

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC149039.3 + phase: 0 /pseudo
(2185 letters)
Database: GMGI
63,676 sequences; 37,918,896 total letters
Searching..................................................done
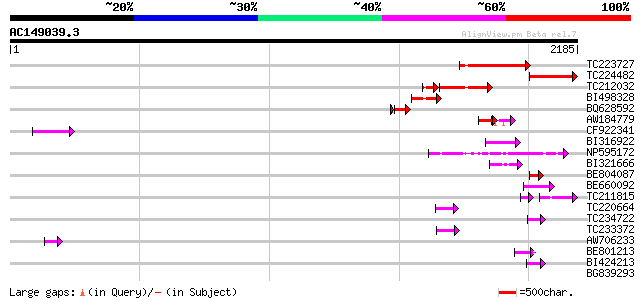
Score E
Sequences producing significant alignments: (bits) Value
TC223727 similar to UP|Q6WAY3 (Q6WAY3) Gag/pol polyprotein, part... 363 e-100
TC224482 similar to UP|Q6WAY3 (Q6WAY3) Gag/pol polyprotein, part... 272 1e-72
TC212032 similar to UP|Q6WAY3 (Q6WAY3) Gag/pol polyprotein, part... 212 4e-69
BI498328 123 1e-27
BQ628592 103 1e-23
AW184779 108 2e-23
CF922341 96 1e-19
BI316922 95 4e-19
NP595172 polyprotein [Glycine max] 91 7e-18
BI321666 83 1e-15
BE804087 81 4e-15
BE660092 weakly similar to GP|9884624|dbj retroelement pol polyp... 78 4e-14
TC211815 48 4e-08
TC220664 weakly similar to UP|Q84TE0 (Q84TE0) At5g51080, partial... 58 4e-08
TC234722 similar to UP|Q6WAY9 (Q6WAY9) Pol (Fragment), partial (... 58 5e-08
TC233372 weakly similar to UP|Q84TE0 (Q84TE0) At5g51080, partial... 54 7e-07
AW706233 54 7e-07
BE801213 weakly similar to GP|6691193|gb| F7F22.17 {Arabidopsis ... 48 5e-05
BI424213 46 2e-04
BG839293 39 0.019
>TC223727 similar to UP|Q6WAY3 (Q6WAY3) Gag/pol polyprotein, partial (9%)
Length = 843
Score = 363 bits (932), Expect = e-100
Identities = 167/272 (61%), Positives = 206/272 (75%)
Frame = +1
Query: 1733 NHWNDVPIIKVQRLERPSHVFAIGDVIDQAGENVVDYKPWYYDIKQFLLSREYPPGASKQ 1792
N D+P I+ +P+H + E D KPWY+DIK++++S+EY P +
Sbjct: 46 NAARDLPYIEFWCRGKPAHCCQV--------EEERDGKPWYFDIKRYVISKEYLPEIADN 201
Query: 1793 DKKTLRRLTSRFLLDGDILYKRNYDMVLLRCVDEHEAEQLMHDVHDGTFRTHATGHTMSR 1852
DK+TLRRL + F + G ILYKRN+DM LRCVD EA ++ +VH+G+F THA GH M+R
Sbjct: 202 DKRTLRRLAAGFFMSGSILYKRNHDMKPLRCVDAREANHMIEEVHEGSFGTHANGHAMAR 381
Query: 1853 KLLRAGYYWMAMEHDCYQHARKCHKCQIYADKIHVPPHALNVISSPWPFSMWGIDMIGRI 1912
K+LRAGYYW+ ME DC H RKCHKCQ +AD ++ PPH LNV+SSPWPFSMWGID+IG I
Sbjct: 382 KILRAGYYWLTMESDCCVHVRKCHKCQAFADNVNAPPHPLNVMSSPWPFSMWGIDVIGAI 561
Query: 1913 EPKASNGHHFILVAIDYFTKWVEAASYTNVTKQVVAKFIKNNIICRYGVPSKIITDNGTN 1972
