

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC149127.6 - phase: 0
(421 letters)
Database: GMGI
63,676 sequences; 37,918,896 total letters
Searching..................................................done
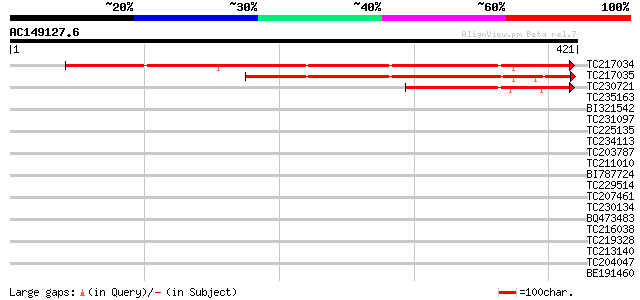
Score E
Sequences producing significant alignments: (bits) Value
TC217034 similar to UP|Q8L732 (Q8L732) Expressed protein, partia... 395 e-110
TC217035 similar to UP|Q8L732 (Q8L732) Expressed protein, partia... 266 1e-71
TC230721 similar to UP|Q8L732 (Q8L732) Expressed protein, partia... 150 1e-36
TC235163 similar to UP|Q9XYL1 (Q9XYL1) Development protein DG110... 34 0.11
BI321542 similar to GP|15626070|gb vitellogenin {Larus argentatu... 33 0.24
TC231097 similar to UP|Q8L732 (Q8L732) Expressed protein, partia... 33 0.24
TC225135 similar to UP|Q9M549 (Q9M549) Poly(A)-binding protein, ... 32 0.41
TC234113 weakly similar to UP|Q7YZI0 (Q7YZI0) MBCTL1 (Fragment),... 32 0.54
TC203787 similar to UP|Q6VBJ3 (Q6VBJ3) Epa4p, partial (6%) 30 1.6
TC211010 weakly similar to UP|Q8L443 (Q8L443) At1g22470/F12K8_18... 30 1.6
BI787724 similar to PIR|T39903|T399 serine-rich protein - fissio... 30 1.6
TC229514 similar to UP|ATE1_ARATH (Q9ZT48) Arginyl-tRNA--protein... 29 3.5
TC207461 similar to GB|AAK96599.1|15450655|AY052695 AT3g52740/F3... 29 3.5
TC230134 weakly similar to UP|NUCL_HUMAN (P19338) Nucleolin (Pro... 29 4.5
BQ473483 29 4.5
TC216038 similar to UP|Q9IZF4 (Q9IZF4) Aphid transmission P5, pa... 29 4.5
TC219328 similar to UP|Q7EZ44 (Q7EZ44) Receptor-like kinase Xa21... 28 5.9
TC213140 weakly similar to UP|Q762A2 (Q762A2) BRI1-KD interactin... 28 7.7
TC204047 hydroxyproline-rich glycoprotein 28 7.7
BE191460 weakly similar to GP|9798599|dbj serine/threonine prote... 28 7.7
>TC217034 similar to UP|Q8L732 (Q8L732) Expressed protein, partial (24%)
Length = 1633
Score = 395 bits (1014), Expect = e-110
Identities = 220/397 (55%), Positives = 269/397 (67%), Gaps = 19/397 (4%)
Frame = +2
Query: 42 DMKWWLHVKTNLAGDTDYTCQHLNTLESKLDAFSTRLHGDNVSIGSDQTVKNFDAVSFVG 101
+MKWWLHVK++L + +YTCQ LN+ ES+L F + +V G DQ VK+FDA+S +
Sbjct: 107 EMKWWLHVKSDLGDEANYTCQQLNSYESELGTFYSEFLNGSVKSGGDQLVKDFDALSNI- 283
Query: 102 NAATAAIEQQWNVY-PKYMKNSDTRTSKIEASLNSDLYLTPKKKNEGEFWFSD------D 154
