

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC149127.7 - phase: 0
(360 letters)
Database: GMGI
63,676 sequences; 37,918,896 total letters
Searching..................................................done
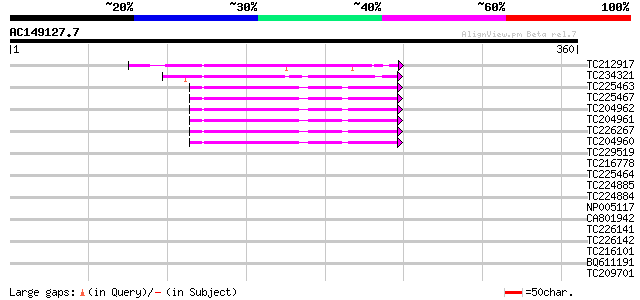
Score E
Sequences producing significant alignments: (bits) Value
TC212917 weakly similar to UP|Q9LX27 (Q9LX27) Calmodulin-like pr... 43 3e-04
TC234321 similar to UP|Q40982 (Q40982) Calmodulin-like protein, ... 41 7e-04
TC225463 UP|Q6LEC4 (Q6LEC4) Calmodulin, complete 41 7e-04
TC225467 UP|O49184 (O49184) Calmodulin, complete 41 7e-04
TC204962 UP|Q6LEC4 (Q6LEC4) Calmodulin, complete 41 7e-04
TC204961 UP|Q6LEC4 (Q6LEC4) Calmodulin, complete 41 7e-04
TC226267 UP|Q6LEC4 (Q6LEC4) Calmodulin, complete 41 7e-04
TC204960 UP|Q6LEC4 (Q6LEC4) Calmodulin, complete 41 7e-04
TC229519 weakly similar to UP|Q93YA8 (Q93YA8) Calcium binding pr... 41 0.001
TC216778 UP|O24431 (O24431) Calmodulin-like domain protein kinas... 39 0.004
TC225464 GB|AAB68399.1|1773321|HAU79736 calmodulin {Helianthus a... 37 0.018
TC224885 similar to UP|Q93YI3 (Q93YI3) Calcium dependent calmodu... 36 0.031
TC224884 similar to UP|Q93YI3 (Q93YI3) Calcium dependent calmodu... 35 0.040
NP005117 51 kDa seed maturation protein 34 0.090
CA801942 34 0.12
TC226141 similar to UP|Q93YA8 (Q93YA8) Calcium binding protein, ... 33 0.26
TC226142 similar to UP|Q93YA8 (Q93YA8) Calcium binding protein, ... 33 0.26
TC216101 similar to UP|Q9SCA1 (Q9SCA1) Calcium-binding protein, ... 31 0.76
BQ611191 similar to GP|4589960|gb|A expressed protein {Arabidops... 31 1.00
TC209701 similar to UP|Q8LDS1 (Q8LDS1) Calcium-dependent protein... 31 1.00
>TC212917 weakly similar to UP|Q9LX27 (Q9LX27) Calmodulin-like protein,
partial (49%)
Length = 1086
Score = 42.7 bits (99), Expect = 3e-04
Identities = 52/183 (28%), Positives = 75/183 (40%), Gaps = 8/183 (4%)
Frame = +1
Query: 76 NVAPRTGLGSTTTVSEIKETYEYLTSGGTLNTTLRLIILFPLLDRDPKDGFVGFNELESW 135
N+AP S TT + G+ L LF D++ DGF+ EL
Sbjct: 244 NLAPSPSPSSPTTK---------MAESGSQKKKEELRKLFSTFDKNG-DGFITKQELRES 393
Query: 136 VTQRALERLDYATQVELESKDKNGDLALSFREYLPDLSE----KDIEKKNMAHG-EAGWL 190
+ + D + D N D + F E+ SE EK+ G E L
Sbjct: 394 LRNIGIFMADKEVDDIVVKYDSNSDGLIDFEEFCLLTSECVGGDHHEKEGGVMGNEEVDL 573
Query: 191 MEKFDVADYDHNGLLNFTELRDFLHP---EDSQNKEMLKWMVNDKFNMDDYEHDGKINFN 247
E FDV D D++GL++ EL L + + E K M+ K +MD DG +NFN
Sbjct: 574 KEAFDVFDKDNDGLISVEELALVLTSLGLREGRKIEECKEMIK-KVDMDG---DGMVNFN 741
Query: 248 QFE 250
+F+
Sbjct: 742 EFK 750