EPKASNGH FILVAIDYFTKWVEAASYT+V + VV +FIK IICRYG+P KIITDNGTN
Sbjct: 562 EPKASNGHRFILVAIDYFTKWVEAASYTDVMRGVVVRFIKKEIICRYGLPRKIITDNGTN 741
Query: 1973 LNNNVVQALCEEFKIEHHNSSPYRPQMNGAVE 2004
LNN ++ +CEEFKI+HHN +PYRP+MN AVE
Sbjct: 742 LNNKMMGEICEEFKIQHHNPTPYRPKMN*AVE 837
>TC224482 similar to UP|Q6WAY3 (Q6WAY3) Gag/pol polyprotein, partial (6%)
Length = 669
Score = 272 bits (695), Expect = 1e-72
Identities = 126/182 (69%), Positives = 156/182 (85%)
Frame = +1
Query: 2004 EATNKNIKRIVQKMVTTYKDWHEMLPYALHGYRTTVRSSTGATPFSLVYGMEAVLPLEVE 2063
EA NKNIK+I+QKM +YKDWHEMLP+ALHGYRT+VR+STGATPFSLVYGMEAVLP EVE
Sbjct: 1 EAANKNIKKIIQKMTVSYKDWHEMLPFALHGYRTSVRTSTGATPFSLVYGMEAVLPFEVE 180
Query: 2064 IPSLRVIMEAKLSEAEWCQSRYDQLNLIEEKRMDAVARGQSYQARMKTAFDKKVHPREFK 2123
+PSLR++ E+ L E+EW Q+RYDQLNLIE KR+ A++ G+ YQ RMK+AFDKKV R+F
Sbjct: 181 VPSLRILAESGLKESEWAQTRYDQLNLIEGKRLTAMSHGRLYQQRMKSAFDKKVCLRKFH 360
Query: 2124 VGELVLKRRISQQPDPRGKWTPNYEGPYVVKKAFSGGALILTHMDGVELPNPVNADIVKK 2183
G+LVLK+ D RGKW PNYEGP+VVK+AFSGGAL+LT+MDG ELP+P+N+D+VK+
Sbjct: 361 EGDLVLKKMSHAVKDHRGKWAPNYEGPFVVKRAFSGGALVLTNMDGEELPSPMNSDVVKR 540
Query: 2184 YF 2185
Y+
Sbjct: 541 YY 546
>TC212032 similar to UP|Q6WAY3 (Q6WAY3) Gag/pol polyprotein, partial (3%)
Length = 803
Score = 212 bits (540), Expect(2) = 4e-69
Identities = 102/207 (49%), Positives = 141/207 (67%)
Frame = +2
Query: 1655 IEEAIDMRIKHLDIYGDSALVINQIKGEWETHHAKLIPYRDYARRLLTYFTKVELHHIPR 1714
++ AID +K L +YGDSALVI+Q++GE ET LIPY+ Y + L +F ++ HH+
Sbjct: 206 VQAAIDSNVKLLKVYGDSALVIHQLRGECETRDPNLIPYQAYIKELAGFFDEISFHHVA* 385
Query: 1715 DENQMADALATLSSMFRVNHWNDVPIIKVQRLERPSHVFAIGDVIDQAGENVVDYKPWYY 1774
+ENQMADALATL SMF++ D+P I+ + RP+H + E D KPWY+
Sbjct: 386 EENQMADALATLVSMFQLTPHGDLPYIEFRCRGRPAHCCLV--------EEERDGKPWYF 541
Query: 1775 DIKQFLLSREYPPGASKQDKKTLRRLTSRFLLDGDILYKRNYDMVLLRCVDEHEAEQLMH 1834
DIK+++ S+EYP AS DK+ RRL + F + G ILYKRN+DMVLL CV+ E E ++
Sbjct: 542 DIKRYVESKEYPLEASDNDKRRKRRLAAGFFMSGSILYKRNHDMVLLHCVNGKEVENMLG 721
Query: 1835 DVHDGTFRTHATGHTMSRKLLRAGYYW 1861
+VH+G+F TH+ GH M+RK+LRAGYYW
Sbjct: 722 EVHEGSFGTHSNGHAMARKILRAGYYW 802
Score = 70.1 bits (170), Expect(2) = 4e-69
