

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
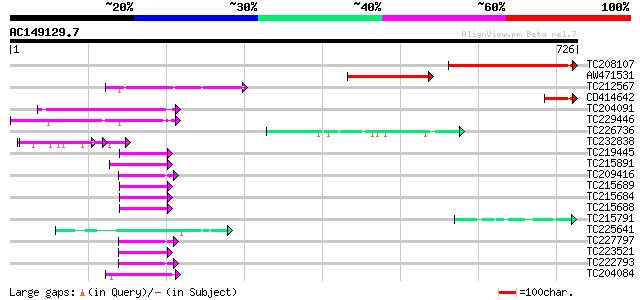
Query= AC149129.7 + phase: 0
(726 letters)
Database: GMGI
63,676 sequences; 37,918,896 total letters
Searching..................................................done
Score E
Sequences producing significant alignments: (bits) Value
TC208107 similar to UP|Q8RWQ1 (Q8RWQ1) At2g44720/F16B22.21, part... 249 3e-66
AW471531 weakly similar to GP|20147226|gb| At2g44720/F16B22.21 {... 145 6e-35
TC212567 weakly similar to UP|PAB3_HUMAN (Q9H361) Polyadenylate-... 81 1e-15
CD414642 similar to GP|22655529|gb| water-stress protein {Zea ma... 68 2e-11
TC204091 similar to UP|Q8LLE6 (Q8LLE6) RNA-binding protein AKIP1... 65 8e-11
TC229446 similar to UP|Q8LLE6 (Q8LLE6) RNA-binding protein AKIP1... 65 1e-10
TC226736 similar to UP|Q9SEE9 (Q9SEE9) Arginine/serine-rich prot... 62 1e-09
TC232838 similar to GB|AAP21377.1|30102918|BT006569 At1g47970 {A... 57 3e-08
TC219445 similar to UP|P93396 (P93396) Transformer-SR ribonucleo... 54 3e-07
TC215891 similar to GB|AAM78058.1|21928059|AY125548 AT5g61030/ma... 53 4e-07
TC209416 similar to UP|Q8LF90 (Q8LF90) RNA recognition motif-con... 53 4e-07
TC215689 similar to UP|P93396 (P93396) Transformer-SR ribonucleo... 52 9e-07
TC215684 similar to UP|P93396 (P93396) Transformer-SR ribonucleo... 52 9e-07
TC215688 similar to UP|P93396 (P93396) Transformer-SR ribonucleo... 52 9e-07
TC215791 similar to GB|CAE76446.1|38567152|BX842633 related to p... 52 1e-06
TC225641 similar to GB|AAA18379.1|475719|ATU08467 RNA-binding pr... 51 2e-06
TC227797 similar to UP|Q8H1N8 (Q8H1N8) At1g20880/F9H16_14, parti... 50 3e-06
TC223521 weakly similar to UP|Q84ZB9 (Q84ZB9) RNA recognition mo... 50 3e-06
TC222793 similar to UP|Q8LF90 (Q8LF90) RNA recognition motif-con... 50 5e-06
TC204084 UP|Q9SWA8 (Q9SWA8) Glycine-rich RNA-binding protein, co... 50 5e-06
>TC208107 similar to UP|Q8RWQ1 (Q8RWQ1) At2g44720/F16B22.21, partial (15%)
Length = 1023
Score = 249 bits (636), Expect = 3e-66
Identities = 128/169 (75%), Positives = 141/169 (82%), Gaps = 4/169 (2%)
Frame = +2
Query: 562 RRLER--PPSPSPRLSYREGRPRDYDALSGSKRPYAAIDDISPRYADAGARQSRSRLDYD 619
+R ER PP P P LSYREGRPRDYDALSGSKRPYAAIDD+ PRYAD GARQSR+RLDYD
Sbjct: 11 QRFERAPPPPPPPHLSYREGRPRDYDALSGSKRPYAAIDDLPPRYADTGARQSRARLDYD 190
Query: 620 YGGSASRYREAYGDRLERSSLGY-SGSRSSLSNQNSHGAYSSRQDPSYGRGSLGGSD-GG 677
YG SAS+Y +AYGDRL RSS+GY GSRSS+S+Q+SHG YSSRQ SYG GS G D GG
Sbjct: 191 YGDSASQYGDAYGDRLGRSSVGYGGGSRSSISSQDSHGMYSSRQGMSYGGGSFGSGDVGG 370
Query: 678 IYSSSYGGDYLSRGSDVGGSSYSSMYSSRGMNGSSSYMSGSGGGSRSYY 726
+YSSSYGGDY+SRGSDVGGSSYSS+YS RG+ G SSYM GGGS SYY
Sbjct: 371 MYSSSYGGDYISRGSDVGGSSYSSVYSGRGVGGGSSYM--GGGGSGSYY 511
>AW471531 weakly similar to GP|20147226|gb| At2g44720/F16B22.21 {Arabidopsis
thaliana}, partial (7%)