+A +IEQ W+V P MKN++TR K EASLN+DL+ TPKKK++ EF+F D D
Sbjct: 284 QSANLSIEQPWHVVSPTCMKNNNTRMPKFEASLNTDLHFTPKKKDQEEFYFQDSHFMDCD 463
Query: 155 ATSFLISEHCKSTSSDFEPHWLGAEKSQPWWRTTGKDDLASLVAKKSFEYVENCDLPEPN 214
++FL+SE K SSD E H +GAEK+ WWRTTGKD+LASLVA+KS E++ NCDLP P
Sbjct: 464 ISNFLVSEQGKMASSDLESHLMGAEKTGAWWRTTGKDELASLVARKSLEHIGNCDLPHPQ 643
Query: 215 IKPFRKIHTLQPRETDKEENQVSSLNQKLEMCSSDSNGCTSTTLTSGCSFQDSDRTFSSS 274
K P+ D + SLN+K E SS ++G T+ TS CSFQDS + SSS
Sbjct: 644 TKHVSP-RPPYPKGVDHHDKTPPSLNRKTETGSSYADGYTTGISTSDCSFQDSSKHSSSS 820
Query: 275 ESKDSDSSCNKNSKVNSESAAKAELLKALCRSQTRAREAEKAAQEACDEKEHILSLFFKQ 334
SKDS SS + ++NSE+ + +ELLKALC SQTRAREAEKAAQ A EKEHILSLFF+Q
Sbjct: 821 HSKDSSSS-TEGCQINSENGSMSELLKALCHSQTRAREAEKAAQRAYSEKEHILSLFFRQ 997
Query: 335 ASQLFAYKQWLHVLQLENLCLQYKSKNQPLLNNLFPYE-----------GKKHRKNR-RK 382
ASQLFAYKQWLH+LQLENLCLQ ++KNQ NLFP K H K R RK
Sbjct: 998 ASQLFAYKQWLHMLQLENLCLQLRNKNQS-FPNLFPASLPWDPCRGRLLKKSHSKARKRK 1174
Query: 383 VKNSRRGIGKCIFAFAVGLALAGAGLLLGWTIGCMFP 419
N R I KC AFAVGL LAGAGLLLGWT+G MFP
Sbjct: 1175NSNRRCVITKCAVAFAVGLGLAGAGLLLGWTMGWMFP 1285
>TC217035 similar to UP|Q8L732 (Q8L732) Expressed protein, partial (22%)
Length = 1128
Score = 266 bits (680), Expect = 1e-71
Identities = 150/257 (58%), Positives = 181/257 (70%), Gaps = 12/257 (4%)
Frame = +3
Query: 176 LGAEKSQPWWRTTGKDDLASLVAKKSFEYVENCDLPEPNIKPFRKIHTLQPRETDKEENQ 235
+GAEK+ WWRTTGKD+LASLVA+KS E++ENCDLP+P K + P+ D +
Sbjct: 6 MGAEKTGAWWRTTGKDELASLVARKSLEHIENCDLPQPQTKHVSQ-RPSYPKGVDHDNTP 182
Query: 236 VSSLNQKLEMCSSDSNGCTSTTLTSGCSFQDSDRTFSSSESKDSDSSCNKNSKVNSESAA 295
SSLN+K E SS ++ T+ TS CSFQDS++ SSS+SKD SS + ++NSE+ +
Sbjct: 183 PSSLNRKTETGSSYADSYTTGISTSDCSFQDSNKHSSSSQSKDFSSS-TEGCQINSENGS 359
Query: 296 KAELLKALCRSQTRAREAEKAAQEACDEKEHILSLFFKQASQLFAYKQWLHVLQLENLCL 355
+ELLKALC SQTRAREAEKAAQ A EKEHILSLFF+QASQLFAYKQWLH+LQLENLCL
Sbjct: 360 VSELLKALCHSQTRAREAEKAAQRAYSEKEHILSLFFRQASQLFAYKQWLHMLQLENLCL 539
Query: 356 QYKSKNQPLLNNLFPYE----------GKKHRKNRRKVKNSRRG--IGKCIFAFAVGLAL 403
Q ++KNQ L NLFP KK RK +NS + I KC+ A AVGL L
Sbjct: 540 QLRNKNQSFL-NLFPASLPWDPCRGMLLKKSNSKARKRRNSNKRCVIAKCVVA-AVGLGL 713
Query: 404 AGAGLLLGWTIGCMFPS 420
AGAGLLLGWT+G MFPS
Sbjct: 714 AGAGLLLGWTMGWMFPS 764
>TC230721 similar to UP|Q8L732 (Q8L732) Expressed protein, partial (23%)