>TC234321 similar to UP|Q40982 (Q40982) Calmodulin-like protein, partial
(96%)
Length = 626
Score = 41.2 bits (95), Expect = 7e-04
Identities = 41/159 (25%), Positives = 70/159 (43%), Gaps = 7/159 (4%)
Frame = +3
Query: 98 YLTSGGTLNTTLR------LIILFPLLDRDPKDGFVGFNELESWVTQRALERLDYATQVE 151
Y G ++ LR + F L D+D DG + EL + + Q+
Sbjct: 48 YYIKGSSMKDALREEQIGEFLEAFCLFDKDG-DGCITIEELSTAIRSLDENPTVEELQIM 224
Query: 152 LESKDKNGDLALSFREYLPDLSEKDIEKKNMAHGEAGWLMEKFDVADYDHNGLLNFTELR 211
+ D +G+ + F E+L ++ K K+ A E L E F V D DH+G ++ +ELR
Sbjct: 225 MNEVDMDGNGTIEFGEFLNLMARK--MKETEAEEE---LKEAFRVFDKDHDGYISPSELR 389
Query: 212 DFLHP-EDSQNKEMLKWMVNDKFNMDDYEHDGKINFNQF 249
+ + E ++ MV + D + DG I++ +F
Sbjct: 390 SVMRTIGEKVTDEEVEQMVKEA----DLDGDGLIDYEEF 494
>TC225463 UP|Q6LEC4 (Q6LEC4) Calmodulin, complete
Length = 946
Score = 41.2 bits (95), Expect = 7e-04
Identities = 34/135 (25%), Positives = 60/135 (44%)
Frame = +1
Query: 115 FPLLDRDPKDGFVGFNELESWVTQRALERLDYATQVELESKDKNGDLALSFREYLPDLSE 174
F L D+D DG + EL + + + Q + D +G+ + F E+L ++
Sbjct: 175 FSLFDKDG-DGCITTKELGTVMRSLGQNPTEAELQDMINEVDADGNGTIDFPEFLNLMAR 351
Query: 175 KDIEKKNMAHGEAGWLMEKFDVADYDHNGLLNFTELRDFLHPEDSQNKEMLKWMVNDKFN 234
K + + L E F V D D NG ++ ELR H + +++ V++
Sbjct: 352 KMKDTDSEEE-----LKEAFRVFDKDQNGFISAAELR---HVMTNLGEKLTDEEVDEMIR 507
Query: 235 MDDYEHDGKINFNQF 249
D + DG+IN+ +F
Sbjct: 508 EADVDGDGQINYEEF 552
>TC225467 UP|O49184 (O49184) Calmodulin, complete
Length = 1048
Score = 41.2 bits (95), Expect = 7e-04
Identities = 34/135 (25%), Positives = 60/135 (44%)
Frame = +3
Query: 115 FPLLDRDPKDGFVGFNELESWVTQRALERLDYATQVELESKDKNGDLALSFREYLPDLSE 174
F L D+D DG + EL + + + Q + D +G+ + F E+L ++
Sbjct: 174 FSLFDKDG-DGCITTKELGTVMRSLGQNPTEAELQDMINEVDADGNGTIDFPEFLNLMAR 350
Query: 175 KDIEKKNMAHGEAGWLMEKFDVADYDHNGLLNFTELRDFLHPEDSQNKEMLKWMVNDKFN 234
K + + L E F V D D NG ++ ELR H + +++ V++
Sbjct: 351 KMKDTDSEEE-----LKEAFRVFDKDQNGFISAAELR---HVMTNLGEKLTDEEVDEMIR 506
Query: 235 MDDYEHDGKINFNQF 249
D + DG+IN+ +F
Sbjct: 507 EADVDGDGQINYEEF 551
>TC204962 UP|Q6LEC4 (Q6LEC4) Calmodulin, complete
Length = 785
Score = 41.2 bits (95), Expect = 7e-04
Identities = 34/135 (25%), Positives = 60/135 (44%)
Frame = +3
Query: 115 FPLLDRDPKDGFVGFNELESWVTQRALERLDYATQVELESKDKNGDLALSFREYLPDLSE 174
F L D+D DG + EL + + + Q + D +G+ + F E+L ++
Sbjct: 144 FSLFDKDG-DGCITTKELGTVMRSLGQNPTEAELQDMINEVDADGNGTIDFPEFLNLMAR 320
Query: 175 KDIEKKNMAHGEAGWLMEKFDVADYDHNGLLNFTELRDFLHPEDSQNKEMLKWMVNDKFN 234
K + + L E F V D D NG ++ ELR H + +++ V++