Length = 342
Score = 145 bits (366), Expect = 6e-35
Identities = 72/112 (64%), Positives = 83/112 (73%), Gaps = 2/112 (1%)
Frame = +3
Query: 433 RSRPAPQSRPAHVFRRLGSRIPPARPSSARNRRPVTSIPVRARPASPPARSYRR--LAAA 490
R RPAP S P+ R +GSR PP R S R+RRP+ SIP R+RP PP+RSY R +A A
Sbjct: 6 RPRPAPHSFPSRSVRGVGSRTPPVRSVSVRDRRPMMSIPARSRPVPPPSRSYDRRPVAPA 185
Query: 491 YPKSSMKRDYGRPVDLPPPRSRVSADYGSQVVSQRGPSYRDYPARDSGYPDL 542
YPKSSMKRDYGR D+PPPRSRV+ DYGS+V S R PSYRDYPAR GY +L
Sbjct: 186 YPKSSMKRDYGRREDIPPPRSRVAVDYGSRVASVRRPSYRDYPARGPGYTEL 341
>TC212567 weakly similar to UP|PAB3_HUMAN (Q9H361) Polyadenylate-binding
protein 3 (Poly(A)-binding protein 3) (PABP 3)
(Testis-specific poly(A)-binding protein), partial (5%)
Length = 615
Score = 81.3 bits (199), Expect = 1e-15
Identities = 55/190 (28%), Positives = 98/190 (50%), Gaps = 8/190 (4%)
Frame = +3
Query: 123 EEEQVEVIKETQKQKEL--------EVFVGGLDKEATEHDLRKVFSKVGEITEVRMTVNP 174
+E +VE E +K EL EV++GG+ A++ DL+ + ++GE+ EVR+
Sbjct: 21 DEAKVEDEDEKRKHAELLSLPPHGSEVYIGGIP-HASDEDLKSLCERIGEVAEVRIMKGK 197
Query: 175 QTKRNKGFALLRFETVEHVKRALAELKNPVINGKQCGITITTCQDSDTLYLDNICKSWKK 234
NKGF + F +VE +A+ EL N GK+ I + Q L++ N+ +SW
Sbjct: 198 XFFENKGFGFVTFTSVELASKAIEELNNTEFMGKK--IKCSKSQAKHRLFIGNVPRSWGV 371
Query: 235 EALKEKLKHYGVESFKDLTLLEDDNNEGTNCGCAFLEFSSHSDSKDAYKRLQKTDVVFGV 294
E LK+ + G + L++D N N G AF+++ +H+ ++ + +++ G
Sbjct: 372 EDLKKIVTEIG-PGVTGVELVKDMKNTNNNRGFAFIDYYNHACAEYSRQKMMSPTFKLGE 548
Query: 295 DKPAEVSFAN 304
+ P VS+A+
Sbjct: 549 NAPT-VSWAD 575
Score = 34.7 bits (78), Expect = 0.15
Identities = 23/78 (29%), Positives = 43/78 (54%), Gaps = 1/78 (1%)
Frame = +3
Query: 320 VFIDLLPPSWDEDYVRALLKKYGAVEKVELAKNMPGARRKNYGFVTFGTHAAAVECAESI 379
V+I +P + DED +++L ++ G V +V + K K +GFVTF + A + E +
Sbjct: 99 VYIGGIPHASDED-LKSLCERIGEVAEVRIMKGKXFFENKGFGFVTFTSVELASKAIEEL 275
Query: 380 TSAG-LGEGNKKAKVRAR 396
+ +G+ K +K +A+
Sbjct: 276 NNTEFMGKKIKCSKSQAK 329
>CD414642 similar to GP|22655529|gb| water-stress protein {Zea mays}, partial
(7%)
Length = 351
Score = 67.8 bits (164), Expect = 2e-11
Identities = 33/42 (78%), Positives = 35/42 (82%)
Frame = -3
Query: 685 GDYLSRGSDVGGSSYSSMYSSRGMNGSSSYMSGSGGGSRSYY 726
GDY+SRGSDVGGSSYSSMYS RG+ G SSYM GGGS SYY
Sbjct: 343 GDYISRGSDVGGSSYSSMYSGRGVGGGSSYM--GGGGSGSYY 224
>TC204091 similar to UP|Q8LLE6 (Q8LLE6) RNA-binding protein AKIP1, partial
(70%)
Length = 2075
Score = 65.5 bits (158), Expect = 8e-11
Identities = 51/189 (26%), Positives = 92/189 (47%), Gaps = 6/189 (3%)
Frame = +2
Query: 36 EYDPEEYVGDDDYDERGIEQDEGQEAGDEVEEEPEENVDEEEGDTGDEEVEEVEYVYEEV 95
E P+EY + E +E++E +E +E EEE EE+ +EEE + +EE +V ++
Sbjct: 107 ENPPQEY----EEVEEEVEEEEVEEEVEEEEEEEEEDEEEEEEEEEEEEENDVVKQQQQQ 274
Query: 96 EGDDDDVAGDD--EHASEENVHEHLAD--VKEEEQVEVIKETQKQKEL--EVFVGGLDKE 149
+ D+DD+ E ++ + L D K + E I++ + ++FV GL +
