division-like protei... 185 2e-47
TC210441 similar to UP|Q9SXH3 (Q9SXH3) FtsJ, partial (80%) 54 9e-08
TC219796 similar to UP|Q9SXH3 (Q9SXH3) FtsJ, partial (35%) 36 0.026
TC220274 30 1.9
TC217938 28 4.2
TC212384 similar to UP|O22512 (O22512) Grr1, partial (10%) 28 7.1
TC216114 weakly similar to UP|Q93VM8 (Q93VM8) AT5g20650/T1M15_50... 27 9.3
TC205827 27 9.3
TC220081 similar to UP|O49356 (O49356) Ferredoxin--NADP+ reducta... 27 9.3
>TC222151 homologue to UP|Q8LAA2 (Q8LAA2) Cell division-like protein, partial
(89%)
Length = 1053
Score = 502 bits (1292), Expect = e-142
Identities = 254/290 (87%), Positives = 262/290 (89%), Gaps = 1/290 (0%)
Frame = +3
Query: 14 RKAKEEGWRARSAFKLLQIDEEFNIFEGVKRVVDLCAAPGSWSQVLSRKLYLPAKLAPDA 73
RKAKEEGWRARSAFKLLQIDEEFN+F+GVKRVVDLCAAPGSWSQVLSRKLYLPAKLAPDA
Sbjct: 6 RKAKEEGWRARSAFKLLQIDEEFNLFDGVKRVVDLCAAPGSWSQVLSRKLYLPAKLAPDA 185
Query: 74 KDENLPLIVAIDLQPMAPIEGVIQVQGDITNARTAEVVIRHFDGCKANLVVCDGAPDVTG 133
KDENLPLIVAIDLQPMAPIEGVIQVQGDITNARTAEVVIRHFDGCKA+LVVCDGAPDVTG
Sbjct: 186 KDENLPLIVAIDLQPMAPIEGVIQVQGDITNARTAEVVIRHFDGCKADLVVCDGAPDVTG 365
Query: 134 LHDMDEFVQSQLILAGLTIVTHVLKEGGKFIAKIFRGKDTSLLYCQLKLFFPVVTFAKPK 193
LHDMDEFVQSQL+LAGLTIVTHVLKEGGKFIAKIFRGKDTSLLYCQLKLFFPVVTFAKPK
Sbjct: 366 LHDMDEFVQSQLLLAGLTIVTHVLKEGGKFIAKIFRGKDTSLLYCQLKLFFPVVTFAKPK 545
Query: 194 SSRNSSIEAFAVCENYSPPEGFNPKDLHRLLEKVGSPSGVDDTDCVSGWLEGPNKVYIPF 253
SSRNSSI AFAV ENYSPPEGF DLHRLLEKVGSPSGVDDTDC GWLEGPNKVYIPF
Sbjct: 546 SSRNSSIXAFAVXENYSPPEGFXXXDLHRLLEKVGSPSGVDDTDCCXGWLEGPNKVYIPF 725
Query: 254 LACGDLTGYDSDRSYPLPKVAGGTYQSLDPVQPPIAPPYKRA-LELKKAS 302
LACGDL+GYDSDRSYPLP G + + P+ P L KKAS
Sbjct: 726 LACGDLSGYDSDRSYPLP*SCRGKHIRAWILCSPLLPHLTXXLLXXKKAS 875
Score = 47.0 bits (110), Expect = 1e-05
Identities = 26/44 (59%), Positives = 29/44 (65%), Gaps = 1/44 (2%)
Frame = +2
Query: 272 KVAGGTYQSLDPVQPPIAPPYKRALELKKA-SPQGFRELENLSL 314
K+ G TYQSLDPVQPPIAPPY A K+ + EL NLSL
Sbjct: 782 KLPGETYQSLDPVQPPIAPPYXXAFGXXKSFXFKDSXELXNLSL 913
>BE660015
Length = 647
Score = 138 bits (347), Expect(2) = 2e-62
Identities = 67/76 (88%), Positives = 69/76 (90%)
Frame = +2
Query: 74 KDENLPLIVAIDLQPMAPIEGVIQVQGDITNARTAEVVIRHFDGCKANLVVCDGAPDVTG 133
+DENLPLIVAIDLQPMAPIEGVIQVQGDITNARTAEVVIRHFDGCKA+LVVCDGAPDVTG
Sbjct: 419 RDENLPLIVAIDLQPMAPIEGVIQVQGDITNARTAEVVIRHFDGCKADLVVCDGAPDVTG 598
Query: 134 LHDMDEFVQSQLILAG 149
LHDMDEFV G
Sbjct: 599 LHDMDEFVNPNSYFQG 646