Sbjct: 321 KMKDTDSEEE-----LKEAFRVFDKDQNGFISAAELR---HVMTNLGEKLTDEEVDEMIR 476
Query: 235 MDDYEHDGKINFNQF 249
D + DG+IN+ +F
Sbjct: 477 EADVDGDGQINYEEF 521
>TC204961 UP|Q6LEC4 (Q6LEC4) Calmodulin, complete
Length = 928
Score = 41.2 bits (95), Expect = 7e-04
Identities = 34/135 (25%), Positives = 60/135 (44%)
Frame = +2
Query: 115 FPLLDRDPKDGFVGFNELESWVTQRALERLDYATQVELESKDKNGDLALSFREYLPDLSE 174
F L D+D DG + EL + + + Q + D +G+ + F E+L ++
Sbjct: 143 FSLFDKDG-DGCITTKELGTVMRSLGQNPTEAELQDMINEVDADGNGTIDFPEFLNLMAR 319
Query: 175 KDIEKKNMAHGEAGWLMEKFDVADYDHNGLLNFTELRDFLHPEDSQNKEMLKWMVNDKFN 234
K + + L E F V D D NG ++ ELR H + +++ V++
Sbjct: 320 KMKDTDSEEE-----LKEAFRVFDKDQNGFISAAELR---HVMTNLGEKLTDEEVDEMIR 475
Query: 235 MDDYEHDGKINFNQF 249
D + DG+IN+ +F
Sbjct: 476 EADVDGDGQINYEEF 520
>TC226267 UP|Q6LEC4 (Q6LEC4) Calmodulin, complete
Length = 895
Score = 41.2 bits (95), Expect = 7e-04
Identities = 34/135 (25%), Positives = 60/135 (44%)
Frame = +3
Query: 115 FPLLDRDPKDGFVGFNELESWVTQRALERLDYATQVELESKDKNGDLALSFREYLPDLSE 174
F L D+D DG + EL + + + Q + D +G+ + F E+L ++
Sbjct: 225 FSLFDKDG-DGCITTKELGTVMRSLGQNPTEAELQDMINEVDADGNGTIDFPEFLNLMAR 401
Query: 175 KDIEKKNMAHGEAGWLMEKFDVADYDHNGLLNFTELRDFLHPEDSQNKEMLKWMVNDKFN 234
K + + L E F V D D NG ++ ELR H + +++ V++
Sbjct: 402 KMKDTDSEEE-----LKEAFRVFDKDQNGFISAAELR---HVMTNLGEKLTDEEVDEMIR 557
Query: 235 MDDYEHDGKINFNQF 249
D + DG+IN+ +F
Sbjct: 558 EADVDGDGQINYEEF 602
>TC204960 UP|Q6LEC4 (Q6LEC4) Calmodulin, complete
Length = 1012
Score = 41.2 bits (95), Expect = 7e-04
Identities = 34/135 (25%), Positives = 60/135 (44%)
Frame = +2
Query: 115 FPLLDRDPKDGFVGFNELESWVTQRALERLDYATQVELESKDKNGDLALSFREYLPDLSE 174
F L D+D DG + EL + + + Q + D +G+ + F E+L ++
Sbjct: 137 FSLFDKDG-DGCITTKELGTVMRSLGQNPTEAELQDMINEVDADGNGTIDFPEFLNLMAR 313
Query: 175 KDIEKKNMAHGEAGWLMEKFDVADYDHNGLLNFTELRDFLHPEDSQNKEMLKWMVNDKFN 234
K + + L E F V D D NG ++ ELR H + +++ V++
Sbjct: 314 KMKDTDSEEE-----LKEAFRVFDKDQNGFISAAELR---HVMTNLGEKLTDEEVDEMIR 469
Query: 235 MDDYEHDGKINFNQF 249
D + DG+IN+ +F
Sbjct: 470 EADVDGDGQINYEEF 514
>TC229519 weakly similar to UP|Q93YA8 (Q93YA8) Calcium binding protein,
partial (67%)
Length = 1060
Score = 40.8 bits (94), Expect = 0.001
Identities = 40/170 (23%), Positives = 69/170 (40%), Gaps = 12/170 (7%)
Frame = +3
Query: 93 KETYEYLTSGGTLNTTLRLIILFPLLDRDPK----------DGFVGFNELESWVTQRALE 142
K E LT + + +ILF ++D + + DG + EL+ + +
Sbjct: 18 KRKGETLTHSRP*DFSSSFLILFLIMDVEVRQIFNKFDKNGDGKISVTELKDMLAALGSK 197
Query: 143 RLDYATQVELESKDKNGDLALSFREYLPDLSEKDIEKKNMAHGEAGWLMEKFDVADYDHN 202
