

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC149134.13 - phase: 0
(163 letters)
Database: GMGI
63,676 sequences; 37,918,896 total letters
Searching..................................................done
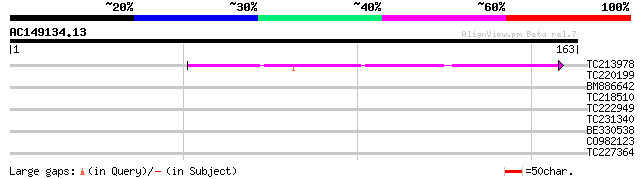
Score E
Sequences producing significant alignments: (bits) Value
TC213978 41 3e-04
TC220199 similar to UP|O65001 (O65001) Terminal ear1, partial (21%) 28 1.5
BM886642 28 2.5
TC218510 similar to UP|Q9FYK6 (Q9FYK6) F21J9.24, partial (49%) 27 3.3
TC222949 27 4.3
TC231340 similar to UP|ECR_AEDAE (P49880) Ecdysone receptor (Ecd... 26 7.4
BE330538 similar to SP|Q9SGG3|ALA Potential phospholipid-transpo... 26 9.6
CO982123 26 9.6
TC227364 similar to UP|Q9SYH5 (Q9SYH5) F15I1.27, partial (8%) 26 9.6
>TC213978
Length = 716
Score = 40.8 bits (94), Expect = 3e-04
Identities = 38/114 (33%), Positives = 55/114 (47%), Gaps = 6/114 (5%)
Frame = +1
Query: 52 QMTLPSFVFSSKTLSILKLKQITLNEVPF------VNLPSLKALYLDVVTFTYYELILKL 105
++ L +FS KTL +LKL LN PF V+LP L L+L ++ +L
Sbjct: 4 RVRLXXALFSCKTLVVLKLIG-GLNRNPFPLDFKSVDLPLLTTLHLQSFILERRDM-AEL 177
Query: 106 LSGCPILQYLGTNNLVVELPYSERPVISLSNLIRANICDIHIEFDWLQNVERLR 159
L G P L+YL ++ P E L L+RA I H+ + + NV+ LR
Sbjct: 178 LRGSPNLEYLFVGHMYFSGP--EARFERLPKLLRATIAFGHVPLEVVNNVQFLR 333
>TC220199 similar to UP|O65001 (O65001) Terminal ear1, partial (21%)
Length = 811
Score = 28.5 bits (62), Expect = 1.5
Identities = 15/39 (38%), Positives = 21/39 (53%)
Frame = +3
Query: 12 SFHLQCWNDYDCRDIYNFLYIAIQRGVENLNIDFSHSLF 50
+FHLQ W ++ R I Y +Q G+E L F +S F
Sbjct: 207 AFHLQHWEVFNSRKICEVTYARVQ-GLEALKEHFKNSKF 320
>BM886642
Length = 421
Score = 27.7 bits (60), Expect = 2.5
Identities = 15/36 (41%), Positives = 22/36 (60%), Gaps = 3/36 (8%)
Frame = +3
Query: 48 SLFSQMTLPSFVFSSKTLSILKLKQITL---NEVPF 80
S S++ PSF + + LKL+Q+TL NE+PF
Sbjct: 66 SQHSKLCKPSFRIENTRIQFLKLQQMTLC*MNELPF 173
>TC218510 similar to UP|Q9FYK6 (Q9FYK6) F21J9.24, partial (49%)
Length = 796
Score = 27.3 bits (59), Expect = 3.3
Identities = 15/31 (48%), Positives = 17/31 (54%), Gaps = 1/31 (3%)
Frame = -1
Query: 132 ISLSNLIRANICDI-HIEFDWLQNVERLRAT 161
I LSNL +N C H DWLQN R+T
Sbjct: 523 ICLSNLCGSNACKATH*PLDWLQNDTNKRST 431
>TC222949
Length = 608
Score = 26.9 bits (58), Expect = 4.3
Identities = 24/81 (29%), Positives = 38/81 (46%)
Frame = -1
Query: 27 YNFLYIAIQRGVENLNIDFSHSLFSQMTLPSFVFSSKTLSILKLKQITLNEVPFVNLPSL 86
Y + Y+ + R DFS L S PS F + I K +++ + FVN PS
Sbjct: 278 YQYQYVNLTRA------DFSPLLPS----PSS*FPFYVIGISKSRELQPRQ--FVNAPSY 135
Query: 87 KALYLDVVTFTYYELILKLLS 107