Score = 119 bits (298), Expect(2) = 2e-62
Identities = 56/57 (98%), Positives = 57/57 (99%)
Frame = +1
Query: 1 MGKASRDKRDIYYRKAKEEGWRARSAFKLLQIDEEFNIFEGVKRVVDLCAAPGSWSQ 57
MGKASRDKRDIYYRKAKEEGWRARSAFKLLQIDEEFN+FEGVKRVVDLCAAPGSWSQ
Sbjct: 250 MGKASRDKRDIYYRKAKEEGWRARSAFKLLQIDEEFNLFEGVKRVVDLCAAPGSWSQ 420
>TC209781 similar to UP|Q8LAA2 (Q8LAA2) Cell division-like protein, partial
(26%)
Length = 476
Score = 185 bits (470), Expect = 2e-47
Identities = 88/92 (95%), Positives = 89/92 (96%)
Frame = +1
Query: 225 EKVGSPSGVDDTDCVSGWLEGPNKVYIPFLACGDLTGYDSDRSYPLPKVAGGTYQSLDPV 284
EKVGSPSGVDDTDC SGWLEGPNKVYIPFLACGDL+GYDSDRSY LPKVAGGTYQSLDPV
Sbjct: 7 EKVGSPSGVDDTDCCSGWLEGPNKVYIPFLACGDLSGYDSDRSYTLPKVAGGTYQSLDPV 186
Query: 285 QPPIAPPYKRALELKKASPQGFRELENLSLDS 316
QPPIAPPYKRALELKKAS QGFRELENLSLDS
Sbjct: 187 QPPIAPPYKRALELKKASSQGFRELENLSLDS 282
>TC210441 similar to UP|Q9SXH3 (Q9SXH3) FtsJ, partial (80%)
Length = 740
Score = 53.9 bits (128), Expect = 9e-08
Identities = 46/179 (25%), Positives = 74/179 (40%), Gaps = 16/179 (8%)
Frame = +3
Query: 10 DIYYRKAKEEGWRARSAFKLLQIDEEFNIFEGVKRVVDLCAAPGSWSQVLSRKLYL---- 65
D +Y++A+ G+ ARSAFKLLQI + I ++DL APG W QV + L
Sbjct: 12 DFFYKEAQRLGYVARSAFKLLQIQNQHKIISPGSSILDLGCAPGGWLQVACQSLGPFRGG 191
Query: 66 PAKLAPDAKDENLPLIVAIDLQPMAPIEGVIQVQGDITNARTAEVVIRHFDGCKANLVVC 125
+ L D K +P P+ V + D+T + ++++
Sbjct: 192 GSVLGVDTKKVKVP--------PLHCDSRVQTISADVTTLPHQRLKALSPKEKGFSVILS 347
Query: 126 DGAPDVTGLHDMDEFVQSQLILAGLTIV------------THVLKEGGKFIAKIFRGKD 172
D P V+G+ D + +L + L + VLK GG + K+ +D
Sbjct: 348 DMCPLVSGITTKDAALSFELGMRALDLALGSRIHLEPSDCVGVLKVGGHLVIKLLESED 524
>TC219796 similar to UP|Q9SXH3 (Q9SXH3) FtsJ, partial (35%)
Length = 763
Score = 35.8 bits (81), Expect = 0.026
Identities = 18/39 (46%), Positives = 22/39 (56%)
Frame = +2
Query: 20 GWRARSAFKLLQIDEEFNIFEGVKRVVDLCAAPGSWSQV 58
G+ A SAFKLLQI + I V+DL APG W +
Sbjct: 23 GYVAGSAFKLLQIQNQH*IMSSGSSVLDLGCAPGGWQSL 139
>TC220274
Length = 941
Score = 29.6 bits (65), Expect = 1.9
Identities = 16/46 (34%), Positives = 26/46 (55%)
Frame = -1
Query: 161 GKFIAKIFRGKDTSLLYCQLKLFFPVVTFAKPKSSRNSSIEAFAVC 206
GK+I+ + TSL++ QL FFPV + + N+S++ VC
Sbjct: 578 GKWISCSYVFNSTSLIHSQLCDFFPVTIWNIRRPLLNASLKRILVC 441
>TC217938
Length = 957
Score = 28.5 bits (62), Expect = 4.2
Identities = 16/41 (39%), Positives = 22/41 (53%), Gaps = 2/41 (4%)
Frame = +1
Query: 161 GKFIAKIFRGKDTSLLYCQLKLFF--PVVTFAKPKSSRNSS 199