Identities = 34/61 (55%), Positives = 41/61 (66%)
Frame = +1
Query: 1591 EPLIDEGPDPNSKWGLVFDGAVNAYGKGIGAVIVSPQGHHIPFTARILFECTNNMAEYEA 1650
E +DE D KW + FD A N G G+GA++VSP IPFT R+ F+CTNNMAEYEA
Sbjct: 22 EEKLDEDRD---KWIVWFDRASNVLGHGVGAILVSPDNQCIPFTTRLGFDCTNNMAEYEA 192
Query: 1651 C 1651
C
Sbjct: 193 C 195
>BI498328
Length = 335
Score = 123 bits (308), Expect = 1e-27
Identities = 64/118 (54%), Positives = 81/118 (68%), Gaps = 1/118 (0%)
Frame = +1
Query: 1548 TQKAIKGSILADHLAYQPLDDYQPIEFDFPDEEIMYL-KSKDCEEPLIDEGPDPNSKWGL 1606
TQKA+KGS LAD+LA PL Y+P+ +FPDE+IM L + K E + +KW +
Sbjct: 4 TQKAVKGSALADYLAQ*PLQGYRPMHPEFPDEDIMALFEEKRTHEDI--------NKWIV 159
Query: 1607 VFDGAVNAYGKGIGAVIVSPQGHHIPFTARILFECTNNMAEYEACIFGIEEAIDMRIK 1664
FDGA NA G G+GAV+VSP IPFTAR+ F+CTNNMAEYEAC G++ AID +K
Sbjct: 160 CFDGASNALGHGVGAVLVSPDDQCIPFTARLGFDCTNNMAEYEACALGVQAAIDFDVK 333
>BQ628592
Length = 423
Score = 103 bits (258), Expect(2) = 1e-23
Identities = 43/61 (70%), Positives = 55/61 (89%)
Frame = -3
Query: 1484 YTMLEKTCCALAWAAKRLRHYLVNHTTWLISRMDPIKYIFEKAVVTGKIARWQVLLSEYD 1543
Y+MLE+TCC L WA+ RLR Y+++HTTWLIS+MDP+KYIFEK +TG+IARWQVLLSE++
Sbjct: 184 YSMLERTCCTLVWASHRLRQYMLSHTTWLISKMDPVKYIFEKPALTGRIARWQVLLSEFN 5
Query: 1544 I 1544
I
Sbjct: 4 I 2
Score = 26.6 bits (57), Expect(2) = 1e-23
Identities = 11/11 (100%), Positives = 11/11 (100%)
Frame = -1
Query: 1469 AIYYLSKKFTD 1479
AIYYLSKKFTD
Sbjct: 228 AIYYLSKKFTD 196
>AW184779
Length = 432
Score = 108 bits (271), Expect = 2e-23
Identities = 47/74 (63%), Positives = 58/74 (77%)
Frame = +1
Query: 1806 LDGDILYKRNYDMVLLRCVDEHEAEQLMHDVHDGTFRTHATGHTMSRKLLRAGYYWMAME 1865
L +ILYKRN+DMVLLRCVD EAEQ++ +VH+G+F THA H M++K+LR GYYW+ ME
Sbjct: 1 LSRNILYKRNHDMVLLRCVDAREAEQMLVEVHEGSFGTHANIHAMAQKILRVGYYWLTME 180
Query: 1866 HDCYQHARKCHKCQ 1879
DC H KCHKCQ
Sbjct: 181 SDCCIHVWKCHKCQ 222
Score = 86.3 bits (212), Expect = 1e-16
Identities = 49/114 (42%), Positives = 60/114 (51%), Gaps = 22/114 (19%)
Frame = +3
Query: 1858 GYYWMAMEHDCY-------------QHARKCHKCQIYADKIHVPPHALNVISSPWPF--- 1901
G W +H C+ R H C + VP ++ +P P+
Sbjct: 99 GILWNTCQHTCHGPEDSESGVLLAHYGERLLHPC---VEMP*VPVRPSLIMLTPHPYL*M 269