D + +E D+NGD + +E+ K + L + FD+ D D N
Sbjct: 198 TTDEELKRMMEELDQNGDGFIDLKEFADFHCNGGAGKDDSKE-----LRDAFDLYDVDKN 362
Query: 203 GLLNFTELRDFLH--PEDSQNKEMLKWMVNDKFNMDDYEHDGKINFNQFE 250
GL++ EL D L E + + + N D + DG +NF +F+
Sbjct: 363 GLISAKELHDVLRNLGEKCSLSDCRRMISN-----VDADGDGNVNFEEFK 497
>TC216778 UP|O24431 (O24431) Calmodulin-like domain protein kinase isoenzyme
gamma , complete
Length = 2377
Score = 38.9 bits (89), Expect = 0.004
Identities = 31/141 (21%), Positives = 71/141 (49%), Gaps = 5/141 (3%)
Frame = +1
Query: 114 LFPLLDRDPKDGFVGFNELESWVTQRALERLDYATQVELESKDKNGDLALSFREYLP-DL 172
+F +D D K G + + EL+S + + + + + +E+ D +G+ ++ + E++ +
Sbjct: 1615 MFTNMDTD-KSGTITYEELKSGLHRLGSKLTEAEVKQLMEAADVDGNGSIDYIEFITATM 1791
Query: 173 SEKDIEKKNMAHGEAGWLMEKFDVADYDHNGLLNFTEL----RDFLHPEDSQNKEMLKWM 228
+E+ + L + F D D++G + EL +++ +D+ KE++
Sbjct: 1792 HRHKLERDDQ-------LFKAFQYFDKDNSGFITRDELESAMKEYGMGDDATIKEIIS-E 1947
Query: 229 VNDKFNMDDYEHDGKINFNQF 249
V+ + D +HDG+IN+ +F
Sbjct: 1948 VDTIISEVDTDHDGRINYEEF 2010
>TC225464 GB|AAB68399.1|1773321|HAU79736 calmodulin {Helianthus annuus;} ,
partial (79%)
Length = 788
Score = 36.6 bits (83), Expect = 0.018
Identities = 28/106 (26%), Positives = 50/106 (46%), Gaps = 4/106 (3%)
Frame = +2
Query: 148 TQVELESK----DKNGDLALSFREYLPDLSEKDIEKKNMAHGEAGWLMEKFDVADYDHNG 203
T+ EL+ D +G+ + F E+L ++ K + + L E F V D D NG
Sbjct: 38 TEAELQDMINEVDADGNGTIDFPEFLNLMARKMKDTDSEEE-----LKEAFRVFDKDQNG 202
Query: 204 LLNFTELRDFLHPEDSQNKEMLKWMVNDKFNMDDYEHDGKINFNQF 249
++ ELR H + +++ V++ D + DG+IN+ +F
Sbjct: 203 FISAAELR---HVMTNLGEKLTDEEVDEMIREADVDGDGQINYEEF 331
>TC224885 similar to UP|Q93YI3 (Q93YI3) Calcium dependent calmodulin
independent protein kinase (Fragment), partial (92%)
Length = 1484
Score = 35.8 bits (81), Expect = 0.031
Identities = 26/136 (19%), Positives = 65/136 (47%)
Frame = +2
Query: 114 LFPLLDRDPKDGFVGFNELESWVTQRALERLDYATQVELESKDKNGDLALSFREYLPDLS 173
+F +D D G + F EL++ + + + + + +E+ D +G+ + + E++
Sbjct: 752 MFKSMDTD-NSGTITFEELKAGLPKLGTKLSESEVRQLMEAADVDGNGTIDYIEFITATM 928
Query: 174 EKDIEKKNMAHGEAGWLMEKFDVADYDHNGLLNFTELRDFLHPEDSQNKEMLKWMVNDKF 233
+ ++ L + F+ D D +G + EL L + +++ +K ++ +
Sbjct: 929 HMNRMERE------DHLYKAFEYFDKDKSGYITMEELESALKKYNMGDEKTIKEIIAEV- 1087
Query: 234 NMDDYEHDGKINFNQF 249
D ++DG+IN+++F
Sbjct: 1088---DADNDGRINYDEF 1126
>TC224884 similar to UP|Q93YI3 (Q93YI3) Calcium dependent calmodulin
independent protein kinase (Fragment), partial (82%)
Length = 1302
Score = 35.4 bits (80), Expect = 0.040
Identities = 26/136 (19%), Positives = 65/136 (47%)