Length = 750
Score = 150 bits (379), Expect = 1e-36
Identities = 80/141 (56%), Positives = 103/141 (72%), Gaps = 16/141 (11%)
Frame = +1
Query: 295 AKAELLKALCRSQTRAREAEKAAQEACDEKEHILSLFFKQASQLFAYKQWLHVLQLENLC 354
+KA+L++ALC SQTRAREAE+AA++A EKE I++L FKQASQLFAYKQWL +LQLE LC
Sbjct: 25 SKAQLMEALCHSQTRAREAEEAAKKAYAEKEDIVTLIFKQASQLFAYKQWLQLLQLETLC 204
Query: 355 LQYKSKNQPLLNNLFP-------YEGKKHRKNRRKVKNSRRGIGKC---------IFAFA 398
+Q KSK+QP ++ LFP YEG+ RK ++K+ NS++G K AFA
Sbjct: 205 IQIKSKDQP-ISTLFPVALPWMSYEGRSSRKRKQKISNSKQGERKANSKCDITTYAVAFA 381
Query: 399 VGLALAGAGLLLGWTIGCMFP 419
+GL+L GAGLLLGWT+G M P
Sbjct: 382 LGLSLVGAGLLLGWTVGWMLP 444
>TC235163 similar to UP|Q9XYL1 (Q9XYL1) Development protein DG1106
(Fragment), partial (5%)
Length = 927
Score = 34.3 bits (77), Expect = 0.11
Identities = 34/121 (28%), Positives = 50/121 (41%), Gaps = 15/121 (12%)
Frame = +3
Query: 233 ENQVSSLNQKLEM---CSSDSNGCTSTTLTSGCSFQDSDRTFSSSESKDSDSSCNKNSKV 289
E +S L++KL CS +G +S + S D + SSS S DSDS C +S
Sbjct: 468 EGDLSGLSKKLSSSCKCSDSDDGFSSL---NSYSRNSDDESTSSSPSSDSDSECCSSSSS 638
Query: 290 NSESAAKAELLKALCRSQTRAREAEKAAQEACDEK------------EHILSLFFKQASQ 337
S S + ++RE E + E+ E + IL+ F K A Q
Sbjct: 639 TSSS------------TPNQSRENEPSIAESSPEYIVNLLRLHIQGIQSILTRFSKSADQ 782
Query: 338 L 338
+
Sbjct: 783 I 785
>BI321542 similar to GP|15626070|gb vitellogenin {Larus argentatus}, partial
(1%)
Length = 358
Score = 33.1 bits (74), Expect = 0.24
Identities = 28/96 (29%), Positives = 47/96 (48%), Gaps = 2/96 (2%)
Frame = +1
Query: 274 SESKDSDSSCNKNSKVNSESAAKAELLKALCRSQTRAREAEKAAQEACDEKEHILSLFFK 333
SES DSDSS K K +S S +++ K + RS+ R + + E D +E S F
Sbjct: 40 SESDDSDSSGRKKKKRSSSSKSRS---KRIRRSRKRTNRSPETEGEGSDLEEENGSGFVD 210
Query: 334 QASQLFAYKQWLHVLQLENLCLQ--YKSKNQPLLNN 367
S+ + + E + L+ ++S+ +P L+N
Sbjct: 211 AKSKTTVDEVMKTEINAEAIKLKELFESQKKPALDN 318
>TC231097 similar to UP|Q8L732 (Q8L732) Expressed protein, partial (9%)
Length = 523
Score = 33.1 bits (74), Expect = 0.24
Identities = 20/80 (25%), Positives = 32/80 (40%), Gaps = 11/80 (13%)
Frame = +3
Query: 4 AARARCAVNHCFTQDFMMPPTSSKQLEPDFTTSDST-----------DSDMKWWLHVKTN 52
A + C + C T + +S E D T + T D +WWLH++ N
Sbjct: 213 APKLACCQSSCATSKSVDTGPASTADESDHNTVNVTHFNRKSSISNLSPDSRWWLHLQPN 392
Query: 53 LAGDTDYTCQHLNTLESKLD 72