Sbjct: 275 QQDEDDIPISKLLEPLGKDQLLNLLCDAAAKHRDVEERIRKAADGDPVHRKIFVHGLGWD 454
Query: 150 ATEHDLRKVFSKVGEITEVRMTVNPQTKRNKGFALLRFETVEHVKRALAELKNPVINGKQ 209
T L F + GEI + + + + ++KG+ + F+T + AL E + K+
Sbjct: 455 TTAGTLISAFRQYGEIEDCKAVTDKVSGKSKGYGFILFKTRRGAQNALKEPQ------KK 616
Query: 210 CGITITTCQ 218
G +T CQ
Sbjct: 617 IGNRMTACQ 643
Score = 35.0 bits (79), Expect = 0.12
Identities = 22/71 (30%), Positives = 35/71 (48%)
Frame = +2
Query: 129 VIKETQKQKELEVFVGGLDKEATEHDLRKVFSKVGEITEVRMTVNPQTKRNKGFALLRFE 188
V + TQK+ ++V + + L FS+ GEI E + ++ T + KGF L +
Sbjct: 716 VSEYTQKK----IYVSNVGADLDPQKLLAFFSRFGEIEEGPLGLDKATGKPKGFCLFVYR 883
Query: 189 TVEHVKRALAE 199
E +RAL E
Sbjct: 884 NPESARRALEE 916
>TC229446 similar to UP|Q8LLE6 (Q8LLE6) RNA-binding protein AKIP1, partial
(20%)
Length = 850
Score = 64.7 bits (156), Expect = 1e-10
Identities = 60/251 (23%), Positives = 110/251 (42%), Gaps = 33/251 (13%)
Frame = +3
Query: 1 MALNIYLIADNGEEVKESIDEYEKDEHLDFEDNYHEYDPEEYVGDDDY----DERGIEQD 56
MA L + + E K + + E ++ + + +PE+ + D + + + E++
Sbjct: 84 MAKKRKLRSSDPEPTKPIEPQQQDQEPVELDQQQQQPNPEQTLEDPNTMAVDEPKQEEEE 263
Query: 57 EGQEAGDEVEEEPEENVDEEEGDTGDEEVEEVEYVYEEVEGDDDDVAGDDEHASEENVHE 116
E +E + EE EE +E++ + E +EE + + +V G +E +EE E
Sbjct: 264 EDEEPQNAESEEEEEEAEEQQEERPPETLEEQQDPETLAAANHAEVNGGNEGNNEEEEEE 443
Query: 117 HLADVKEEEQVEVIKETQKQKEL-----------------------------EVFVGGLD 147
L EEE VE + E +++L ++FV GL
Sbjct: 444 DLT--LEEEPVEKLLEPFTKEQLHSLVTQAVDMFPEFVDSVRRLADVDPAHRKIFVHGLG 617
Query: 148 KEATEHDLRKVFSKVGEITEVRMTVNPQTKRNKGFALLRFETVEHVKRALAELKNPVING 207
+AT L VF K GEI + + + + ++KG+A + F+ + ++A LK+P
Sbjct: 618 WDATADTLTAVFGKYGEIEDCKAVTDKVSGKSKGYAFILFKHRDDARKA---LKHP---Q 779
Query: 208 KQCGITITTCQ 218
K+ G T+CQ
Sbjct: 780 KKIGNRTTSCQ 812
>TC226736 similar to UP|Q9SEE9 (Q9SEE9) Arginine/serine-rich protein, partial
(63%)
Length = 1243
Score = 61.6 bits (148), Expect = 1e-09
Identities = 86/287 (29%), Positives = 115/287 (39%), Gaps = 34/287 (11%)
Frame = +2
Query: 330 DEDYVRALLKKYGAVEKVELAKNMPGARRKNYGFVTFGTHAAAVECAESITSAGLGEGNK 389
+E +++ + +G V VELA + K YG+V F T A + + A +
Sbjct: 26 NEGHLKEIFSNFGEVISVELAMDRTVNLPKGYGYVQFKTRGEAEKALLYMDGAQIDGNVI 205
Query: 390 KAKV----RARLSRPLPKVRG----KHVPHRDISGRKIGRLERPSRSR-PAPRSRPAPQ- 439
KA+ R ++S P PK + P D + + + + P R R +PR +P P
Sbjct: 206 KARFTLPPRQKVSPP-PKTSAVAPKRDTPRTDNASADVDK-DGPKRQRESSPRRKPPPSP 379
Query: 440 SRPAHVFRRLGSRIPPARPSSARN------RRPVTS-------IPVRARPASP------- 479
R + V RR GS P RP S R RR V S P R RP SP
Sbjct: 380 RRRSPVPRRAGS---PRRPDSPRRRADSPVRRRVDSPYRRGGDTPPRRRPVSPGRGRSPS 550
Query: 480 PARSYRRLAAAYPKSSMKRDYGRPVDLPPPRSRVSADYGSQVVSQRGPSYR---DYPARD 536
P R R A P+ M+ GR PPP R S + +R P R P R
Sbjct: 551 PPRRLRSPARLSPR-RMRGSPGRRRSPPPPPRRRSPPRRPRSPPRRSPIGRRRTRSPIRR 727
Query: 537 SGYPDLHRDTSRT-APRRGYLDDGYDRRLERPPSPSPRLSYREGRPR 582
S R SR+ +PRRG R SPSPR R + R