Query: 1902 ------SMWGIDMIGRIEPKASNGHHFILVAIDYFTKWVEAASYTNVTKQVVAK 1949
MWGID+IG IEPKASNGHHFILVAIDYFTKWVEA SY +VT+ VV +
Sbjct: 270 SWQHLGHMWGIDVIGAIEPKASNGHHFILVAIDYFTKWVEAVSYASVTRSVVIR 431
>CF922341
Length = 675
Score = 96.3 bits (238), Expect = 1e-19
Identities = 63/170 (37%), Positives = 86/170 (50%), Gaps = 9/170 (5%)
Frame = +1
Query: 87 NLTPVSTALTQAATTVTE---PIVNTV--PQFVHANAHRGSIATTGTMEERMEELAK--E 139
NL T LT A P+ NT+ PQ+ H H +TT M E+ K
Sbjct: 142 NLADFETCLTYATEGQAVGGIPLRNTLEGPQY-HPQLHLLH-STTSKNPHVMAEMGKLDH 315
Query: 140 LRREIKANRGNGDSI--KTQDLCLVSKVDVPKKFKVPEFDKYNGLTCPQNHIVKYVRKMS 197
L ++A G D ++L LV + P KFKV +FDKY G TCP+NH+ Y +KM
Sbjct: 316 LEEGLRAIEGGEDYAFANLEELFLVPNIITPPKFKVLDFDKYKGTTCPKNHLKMYCQKMG 495
Query: 198 NYKDNDSLMIHCFQDSLMEDAAEWYTSLSKDDVHTFDELAAAFKSHYGFN 247
Y ++ L+IH FQ+SL A WYT+L VH++ +L AF Y +N
Sbjct: 496 AYAKDEELLIHSFQESLTGVAVTWYTNLEPSRVHSWKDLMVAFVRQYQYN 645
>BI316922
Length = 405
Score = 94.7 bits (234), Expect = 4e-19
Identities = 45/135 (33%), Positives = 78/135 (57%)
Frame = +3
Query: 1835 DVHDGTFRTHATGHTMSRKLLRAGYYWMAMEHDCYQHARKCHKCQIYADKIHVPPHALNV 1894
++H G H+ M+ ++LR GYY M C ++ +KC +C + + H+ L+
Sbjct: 3 EMHRGICGMHSKSQLMTTRVLRVGYY**TMRKYCTEYVKKCEEC*KFGNISHLLVEELHN 182
Query: 1895 ISSPWPFSMWGIDMIGRIEPKASNGHHFILVAIDYFTKWVEAASYTNVTKQVVAKFIKNN 1954
I +PWPF++ G+D++ R P + ++LV ID FTKW+E ++ V KF+ N
Sbjct: 183 IVAPWPFAI*GVDIL-RPFPLSKRQVKYLLVGIDQFTKWIETEHIAIISIANVRKFV*RN 359
Query: 1955 IICRYGVPSKIITDN 1969
I+C +G+P+ +I+DN
Sbjct: 360 IVC*FGIPNTLISDN 404
>NP595172 polyprotein [Glycine max]
Length = 4659
Score = 90.5 bits (223), Expect = 7e-18
Identities = 138/555 (24%), Positives = 228/555 (40%), Gaps = 14/555 (2%)
Frame = +1
Query: 1613 NAYGKGIGAVIVSPQGHHIPFTARILFECTNNMAEYEACIFGIEEAIDMRIKHLDIYGDS 1672
+A G G+GAV+ GH I + ++ L + Y + I EA+ + +H +
Sbjct: 2689 DASGIGVGAVL-GQNGHPIAYFSKKLAPRMQKQSAYTRELLAITEALS-KFRHYLLGNKF 2862
Query: 1673 ALVINQIKGEWETHHAKLIPYRD-YARRLLTYFTKVELHHIPRDENQMADALATLSSMFR 1731
+ +Q + + P + + + L Y K+E + P +NQ ADAL S MF
Sbjct: 2863 IIRTDQRSLKSLMDQSLQTPEQQAWLHKFLGYDFKIE--YKPGKDNQAADAL---SRMFM 3027