T + LN LE +++
Sbjct: 393 YGYQKGLTYEQLNALEDEVE 452
>TC225135 similar to UP|Q9M549 (Q9M549) Poly(A)-binding protein, partial (97%)
Length = 2392
Score = 32.3 bits (72), Expect = 0.41
Identities = 20/44 (45%), Positives = 24/44 (54%)
Frame = -1
Query: 205 VENCDLPEPNIKPFRKIHTLQPRETDKEENQVSSLNQKLEMCSS 248
VEN P+P +KPF + TLQ K EN SSL+ M SS
Sbjct: 1174 VENATKPDPRLKPFGSLITLQEVMVPKGEN--SSLSFSSPMLSS 1049
>TC234113 weakly similar to UP|Q7YZI0 (Q7YZI0) MBCTL1 (Fragment), partial
(5%)
Length = 460
Score = 32.0 bits (71), Expect = 0.54
Identities = 18/53 (33%), Positives = 28/53 (51%), Gaps = 1/53 (1%)
Frame = +1
Query: 246 CSSDSNGCTSTTLT-SGCSFQDSDRTFSSSESKDSDSSCNKNSKVNSESAAKA 297
CSS + +S L+ S CS S R SS+ S + SSC+ S + S+ ++
Sbjct: 295 CSSSCSSASSGNLSPSSCSSPSSPRGSSSTSSATAPSSCSATSSTTASSSRRS 453
>TC203787 similar to UP|Q6VBJ3 (Q6VBJ3) Epa4p, partial (6%)
Length = 1056
Score = 30.4 bits (67), Expect = 1.6
Identities = 16/42 (38%), Positives = 23/42 (54%)
Frame = +3
Query: 252 GCTSTTLTSGCSFQDSDRTFSSSESKDSDSSCNKNSKVNSES 293
GC +++ +S SF S SSS S S SS + +S +S S
Sbjct: 294 GCNASSSSSSSSFSSSSAPSSSSSSPASSSSSSSSSSSSSSS 419
>TC211010 weakly similar to UP|Q8L443 (Q8L443) At1g22470/F12K8_18, partial
(36%)
Length = 492
Score = 30.4 bits (67), Expect = 1.6
Identities = 30/103 (29%), Positives = 39/103 (37%), Gaps = 16/103 (15%)
Frame = +2
Query: 239 LNQKLEMCSSDSNGCTSTTLTSGCSFQDSDRTFSSSESKDSDSSC---------NKNSKV 289
L+Q CSS S+ TT S C S + +S S S SSC +K+SK
Sbjct: 173 LSQLSSACSSSSSSSRKTTFISSCHSSSSSSSLTSYCS-SSASSCASPTLPFPFHKDSKS 349
Query: 290 NSE-------SAAKAELLKALCRSQTRAREAEKAAQEACDEKE 325
S SA +LK+ S R K E D +
Sbjct: 350 GSNSVSIFLLSAKPGGILKSRSMSVVATRRRTKGQDEGDDRNK 478
>BI787724 similar to PIR|T39903|T399 serine-rich protein - fission yeast
(Schizosaccharomyces pombe), partial (5%)
Length = 283
Score = 30.4 bits (67), Expect = 1.6
Identities = 20/62 (32%), Positives = 31/62 (49%)
Frame = -3
Query: 248 SDSNGCTSTTLTSGCSFQDSDRTFSSSESKDSDSSCNKNSKVNSESAAKAELLKALCRSQ 307
S S C+S+++ S SSS S S+SS + +S + S+ K +LL C S
Sbjct: 269 SSSTSCSSSSMLSSIRM*SVSDPLSSSSSPTSNSSVSSSSS-SFSSSFKTKLLPFSCPSS 93
Query: 308 TR 309
T+
Sbjct: 92 TK 87
>TC229514 similar to UP|ATE1_ARATH (Q9ZT48) Arginyl-tRNA--protein transferase
1 (R-transferase 1) (Arginyltransferase 1)