Sbjct: 728 SA-----RSRSRSFSPRRGRPPVRRGRSSSYSDSPSPRKVSRRSKSR 853
Score = 35.4 bits (80), Expect = 0.090
Identities = 16/62 (25%), Positives = 32/62 (50%)
Frame = +2
Query: 146 LDKEATEHDLRKVFSKVGEITEVRMTVNPQTKRNKGFALLRFETVEHVKRALAELKNPVI 205
L + E L+++FS GE+ V + ++ KG+ ++F+T ++AL + I
Sbjct: 11 LSRNVNEGHLKEIFSNFGEVISVELAMDRTVNLPKGYGYVQFKTRGEAEKALLYMDGAQI 190
Query: 206 NG 207
+G
Sbjct: 191 DG 196
>TC232838 similar to GB|AAP21377.1|30102918|BT006569 At1g47970 {Arabidopsis
thaliana;} , partial (46%)
Length = 902
Score = 57.0 bits (136), Expect = 3e-08
Identities = 37/122 (30%), Positives = 60/122 (48%), Gaps = 9/122 (7%)
Frame = +2
Query: 13 EEVKESIDEYEKDEHLDFEDNYHEYDPEEYVGDDDYDE---------RGIEQDEGQEAGD 63
++ ++ DE E D+ + +DN E D GDDD DE RG E D+ D
Sbjct: 323 DDAEDDEDEEEDDDDDEGDDNDDEEDDAPDGGDDDDDEDDEEEGDVQRGGEPDDDDNDSD 502
Query: 64 EVEEEPEENVDEEEGDTGDEEVEEVEYVYEEVEGDDDDVAGDDEHASEENVHEHLADVKE 123
+ ++ E+ +E+E + G+EE EY+ +E +++ A D EEN E DV +
Sbjct: 503 DDSDDDEDEDEEDEEEQGEEEDLGTEYLIRPLETAEEEEASSD-FEPEENGEEEEEDVDD 679
Query: 124 EE 125
E+
Sbjct: 680 ED 685
Score = 54.3 bits (129), Expect = 2e-07
Identities = 40/152 (26%), Positives = 72/152 (47%), Gaps = 10/152 (6%)
Frame = +2
Query: 13 EEVKESIDEYEKDEHLDFEDNYHEYDPEEYVGDDDYDERGIEQDEGQEAGDEVEEEPEEN 72
+E+ E ++E E ++ +D EY+ + V DD D+ E+D+ + GD+ ++E E++
Sbjct: 233 KELSEQVEENEVKVLVEEDDE--EYESVDEVVDDAEDDEDEEEDDDDDEGDDNDDE-EDD 403
Query: 73 VDEEEGDTGDEEVEEVEYVYEEVEGDDDDVAGDDEHASEENVHEHLADVKEEEQ------ 126
+ D DE+ EE V E DDDD DD+ +E+ E + + EE+
Sbjct: 404 APDGGDDDDDEDDEEEGDVQRGGEPDDDDNDSDDDSDDDEDEDEEDEEEQGEEEDLGTEY 583
Query: 127 ----VEVIKETQKQKELEVFVGGLDKEATEHD 154
+E +E + + E G ++E D
Sbjct: 584 LIRPLETAEEEEASSDFEPEENGEEEEEDVDD 679
Score = 52.0 bits (123), Expect = 9e-07
Identities = 40/124 (32%), Positives = 59/124 (47%), Gaps = 22/124 (17%)
Frame = +2
Query: 10 DNGEEVKESIDEYEKDEHL------DFEDNYHEYDPEEYVGDDDYDERGIEQDEGQE--- 60
D G++ + DE E D D D+ + D +E ++D +E+G E+D G E
Sbjct: 410 DGGDDDDDEDDEEEGDVQRGGEPDDDDNDSDDDSDDDEDEDEEDEEEQGEEEDLGTEYLI 589
Query: 61 ----AGDEVEE----EPEENVDEEEGDTGDEEVEEVEYV-----YEEVEGDDDDVAGDDE 107
+E E EPEEN +EEE D DE+ E+ E ++ + DDDD DDE
Sbjct: 590 RPLETAEEEEASSDFEPEENGEEEEEDVDDEDCEKAEAPPKRKRSDKDDSDDDDGGEDDE 769
Query: 108 HASE 111
S+
Sbjct: 770 RPSK 781
Score = 39.7 bits (91), Expect = 0.005
Identities = 20/61 (32%), Positives = 33/61 (53%)
Frame = +2
Query: 56 DEGQEAGDEVEEEPEENVDEEEGDTGDEEVEEVEYVYEEVEGDDDDVAGDDEHASEENVH 115
D +A E+ E+ EEN + + DEE E V+ V ++ E D+D+ DD+ ++N
Sbjct: 212 DSKMKATKELSEQVEENEVKVLVEEDDEEYESVDEVVDDAEDDEDEEEDDDDDEGDDNDD 391
Query: 116 E 116
E
Sbjct: 392 E 394
Score = 30.4 bits (67), Expect = 2.9
Identities = 47/160 (29%), Positives = 59/160 (36%)
Frame = -3
Query: 563 RLERPPSPSPRLSYREGRPRDYDALSGSKRPYAAIDDISPRYADAGARQSRSRLDYDYGG 622
RLE S SP S E D G+ + S + + S+S LD