Query: 1732 VNHWNDVPIIKVQRLERPSHVFAIGDVIDQAGENVVDYKPWYYDIKQFLLSREYPPGASK 1791
+ W++ I ++ L I D +KQ L Y GA
Sbjct: 3028 LA-WSEPHSIFLEELRAR----LISDP----------------HLKQ--LMETYKQGAD- 3135
Query: 1792 QDKKTLRRLTSRFLLDGDILYKRNYDMVLLRCVDEHEAEQLMHDVHDGTFRTHATGHTMS 1851
S + + +LY + D V++ +E +++ + H HA G T +
Sbjct: 3136 ---------ASHYTVREGLLYWK--DRVVIPA-EEEIVNKILQEYHSSPIGGHA-GITRT 3276
Query: 1852 RKLLRAGYYWMAMEHDCYQHARKCHKCQIYADKIHVPPHALNVISSPWPFSMW---GIDM 1908
L+A +YW M+ D + +KC CQ +P L + P P +W +D
Sbjct: 3277 LARLKAQFYWPKMQEDVKAYIQKCLICQQAKSNNTLPAGLLQPL--PIPQQVWEDVAMDF 3450
Query: 1909 IGRIEPKASNGHHFILVAIDYFTKWVEAASY-TNVTKQVVAKFIKNNIICRYGVPSKIIT 1967
I + S G I+V ID TK+ + +VVA+ ++I+ +G+P I++
Sbjct: 3451 ITGLPN--SFGLSVIMVVIDRLTKYAHFIPLKADYNSKVVAEAFMSHIVKLHGIPRSIVS 3624
Query: 1968 DNGTNLNNNVVQALCEEFKIEHHN---SSPYRPQMNGAVEATNKNIKRIVQKMVTTY-KD 2023
D + Q L FK++ SS Y PQ +G E NK ++ ++ + K
Sbjct: 3625 DRDRVFTSTFWQHL---FKLQGTTLAMSSAYHPQSDGQSEVLNKCLEMYLRCFTYEHPKG 3795
Query: 2024 WHEMLPYALHGYRTTVRSSTGATPFSLVYGMEAVLPLEVEIPSLRVIMEAKLSEAEWCQS 2083
W + LP+A Y T S G TPF +YG E P+L + AE +
Sbjct: 3796 WVKALPWAEFWYNTAYHMSLGMTPFRALYGREP--------PTLTRQACSIDDPAEVREQ 3951
Query: 2084 RYDQLNLIEEKRMDAVARGQSYQARMKTAFDKKVHPREFKVGELVL-----KRRISQQPD 2138
D+ L+ + +++ + R Q MK DKK F++G+ VL R+ S
Sbjct: 3952 LTDRDALLAKLKIN-LTRAQQV---MKRQADKKRLDVSFQIGDEVLVKLQPYRQHSAVLR 4119
Query: 2139 PRGKWTPNYEGPYVV 2153
K + Y GP+ V
Sbjct: 4120 KNQKLSMRYFGPFKV 4164
>BI321666
Length = 430
Score = 83.2 bits (204), Expect = 1e-15
Identities = 45/129 (34%), Positives = 71/129 (54%), Gaps = 3/129 (2%)
Frame = +2
Query: 1850 MSRKLLRAGYYWMAMEHDCYQHARKCHKCQ---IYADKIHVPPHALNVISSPWPFSMWGI 1906
+S +L++ +Y ++ D Y HA+ C+KCQ + + +P H + + F WGI
Sbjct: 2 ISTNVLQSRFYLPSIFKDAYVHAQSCNKCQRTRSVSKRNELPLHTILEVEI---FDYWGI 172
Query: 1907 DMIGRIEPKASNGHHFILVAIDYFTKWVEAASYTNVTKQVVAKFIKNNIICRYGVPSKII 1966
D +G P SN +ILV +DY +KWVEA + ++V KF+K I R GVP +I
Sbjct: 173 DFVGPFPPSFSN--EYILVVVDYVSKWVEAVACQKSDAKIVIKFLKKQIFSRLGVPWVLI 346
Query: 1967 TDNGTNLNN 1975
+ G++L N
Sbjct: 347 DNGGSHLCN 373
>BE804087
Length = 160