Frame = +1
Query: 114 LFPLLDRDPKDGFVGFNELESWVTQRALERLDYATQVELESKDKNGDLALSFREYLPDLS 173
+F +D D G + F EL++ + + + + + +E+ D +G+ + + E++
Sbjct: 595 MFKSMDTD-NSGTITFEELKAGLPKLGTKVSESEVRQLMEAADVDGNGTIDYIEFITATM 771
Query: 174 EKDIEKKNMAHGEAGWLMEKFDVADYDHNGLLNFTELRDFLHPEDSQNKEMLKWMVNDKF 233
+ ++ L + F+ D D +G + EL L + +++ +K ++ +
Sbjct: 772 HMNRMERE------DHLYKAFEYFDKDRSGYITMEELESTLKKYNMGDEKTIKEIIVEV- 930
Query: 234 NMDDYEHDGKINFNQF 249
D ++DG+IN+++F
Sbjct: 931 ---DTDNDGRINYDEF 969
>NP005117 51 kDa seed maturation protein
Length = 1422
Score = 34.3 bits (77), Expect = 0.090
Identities = 16/46 (34%), Positives = 25/46 (53%)
Frame = +1
Query: 130 NELESWVTQRALERLDYATQVELESKDKNGDLALSFREYLPDLSEK 175
N++ + T A + DYAT +SKD GD A ++Y D ++K
Sbjct: 274 NKVSDYATDTAQKSKDYATDTAQKSKDYAGDAAQKSKDYAGDAAQK 411
>CA801942
Length = 446
Score = 33.9 bits (76), Expect = 0.12
Identities = 30/127 (23%), Positives = 52/127 (40%), Gaps = 2/127 (1%)
Frame = +2
Query: 124 DGFVGFNELESWVTQRALERLDYATQVELESKDKNGDLALSFREYLPDLSEKDIEKKNMA 183
DG + EL+ ++ + D + +E D+NGD + +E+ K +
Sbjct: 95 DGKISMAELKDMLSALGSKTTDEELKRMIEELDQNGDGFIDLKEFADFHCNGGAGKDDSK 274
Query: 184 HGEAGWLMEKFDVADYDHNGLLNFTELRDFLH--PEDSQNKEMLKWMVNDKFNMDDYEHD 241
L + FD+ D D NGL++ EL L E + + + N D + D
Sbjct: 275 E-----LRDAFDLYDVDKNGLISAKELHHVLRNLGEKCSLSDCRRMISN-----VDGDGD 424
Query: 242 GKINFNQ 248
G +NF +
Sbjct: 425 GNVNFEE 445
>TC226141 similar to UP|Q93YA8 (Q93YA8) Calcium binding protein, partial
(57%)
Length = 1062
Score = 32.7 bits (73), Expect = 0.26
Identities = 23/75 (30%), Positives = 38/75 (50%), Gaps = 3/75 (4%)
Frame = +2
Query: 179 KKNMAHGEAGWLMEKFDVADYDHNGLLNFTELRDFLHPEDSQNKEMLKWMVNDKFNM--- 235
+ + A G L + F++ D+D NG ++ TEL L N+ +K V + NM
Sbjct: 590 RSDTADGGDAELHDAFNLYDHDKNGHISATELCQVL------NRLGMKCSVEECHNMIKS 751
Query: 236 DDYEHDGKINFNQFE 250
D + DG +NF +F+
Sbjct: 752 VDSDGDGNVNFPEFK 796
>TC226142 similar to UP|Q93YA8 (Q93YA8) Calcium binding protein, partial
(89%)
Length = 572
Score = 32.7 bits (73), Expect = 0.26
Identities = 23/75 (30%), Positives = 38/75 (50%), Gaps = 3/75 (4%)
Frame = +3
Query: 179 KKNMAHGEAGWLMEKFDVADYDHNGLLNFTELRDFLHPEDSQNKEMLKWMVNDKFNM--- 235
+ + A G L + F++ D+D NG ++ TEL L N+ +K V + NM
Sbjct: 345 RSDTADGGDAELHDAFNLYDHDKNGHISATELCQVL------NRLGMKCSVEECHNMIKS 506
Query: 236 DDYEHDGKINFNQFE 250
D + DG +NF +F+
Sbjct: 507 VDSDGDGNVNFPEFK 551
>TC216101 similar to UP|Q9SCA1 (Q9SCA1) Calcium-binding protein, partial
(65%)
Length = 796
Score = 31.2 bits (69), Expect = 0.76