Sbjct: 783 RLEGLSSSSPPSSSSESSLSDLFLFGGASAFSQSSSSTSSSSSSPFSSGSKS-LDASSSS 607
Query: 623 SASRYREAYGDRLERSSLGYSGSRSSLSNQNSHGAYSSRQDPSYGRGSLGGSDGGIYSSS 682
+ S+ R Y + SS S S SS S+ +S + S S G S SSS
Sbjct: 606 AVSKGRIRYSVP-KSSSSPCSSSSSSSSSSSSSESSSESLSSSSGSPPRWTSPSSSSSSS 430
Query: 683 YGGDYLSRGSDVGGSSYSSMYSSRGMNGSSSYMSGSGGGS 722
S G SS SS SS + SSS S S S
Sbjct: 429 -----SSSSPPSGASSSSSSLSSPSSSSSSSSSSSSSSAS 325
>TC219445 similar to UP|P93396 (P93396) Transformer-SR ribonucleoprotein
(Fragment), partial (51%)
Length = 735
Score = 53.5 bits (127), Expect = 3e-07
Identities = 24/68 (35%), Positives = 39/68 (57%)
Frame = +1
Query: 141 VFVGGLDKEATEHDLRKVFSKVGEITEVRMTVNPQTKRNKGFALLRFETVEHVKRALAEL 200
++V GL TE DL + FSK G++ + V P+T+ ++GFA + +TVE R + L
Sbjct: 250 LYVTGLSSRVTERDLEEHFSKEGKVASCFLVVEPRTRISRGFAFITMDTVEDANRCIKYL 429
Query: 201 KNPVINGK 208
V+ G+
Sbjct: 430 NQSVLEGR 453
>TC215891 similar to GB|AAM78058.1|21928059|AY125548 AT5g61030/maf19_30
{Arabidopsis thaliana;} , partial (41%)
Length = 782
Score = 53.1 bits (126), Expect = 4e-07
Identities = 22/81 (27%), Positives = 44/81 (54%)
Frame = +2
Query: 128 EVIKETQKQKELEVFVGGLDKEATEHDLRKVFSKVGEITEVRMTVNPQTKRNKGFALLRF 187
+ I+ ++F+GG+ E LR+ FSK GE+ + R+ ++ +T R++GF + +
Sbjct: 140 QAIRSMSSAPSTKLFIGGVSYSTDEQSLREAFSKYGEVVDARIIMDRETGRSRGFGFITY 319
Query: 188 ETVEHVKRALAELKNPVINGK 208
+VE A+ L ++G+
Sbjct: 320 TSVEEASSAIQALDGQDLHGR 382
Score = 32.3 bits (72), Expect = 0.76
Identities = 23/87 (26%), Positives = 37/87 (42%)
Frame = +2
Query: 320 VFIDLLPPSWDEDYVRALLKKYGAVEKVELAKNMPGARRKNYGFVTFGTHAAAVECAESI 379
+FI + S DE +R KYG V + + R + +GF+T+ +VE A S
Sbjct: 179 LFIGGVSYSTDEQSLREAFSKYGEVVDARIIMDRETGRSRGFGFITY----TSVEEASSA 346
Query: 380 TSAGLGEGNKKAKVRARLSRPLPKVRG 406
A G+ +R + P+ G
Sbjct: 347 IQALDGQDLHGRPIRVNYANERPRGYG 427
Score = 30.4 bits (67), Expect = 2.9
Identities = 24/85 (28%), Positives = 34/85 (39%), Gaps = 5/85 (5%)
Frame = +2
Query: 643 SGSRSSLSNQNSHGA-----YSSRQDPSYGRGSLGGSDGGIYSSSYGGDYLSRGSDVGGS 697
S + +L Q+ HG Y++ + YG G G Y + GG Y GGS
Sbjct: 338 SSAIQALDGQDLHGRPIRVNYANERPRGYGGGGFGS-----YGAVGGGGY------EGGS 484
Query: 698 SYSSMYSSRGMNGSSSYMSGSGGGS 722
SY Y + + G GGG+
Sbjct: 485 SYRGGYGGDNYSRNDGSGYGYGGGN 559
>TC209416 similar to UP|Q8LF90 (Q8LF90) RNA recognition motif-containing
protein SEB-4, partial (42%)
Length = 1327
Score = 53.1 bits (126), Expect = 4e-07
Identities = 27/77 (35%), Positives = 44/77 (57%)
Frame = +1
Query: 140 EVFVGGLDKEATEHDLRKVFSKVGEITEVRMTVNPQTKRNKGFALLRFETVEHVKRALAE 199
+VFVGGL E ++RK F + G+I E + + T ++KG+ + F E +RA A+
Sbjct: 559 KVFVGGLAWETPTEEMRKYFEQFGDILEAVIITDKSTGKSKGYGFVTFRDPESARRACAD 738
Query: 200 LKNPVINGKQCGITITT 216
NPVI+G++ I +
Sbjct: 739 -PNPVIDGRRANCNIAS 786
Score = 30.8 bits (68), Expect = 2.2
Identities = 57/264 (21%), Positives = 88/264 (32%), Gaps = 3/264 (1%)