(Arginine-tRNA--protein transferase 1) , partial (27%)
Length = 1035
Score = 29.3 bits (64), Expect = 3.5
Identities = 15/40 (37%), Positives = 24/40 (59%)
Frame = -1
Query: 256 TTLTSGCSFQDSDRTFSSSESKDSDSSCNKNSKVNSESAA 295
T+ SG S + R+ SSSE DSDS+ + +S + + A+
Sbjct: 684 TSAISGSSSEVVSRSGSSSEHSDSDSTISSSSSIGTSFAS 565
>TC207461 similar to GB|AAK96599.1|15450655|AY052695 AT3g52740/F3C22_140
{Arabidopsis thaliana;} , partial (48%)
Length = 1713
Score = 29.3 bits (64), Expect = 3.5
Identities = 18/65 (27%), Positives = 36/65 (54%)
Frame = -3
Query: 239 LNQKLEMCSSDSNGCTSTTLTSGCSFQDSDRTFSSSESKDSDSSCNKNSKVNSESAAKAE 298
L+ + M S +N CT+ + + C+F D+++ S+++K SD+S K + E+ +
Sbjct: 1042 LDFNVHMDLSLNNNCTNVEVQNTCTFSDTEK---STKTKGSDAS----PKTHEENVQVIQ 884
Query: 299 LLKAL 303
L+ L
Sbjct: 883 LVSKL 869
>TC230134 weakly similar to UP|NUCL_HUMAN (P19338) Nucleolin (Protein C23),
partial (8%)
Length = 581
Score = 28.9 bits (63), Expect = 4.5
Identities = 15/42 (35%), Positives = 22/42 (51%)
Frame = -3
Query: 258 LTSGCSFQDSDRTFSSSESKDSDSSCNKNSKVNSESAAKAEL 299
L GC S + SSS S S SS + +S +S S++ + L
Sbjct: 546 LEGGCGVSSSSSSSSSSSSSSSPSSSSLSSSSSSSSSSSSSL 421
>BQ473483
Length = 421
Score = 28.9 bits (63), Expect = 4.5
Identities = 19/69 (27%), Positives = 34/69 (48%), Gaps = 5/69 (7%)
Frame = -3
Query: 233 ENQVSSLNQKLEMCSSDSNG--CTSTTLTSGCSFQ---DSDRTFSSSESKDSDSSCNKNS 287
+ +SS + C+S C+S++ +S C+ Q + R+ S+ S D S +K
Sbjct: 347 KTHLSSFHSSSHPCASKQTPATCSSSSSSSACTRQIPPRAKRSSGSAASGDRRSRVSKKK 168
Query: 288 KVNSESAAK 296
K++ E AK
Sbjct: 167 KIDEEKQAK 141
>TC216038 similar to UP|Q9IZF4 (Q9IZF4) Aphid transmission P5, partial (3%)
Length = 1754
Score = 28.9 bits (63), Expect = 4.5
Identities = 20/55 (36%), Positives = 28/55 (50%), Gaps = 4/55 (7%)
Frame = -2
Query: 243 LEMCSSDS----NGCTSTTLTSGCSFQDSDRTFSSSESKDSDSSCNKNSKVNSES 293
LE C S S G +++ LTSG F+DS + S S S S S +S ++ S
Sbjct: 943 LESCLSVSFGGGKGTSASFLTSGSGFKDSSGS*SCSSSTSSSPSSKISSSLSLSS 779
>TC219328 similar to UP|Q7EZ44 (Q7EZ44) Receptor-like kinase Xa21-binding
protein 3-like, partial (51%)
Length = 1424
Score = 28.5 bits (62), Expect = 5.9
Identities = 26/77 (33%), Positives = 35/77 (44%), Gaps = 1/77 (1%)
Frame = +3
Query: 208 CDLPEPNIKPFRKIHTLQPRETDKEENQVSSLNQKLEMCSSDSNGCTSTTLTSGCSFQDS 267