Score = 81.3 bits (199), Expect = 4e-15
Identities = 37/53 (69%), Positives = 43/53 (80%)
Frame = +2
Query: 2004 EATNKNIKRIVQKMVTTYKDWHEMLPYALHGYRTTVRSSTGATPFSLVYGMEA 2056
EA NKNIK+ + KM +YKDWHEM +ALH YRT VR+STGATP+SLVYG EA
Sbjct: 2 EAANKNIKKNI*KMTVSYKDWHEMFSFALHMYRTLVRTSTGATPYSLVYGKEA 160
>BE660092 weakly similar to GP|9884624|dbj retroelement pol polyprotein-like
{Arabidopsis thaliana}, partial (13%)
Length = 378
Score = 78.2 bits (191), Expect = 4e-14
Identities = 41/122 (33%), Positives = 71/122 (57%), Gaps = 1/122 (0%)
Frame = -3
Query: 1978 VQALCEEFKIEHHNSSPYRPQMNGAVEATNKNIKRIVQKMVT-TYKDWHEMLPYALHGYR 2036
+ AL +++ + H S+PY PQ NG E +N+ IKRI++K+V + KDW L AL +R
Sbjct: 376 MHALLKKYGVVHRVSTPYHPQTNGQAEISNREIKRILEKIVQPSRKDWSTRLDDALWAHR 197
Query: 2037 TTVRSSTGATPFSLVYGMEAVLPLEVEIPSLRVIMEAKLSEAEWCQSRYDQLNLIEEKRM 2096
T ++ G +P+ +V+G LP+E+E + + S + + R QL+ ++E R+
Sbjct: 196 TAYKAPIGMSPYRVVFGKACHLPVEIEHKAYWAVKTCNFSMDQAGEERKLQLSELDEIRL 17
Query: 2097 DA 2098
+A
Sbjct: 16 EA 11
>TC211815
Length = 704
Score = 48.1 bits (113), Expect(2) = 4e-08
Identities = 42/146 (28%), Positives = 70/146 (47%)
Frame = +2
Query: 2040 RSSTGATPFSLVYGMEAVLPLEVEIPSLRVIMEAKLSEAEWCQSRYDQLNLIEEKRMDAV 2099
+S+T TPF L+YG+ +LP+EV L A++ E Q L+LI++ R D V
Sbjct: 230 QSTTHETPF*LIYGISVMLPIEVGEVFL*RHYFAEV*NKEALQI---DLDLIKQVREDTV 400
Query: 2100 ARGQSYQARMKTAFDKKVHPREFKVGELVLKRRISQQPDPRGKWTPNYEGPYVVKKAFSG 2159
+++ RM F+ K+ G + + S + R K+T N EGP+ ++
Sbjct: 401 IMT*AFKQRMTRCFNSKLPSTV*GRGPSMEGIQRSLEVLVRSKFTTN*EGPFKIRHNSKN 580
Query: 2160 GALILTHMDGVELPNPVNADIVKKYF 2185
GA L + G + N+ +K Y+
Sbjct: 581 GAYKLEELSGKVVLRIWNSMHLKVYY 658
Score = 29.6 bits (65), Expect(2) = 4e-08
Identities = 16/51 (31%), Positives = 23/51 (44%)
Frame = +3
Query: 1968 DNGTNLNNNVVQALCEEFKIEHHNSSPYRPQMNGAVEATNKNIKRIVQKMV 2018
DNG N + I+H +S PQ N EA NK I ++K++
Sbjct: 3 DNGLQFTNRKLNEFPSGLNIKHRVTSVKHPQTNRRAEAANKVILGDLKKLL 155
>TC220664 weakly similar to UP|Q84TE0 (Q84TE0) At5g51080, partial (31%)
Length = 724
Score = 58.2 bits (139), Expect = 4e-08
Identities = 32/86 (37%), Positives = 46/86 (53%)
Frame = +1