Identities = 31/110 (28%), Positives = 51/110 (46%), Gaps = 5/110 (4%)
Frame = +1
Query: 190 LMEKFDVADYDHNGLLNFTELRDFLHPEDSQNKEMLKWMVNDK-----FNMDDYEHDGKI 244
L F + D + +G ++ EL D L E L ++ DK D DG +
Sbjct: 235 LKRVFQMFDRNGDGRISLKELSDSL--------ENLGILIPDKDLAQMIERIDVNGDGCV 390
Query: 245 NFNQFEDNVYVTYESYVDFETNGEGDIPTAKDKFAELDVNKDQFLSPEEL 294
+ ++F D YES ++ E + E D+ ++ F D N+D F++ EEL
Sbjct: 391 DMDEFGD----LYESIME-ERDEEEDM---REAFNVFDQNRDGFITVEEL 516
>BQ611191 similar to GP|4589960|gb|A expressed protein {Arabidopsis
thaliana}, partial (30%)
Length = 412
Score = 30.8 bits (68), Expect = 1.00
Identities = 21/81 (25%), Positives = 36/81 (43%)
Frame = -3
Query: 42 ILERAPTFDPLVTKIERESEQKNQQHKNDFDNNKNVAPRTGLGSTTTVSEIKETYEYLTS 101
IL++ P + ++ +QKN K NNK + P G G ++ E+
Sbjct: 308 ILQKLPPKSNI*KTN*KKKQQKNLFPKQLIRNNKKLLPLLGFGDLLQRIRVRNLLEF--- 138
Query: 102 GGTLNTTLRLIILFPLLDRDP 122
G+ + + L++L PLL P
Sbjct: 137 -GSRSDNILLLLLLPLLSLLP 78
>TC209701 similar to UP|Q8LDS1 (Q8LDS1) Calcium-dependent protein kinase,
partial (30%)
Length = 799
Score = 30.8 bits (68), Expect = 1.00
Identities = 35/141 (24%), Positives = 62/141 (43%), Gaps = 6/141 (4%)
Frame = +3
Query: 115 FPLLDRDPKDGFVGFNELESWVTQRALERLDYATQVE-LESKDKNGDLALSFREYLPDLS 173
F +D D K+G + E+ + + +L + +E L++ D N D + FRE++
Sbjct: 33 FDAIDVD-KNGSISLEEMRQALAKDLPWKLKESRVLEILQAIDNNTDGLVDFREFVAATL 209
Query: 174 -----EKDIEKKNMAHGEAGWLMEKFDVADYDHNGLLNFTELRDFLHPEDSQNKEMLKWM 228
E+D +K A EKFD+ D +G ++ PE+ + L+
Sbjct: 210 HVHQLEEDSDKWQQLSQAA---FEKFDL---DKDG---------YITPEELRMHTCLRGS 344
Query: 229 VNDKFNMDDYEHDGKINFNQF 249
V+ D + DGKI+ +F
Sbjct: 345 VDPLLEEADIDKDGKISLPEF 407
Database: GMGI
Posted date: Oct 22, 2004 4:58 PM
Number of letters in database: 37,918,896
Number of sequences in database: 63,676
Lambda K H
0.315 0.136 0.392
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 12,563,442
Number of Sequences: 63676
Number of extensions: 142516
Number of successful extensions: 554
Number of sequences better than 10.0: 52
Number of HSP's better than 10.0 without gapping: 531
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 553
length of query: 360
length of database: 12,639,632
effective HSP length: 98
effective length of query: 262
effective length of database: 6,399,384
effective search space: 1676638608
effective search space used: 1676638608
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 16 ( 7.3 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 42 (22.0 bits)
S2: 59 (27.3 bits)
Medicago: description of AC149127.7