C L I F K+ +P KE + S+L+ L CSS+ G +S T C S
Sbjct: 837 CPLCRHGIVSFVKLPDARPLH--KEMQRTSNLSLTLCTCSSEVLGDSSDMTTPFCKPTSS 1010
Query: 268 DRTFSSSESKDSDS-SC 283
+ SSS S+ S SC
Sbjct: 1011RGSKSSSPSRSFRSMSC 1061
>TC213140 weakly similar to UP|Q762A2 (Q762A2) BRI1-KD interacting protein
112 (Fragment), partial (30%)
Length = 795
Score = 28.1 bits (61), Expect = 7.7
Identities = 19/61 (31%), Positives = 29/61 (47%), Gaps = 1/61 (1%)
Frame = +2
Query: 267 SDRTFSSSESKDSDSSCNKNSK-VNSESAAKAELLKALCRSQTRAREAEKAAQEACDEKE 325
+D T S + K + N K N + K+ LKAL + Q + + +EKA E E+
Sbjct: 23 TDSTSKSKKQKSVKENLNSKRKDANKKQTDKS--LKALLKDQGKDKSSEKAKAEPSKEEL 196
Query: 326 H 326
H
Sbjct: 197 H 199
>TC204047 hydroxyproline-rich glycoprotein
Length = 887
Score = 28.1 bits (61), Expect = 7.7
Identities = 15/47 (31%), Positives = 21/47 (43%)
Frame = +3
Query: 353 LCLQYKSKNQPLLNNLFPYEGKKHRKNRRKVKNSRRGIGKCIFAFAV 399
L LQ PLL + F H RK +N G+G+CI + +
Sbjct: 459 LLLQITPSTLPLLGSTF--SSHPHMDAYRKCENESHGVGRCICSLQI 593
>BE191460 weakly similar to GP|9798599|dbj serine/threonine protein kinase
{Arabidopsis thaliana}, partial (26%)
Length = 523
Score = 28.1 bits (61), Expect = 7.7
Identities = 17/52 (32%), Positives = 21/52 (39%)
Frame = +3
Query: 132 SLNSDLYLTPKKKNEGEFWFSDDATSFLISEHCKSTSSDFEPHWLGAEKSQP 183
SL S Y PK+K+ W S A S + H S PH+ S P
Sbjct: 195 SLTSSKYSRPKQKSTSS-WSSPPAGSSFTTSHATCCSPMKPPHFTSCNSSPP 347
Database: GMGI
Posted date: Oct 22, 2004 4:58 PM
Number of letters in database: 37,918,896
Number of sequences in database: 63,676
Lambda K H
0.314 0.127 0.382
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 21,018,118
Number of Sequences: 63676
Number of extensions: 299316
Number of successful extensions: 1890
Number of sequences better than 10.0: 42
Number of HSP's better than 10.0 without gapping: 1863
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 1878
length of query: 421
length of database: 12,639,632
effective HSP length: 100
effective length of query: 321
effective length of database: 6,272,032
effective search space: 2013322272
effective search space used: 2013322272
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 16 ( 7.3 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.6 bits)
S2: 60 (27.7 bits)
Medicago: description of AC149127.6