+LY+ + Y + L++LS
Sbjct: 134 LSLYIFAPFYDYLD*HLQMLS 72
>TC231340 similar to UP|ECR_AEDAE (P49880) Ecdysone receptor (Ecdysteroid
receptor) (20-hydroxy-ecdysone receptor) (20E receptor),
partial (4%)
Length = 971
Score = 26.2 bits (56), Expect = 7.4
Identities = 24/102 (23%), Positives = 42/102 (40%), Gaps = 7/102 (6%)
Frame = +2
Query: 2 AMRDITLPILSFHLQCWNDYDCRDIYNFLYIAIQRGVENLNIDFSHSLFSQMTLPSFVFS 61
++RD +L +L + + + + + A+ V L I + +P VFS
Sbjct: 2 SLRDASLALLKLDFERHGCIEPQLLKRIVKYAVSHNVRQLGISVKCDI---RDVPQCVFS 172
Query: 62 SKTLSILKLKQITLNEV-------PFVNLPSLKALYLDVVTF 96
+TL+ LKL + +NL +L L+L TF
Sbjct: 173 CQTLTSLKLSVCPRGYIYGSTLFPKSLNLTALTTLHLQHFTF 298
>BE330538 similar to SP|Q9SGG3|ALA Potential phospholipid-transporting ATPase
5 (EC 3.6.3.1) (Aminophospholipid flippase 5)., partial
(12%)
Length = 472
Score = 25.8 bits (55), Expect = 9.6
Identities = 14/42 (33%), Positives = 23/42 (54%)
Frame = -3
Query: 111 ILQYLGTNNLVVELPYSERPVISLSNLIRANICDIHIEFDWL 152
+L+Y G + +PY ER + ++ L N C IH++ WL
Sbjct: 398 LLEYSGES-----IPYKERKMYQVAMLPHTNRC*IHVK--WL 294
>CO982123
Length = 790
Score = 25.8 bits (55), Expect = 9.6
Identities = 15/45 (33%), Positives = 24/45 (53%)
Frame = +2
Query: 36 RGVENLNIDFSHSLFSQMTLPSFVFSSKTLSILKLKQITLNEVPF 80
R +L I+ SLF+ + L F++ +SI L +TL +PF
Sbjct: 491 RWAVSLAINL*TSLFTSIQLQPGSFTTLLMSIFSLTILTLAHIPF 625
>TC227364 similar to UP|Q9SYH5 (Q9SYH5) F15I1.27, partial (8%)
Length = 1376
Score = 25.8 bits (55), Expect = 9.6
Identities = 11/33 (33%), Positives = 19/33 (57%)
Frame = +3
Query: 18 WNDYDCRDIYNFLYIAIQRGVENLNIDFSHSLF 50
W D+D + YNF + G + +N+ F+H+ F
Sbjct: 1065 WEDFDKKKNYNF*ASWLIIGYKLINLIFTHTSF 1163
Database: GMGI
Posted date: Oct 22, 2004 4:58 PM
Number of letters in database: 37,918,896
Number of sequences in database: 63,676
Lambda K H
0.327 0.142 0.427
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 7,685,214
Number of Sequences: 63676
Number of extensions: 103502
Number of successful extensions: 799
Number of sequences better than 10.0: 18
Number of HSP's better than 10.0 without gapping: 797
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 799
length of query: 163
length of database: 12,639,632
effective HSP length: 90
effective length of query: 73
effective length of database: 6,908,792
effective search space: 504341816
effective search space used: 504341816
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 15 ( 7.1 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 40 (21.7 bits)
S2: 55 (25.8 bits)
Medicago: description of AC149134.13