G+F++ RG T LLY +FF PVV +R+SS
Sbjct: 385 GRFVSSSSRGPGTELLYTGRSIFFGSPVVNEQPSTGTRSSS 507
>TC212384 similar to UP|O22512 (O22512) Grr1, partial (10%)
Length = 709
Score = 27.7 bits (60), Expect = 7.1
Identities = 15/39 (38%), Positives = 22/39 (55%), Gaps = 2/39 (5%)
Frame = -1
Query: 186 VVTFAKPKSSRNSSIEAFAVCENYSPPE--GFNPKDLHR 222
++ +K SS +SIE F C N+SP + G +DL R
Sbjct: 475 LLKISKRHSSGKASIEVFCFCSNHSPSKTNGALTRDLFR 359
>TC216114 weakly similar to UP|Q93VM8 (Q93VM8) AT5g20650/T1M15_50 (COPT5),
partial (60%)
Length = 678
Score = 27.3 bits (59), Expect = 9.3
Identities = 16/42 (38%), Positives = 22/42 (52%)
Frame = -1
Query: 269 PLPKVAGGTYQSLDPVQPPIAPPYKRALELKKASPQGFRELE 310
P K +G + LD V + PP KR L+L++ G ELE
Sbjct: 312 PEEKRSGDLHAHLDAVSGEL-PPQKRGLDLRREGLSGADELE 190
>TC205827
Length = 2695
Score = 27.3 bits (59), Expect = 9.3
Identities = 20/58 (34%), Positives = 31/58 (52%), Gaps = 2/58 (3%)
Frame = +1
Query: 150 LTIVTHVLKEGGKFIA-KIFRGKDTSLLYCQLKLFFPVVTFAKP-KSSRNSSIEAFAV 205
L+ V VLK GGKF+ + +LL+ + +L + + A P KSS S++ F V
Sbjct: 481 LSEVKRVLKPGGKFVCLTLAESHVLNLLFSKFRLGWKMSVDAIPLKSSGKPSLQTFMV 654
>TC220081 similar to UP|O49356 (O49356) Ferredoxin--NADP+ reductase - like
protein, partial (24%)
Length = 548
Score = 27.3 bits (59), Expect = 9.3
Identities = 16/49 (32%), Positives = 23/49 (46%), Gaps = 1/49 (2%)
Frame = +2
Query: 213 EGFNPKDLHRLLEKVGSPSGVDDTDCVSGWLE-GPNKVYIPFLACGDLT 260
+G P D R+L PS ++ V GWL+ GP + L C + T
Sbjct: 68 KGVVPNDRGRVLSDTSDPSVLEKGLYVCGWLKRGPTGIIATNLYCAEET 214
Database: GMGI
Posted date: Oct 22, 2004 4:58 PM
Number of letters in database: 37,918,896
Number of sequences in database: 63,676
Lambda K H
0.319 0.139 0.416
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 13,891,288
Number of Sequences: 63676
Number of extensions: 203046
Number of successful extensions: 903
Number of sequences better than 10.0: 22
Number of HSP's better than 10.0 without gapping: 897
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 903
length of query: 316
length of database: 12,639,632
effective HSP length: 97
effective length of query: 219
effective length of database: 6,463,060
effective search space: 1415410140
effective search space used: 1415410140
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 16 ( 7.4 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.7 bits)
S2: 59 (27.3 bits)
Medicago: description of AC149135.13