Frame = +1
Query: 320 VFIDLLPPSWDEDYVRALLKKYGAVEKVELAKNMPGARRKNYGFVTFGTHAAAVECAESI 379
VF+ L + +R +++G + + + + + K YGFVTF +A
Sbjct: 562 VFVGGLAWETPTEEMRKYFEQFGDILEAVIITDKSTGKSKGYGFVTFRDPESA------- 720
Query: 380 TSAGLGEGNKKAKVRARLSRPLPKVRGKHVPHRDISGRKIGRLERPSRSRPAPRSRPAPQ 439
R + P P + G+ + I L RP S PR R Q
Sbjct: 721 --------------RRACADPNPVIDGRR------ANCNIASLGRPRPS--PPRGRGTFQ 834
Query: 440 SRPAHVFRRLGSRIPPARPSSARNRRPVTSIPVRARPASPPARSYRRLAAAYPKSSMKRD 499
+P A P + P P P P Y + + P+ +
Sbjct: 835 GGAPVTAAASYGGVPAAGPPALAPPPPPLVYPPYGYPTYTPDYGYHQASLYNPQIQQPQY 1014
Query: 500 YGRPVDLPPPRSRVSA-DYGSQVVSQRGPSYRDYPARDSGYPD-LHRDTSRTAPRRGYLD 557
Y + + P + S YG V + RG P R P L+ T + Y
Sbjct: 1015YHQQLYGPSSSTMGSPYYYGYSVQAPRGTFSSTQPHRIPAGPSYLYYPTPMESSFSAYRP 1194
Query: 558 DGYDRRLE-RPPSPSPRLSYREGR 580
++L R P PSP S + R
Sbjct: 1195PPQLQQLPIRQPPPSPSDSQTQQR 1266
>TC215689 similar to UP|P93396 (P93396) Transformer-SR ribonucleoprotein
(Fragment), partial (51%)
Length = 545
Score = 52.0 bits (123), Expect = 9e-07
Identities = 23/68 (33%), Positives = 40/68 (58%)
Frame = +1
Query: 141 VFVGGLDKEATEHDLRKVFSKVGEITEVRMTVNPQTKRNKGFALLRFETVEHVKRALAEL 200
++V GL TE DL + FSK G+++ + V P+T+ ++GFA + E+ E +R + L
Sbjct: 52 LYVTGLSSRVTERDLEEHFSKEGKVSSCFLVVEPRTRISRGFAFVTMESAEDAERCIKYL 231
Query: 201 KNPVINGK 208
V+ G+
Sbjct: 232 NQSVLEGR 255
>TC215684 similar to UP|P93396 (P93396) Transformer-SR ribonucleoprotein
(Fragment), partial (84%)
Length = 1263
Score = 52.0 bits (123), Expect = 9e-07
Identities = 23/68 (33%), Positives = 40/68 (58%)
Frame = +2
Query: 141 VFVGGLDKEATEHDLRKVFSKVGEITEVRMTVNPQTKRNKGFALLRFETVEHVKRALAEL 200
++V GL TE DL + FSK G+++ + V P+T+ ++GFA + E+ E +R + L
Sbjct: 269 LYVTGLSSRVTERDLEEHFSKEGKVSSCFLVVEPRTRISRGFAFVTMESAEDAERCIKYL 448
Query: 201 KNPVINGK 208
V+ G+
Sbjct: 449 NQSVLEGR 472
Score = 29.3 bits (64), Expect = 6.5
Identities = 20/54 (37%), Positives = 25/54 (46%)
Frame = +2
Query: 426 SRSRPAPRSRPAPQSRPAHVFRRLGSRIPPARPSSARNRRPVTSIPVRARPASP 479
SRSR RSR P+ RP R G P R R+R P+ R+ P +P
Sbjct: 116 SRSRSRSRSRSRPRQRPRSRSRSRGRSRSPIR---GRSRSPIHG---RSEPTNP 259
>TC215688 similar to UP|P93396 (P93396) Transformer-SR ribonucleoprotein
(Fragment), partial (61%)
Length = 614
Score = 52.0 bits (123), Expect = 9e-07
Identities = 23/68 (33%), Positives = 40/68 (58%)
Frame = +1
Query: 141 VFVGGLDKEATEHDLRKVFSKVGEITEVRMTVNPQTKRNKGFALLRFETVEHVKRALAEL 200
++V GL TE DL + FSK G+++ + V P+T+ ++GFA + E+ E +R + L
Sbjct: 154 LYVTGLSSRVTERDLEEHFSKEGKVSSCFLVVEPRTRISRGFAFVTMESAEDAERCIKYL 333
Query: 201 KNPVINGK 208
V+ G+
Sbjct: 334 NQSVLEGR 357
>TC215791 similar to GB|CAE76446.1|38567152|BX842633 related to peroxisomal
membrane protein pex13 {Neurospora crassa;} , partial
(9%)
Length = 1440
Score = 51.6 bits (122), Expect = 1e-06
Identities = 46/156 (29%), Positives = 61/156 (38%)
Frame = +3
Query: 570 PSPRLSYREGRPRDYDALSGSKRPYAAIDDISPRYADAGARQSRSRLDYDYGGSASRYRE 629
P+P G D SG+ +P + +D A +R+ L + +
Sbjct: 189 PAPFKPPSAGNTSDVVEASGTAKPGEIVSS-----SDRSAAVNRNTLGRPV--PTRPWEQ 347
Query: 630 AYGDRLERSSLGYSGSRSSLSNQNSHGAYSSRQDPSYGRGSLGGSDGGIYSSSYGGDYLS 689
YG+ S+ G GS + ++ G Y S S GG GG+Y SSYGG
Sbjct: 348 NYGN----STYGGYGSTMNYNSGYGSGMYGS---------SYGGLGGGMYGSSYGGGMYG 488
Query: 690 RGSDVGGSSYSSMYSSRGMNGSSSYMSGSGGGSRSY 725
GG Y MY S GM G Y SG GG Y
Sbjct: 489 NSMYRGG--YGGMYGSSGMYGGGMYNSGLGGPMGGY 590
>TC225641 similar to GB|AAA18379.1|475719|ATU08467 RNA-binding protein 2
{Arabidopsis thaliana;} , partial (59%)
Length = 1223
Score = 51.2 bits (121), Expect = 2e-06
Identities = 54/240 (22%), Positives = 91/240 (37%), Gaps = 13/240 (5%)
Frame = +3
Query: 59 QEAGDEVEEEPEENVDEEEGDT--GDEEVEEVEYVYEEVEGDDDDVAGDDEHASEENVHE 116
Q +G E E EE E +GDT G EE GDD + G+ A EE
Sbjct: 282 QTSGWAKEGEEEEAAWENQGDTAWGTEE-----------GGDDIEDGGEGGFAEEE---- 416
Query: 117 HLADVKEEEQVEVIKETQKQKELEVFVGGLDKEATEHDLRKVFSKVGEITEVRMTVNPQT 176
+E+++FVG L + L +F + G + + N T
Sbjct: 417 -------------------AEEVKIFVGNLPFDFDSEKLASLFEQAGTVEVAEVIYNRAT 539
Query: 177 KRNKGFALLRFETVEHVKRALAELKNPVINGKQCGITITTCQ-----------DSDTLYL 225
R++GF + T+E +++A+ +NG+ + + S +Y+
Sbjct: 540 DRSRGFGFVTMSTIEELEKAVKMFSGYELNGRVLTVNKAAPKGAQPERPPRPPQSFRVYV 719
Query: 226 DNICKSWKKEALKEKLKHYGVESFKDLTLLEDDNNEGTNCGCAFLEFSSHSDSKDAYKRL 285
N+ L++ +G +D ++ D G + G F+ SS +D DA L
Sbjct: 720 GNLPWDVDNSRLEQIFSEHG--KVEDARVVY-DRETGRSRGFGFVTMSSETDMNDAIAAL 890
Score = 39.3 bits (90), Expect = 0.006
Identities = 17/78 (21%), Positives = 40/78 (50%)
Frame = +3
Query: 137 KELEVFVGGLDKEATEHDLRKVFSKVGEITEVRMTVNPQTKRNKGFALLRFETVEHVKRA 196
+ V+VG L + L ++FS+ G++ + R+ + +T R++GF + + + A
Sbjct: 699 QSFRVYVGNLPWDVDNSRLEQIFSEHGKVEDARVVYDRETGRSRGFGFVTMSSETDMNDA 878
Query: 197 LAELKNPVINGKQCGITI 214
+A L ++G+ + +
Sbjct: 879 IAALDGQSLDGRAIRVNV 932
>TC227797 similar to UP|Q8H1N8 (Q8H1N8) At1g20880/F9H16_14, partial (59%)
Length = 1737
Score = 50.4 bits (119), Expect = 3e-06
Identities = 26/77 (33%), Positives = 43/77 (55%)
Frame = +2
Query: 140 EVFVGGLDKEATEHDLRKVFSKVGEITEVRMTVNPQTKRNKGFALLRFETVEHVKRALAE 199
+VFVGGL E +R+ F + GEI E + + T R+KG+ + F E +RA A+
Sbjct: 356 KVFVGGLAWETQSETMRRYFDQFGEILEAVVITDKNTGRSKGYGFVTFRDPEAARRACAD 535
Query: 200 LKNPVINGKQCGITITT 216
+PVI+G++ + +
Sbjct: 536 -PSPVIDGRRANCNLAS 583
>TC223521 weakly similar to UP|Q84ZB9 (Q84ZB9) RNA recognition motif
(RRM)-containing protein-like, partial (57%)
Length = 651
Score = 50.4 bits (119), Expect = 3e-06
Identities = 21/69 (30%), Positives = 41/69 (58%)
Frame = +1
Query: 140 EVFVGGLDKEATEHDLRKVFSKVGEITEVRMTVNPQTKRNKGFALLRFETVEHVKRALAE 199
++FVGGL TE+ L + FS G++ E ++ + + R+KGF + F + + + A+ +
Sbjct: 82 KLFVGGLSFYTTENALSEAFSNYGQVIEAKIVTDRVSDRSKGFGFVTFASQDEAENAIED 261
Query: 200 LKNPVINGK 208
+K +NG+
Sbjct: 262 MKGKTLNGR 288
>TC222793 similar to UP|Q8LF90 (Q8LF90) RNA recognition motif-containing
protein SEB-4, partial (34%)
Length = 566
Score = 49.7 bits (117), Expect = 5e-06
Identities = 25/77 (32%), Positives = 44/77 (56%)
Frame = +2
Query: 140 EVFVGGLDKEATEHDLRKVFSKVGEITEVRMTVNPQTKRNKGFALLRFETVEHVKRALAE 199
++FVGGL E ++RK F + G+I E + + T ++KG+ + F E +RA A+
Sbjct: 287 KLFVGGLAWETPTEEMRKYFEQFGDILEAVIITDKNTGKSKGYGFVTFCGQESARRACAD 466
Query: 200 LKNPVINGKQCGITITT 216
NP+I+G++ I +
Sbjct: 467 -PNPIIDGRRANCNIAS 514
>TC204084 UP|Q9SWA8 (Q9SWA8) Glycine-rich RNA-binding protein, complete
Length = 1937
Score = 49.7 bits (117), Expect = 5e-06
Identities = 29/100 (29%), Positives = 50/100 (50%), Gaps = 4/100 (4%)
Frame = +2
Query: 123 EEEQVEV----IKETQKQKELEVFVGGLDKEATEHDLRKVFSKVGEITEVRMTVNPQTKR 178
E+ +VEV E FVGGL +H L + FS+ GEI E ++ + +T R
Sbjct: 1181 EQREVEVGFMQFSMASADVEYRCFVGGLAWATDDHALERAFSQYGEIVETKIINDRETGR 1360
Query: 179 NKGFALLRFETVEHVKRALAELKNPVINGKQCGITITTCQ 218
++GF + F + + +K A+ + ++G+ IT+ Q
Sbjct: 1361 SRGFGFVTFASEQSMKDAIGAMNGQNLDGR--NITVNEAQ 1474
Score = 36.6 bits (83), Expect = 0.040
Identities = 17/72 (23%), Positives = 35/72 (48%)
Frame = -1
Query: 137 KELEVFVGGLDKEATEHDLRKVFSKVGEITEVRMTVNPQTKRNKGFALLRFETVEHVKRA 196
++L++FVG L L ++F G + V + + T R++GF + +VE + A
Sbjct: 887 RDLKLFVGNLPFSVDSARLAELFESAGNVEVVEVIYDKTTGRSRGFGFVTMSSVEEAEAA 708
Query: 197 LAELKNPVINGK 208
+ ++G+
Sbjct: 707 AKQFNGYELDGR 672
Score = 34.3 bits (77), Expect = 0.20
Identities = 20/78 (25%), Positives = 40/78 (50%), Gaps = 1/78 (1%)
Frame = -1
Query: 138 ELEVFVGGLDKEATEHDLRKVFSKVGE-ITEVRMTVNPQTKRNKGFALLRFETVEHVKRA 196
E V VG L + L +F + G+ + E R+ + ++ R++GF + F + + VK A
Sbjct: 566 ENRVHVGNLAWGVDDVALESLFREQGKKVLEARVIYDRESGRSRGFGFVTFGSPDEVKSA 387
Query: 197 LAELKNPVINGKQCGITI 214
+ L +NG+ +++
Sbjct: 386 IQSLDGVDLNGRAIRVSL 333
Database: GMGI
Posted date: Oct 22, 2004 4:58 PM
Number of letters in database: 37,918,896
Number of sequences in database: 63,676
Lambda K H
0.311 0.133 0.379
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 28,214,908
Number of Sequences: 63676
Number of extensions: 451004
Number of successful extensions: 5448
Number of sequences better than 10.0: 552
Number of HSP's better than 10.0 without gapping: 4025
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 4813
length of query: 726
length of database: 12,639,632
effective HSP length: 104
effective length of query: 622
effective length of database: 6,017,328
effective search space: 3742778016
effective search space used: 3742778016
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 16 ( 7.2 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 42 (21.8 bits)
S2: 62 (28.5 bits)
Medicago: description of AC149129.7