Query: 1642 TNNMAEYEACIFGIEEAIDMRIKHLDIYGDSALVINQIKGEWETHHAKLIPYRDYARRLL 1701
TNN AEY A I G++ A+ + I GDS LV QI G W+ + L + A+ L
Sbjct: 64 TNNAAEYRAMILGMKYALKKGFTGIRIQGDSKLVCMQIDGSWKVKNENLSTLYNVAKELK 243
Query: 1702 TYFTKVELHHIPRDENQMADALATLS 1727
F+ ++ H+ R+ N ADA A L+
Sbjct: 244 DKFSSFQISHVLRNFNSDADAQANLA 321
>TC234722 similar to UP|Q6WAY9 (Q6WAY9) Pol (Fragment), partial (32%)
Length = 482
Score = 57.8 bits (138), Expect = 5e-08
Identities = 29/70 (41%), Positives = 41/70 (58%), Gaps = 1/70 (1%)
Frame = -3
Query: 1995 YRPQMNGAVEATNKNIKRIVQKMV-TTYKDWHEMLPYALHGYRTTVRSSTGATPFSLVYG 2053
Y PQ NG E +NK IKR+++ +V ++ KDW L A YR ++ G +PF LVYG
Sbjct: 234 YHPQTNGQAEVSNKEIKRVLENIVVSSRKDWALKLDDAFWAYRIAFKTPIGLSPFQLVYG 55
Query: 2054 MEAVLPLEVE 2063
L +E+E
Sbjct: 54 KACHLSVELE 25
>TC233372 weakly similar to UP|Q84TE0 (Q84TE0) At5g51080, partial (12%)
Length = 561
Score = 53.9 bits (128), Expect = 7e-07
Identities = 30/86 (34%), Positives = 43/86 (49%)
Frame = +1
Query: 1646 AEYEACIFGIEEAIDMRIKHLDIYGDSALVINQIKGEWETHHAKLIPYRDYARRLLTYFT 1705
AEY + I G+ K + + GDS LV NQI+G W+ + + A+ L F
Sbjct: 1 AEYRSLILGLXHXXKKGYKXIIVQGDSLLVCNQIQGLWKIKNQNMGTLCXEAKELKDKFL 180
Query: 1706 KVELHHIPRDENQMADALATLSSMFR 1731
++ HIPR+ N ADA A L+ R
Sbjct: 181 SFKISHIPREYNSEADAQANLAINLR 258
>AW706233
Length = 376
Score = 53.9 bits (128), Expect = 7e-07
Identities = 28/71 (39%), Positives = 37/71 (51%), Gaps = 2/71 (2%)
Frame = -3
Query: 135 ELAKELRREIKANRGNGDSI--KTQDLCLVSKVDVPKKFKVPEFDKYNGLTCPQNHIVKY 192
E L+ KA G D ++L LV + P KFKV FDKY G TCP+NH+ Y
Sbjct: 347 EKLDHLKERFKAIEGGQDYAFANLEELFLVXNIISPPKFKVLNFDKYKGTTCPKNHLKMY 168
Query: 193 VRKMSNYKDND 203
+KM Y ++
Sbjct: 167 CQKMGAYAKDE 135
>BE801213 weakly similar to GP|6691193|gb| F7F22.17 {Arabidopsis thaliana},
partial (3%)
Length = 416
Score = 47.8 bits (112), Expect = 5e-05
Identities = 23/81 (28%), Positives = 43/81 (52%)
Frame = +3
Query: 1945 QVVAKFIKNNIICRYGVPSKIITDNGTNLNNNVVQALCEEFKIEHHNSSPYRPQMNGAVE 2004
++V KF+K NI R+G+P +I+D G++ + + + + + H + Y PQ NG +
Sbjct: 12 KIVIKFLKKNIFSRFGMPRILISDGGSHFYYSQLNKVLKHDSVRHKVETSYHPQTNGQAK 191
Query: 2005 ATNKNIKRIVQKMVTTYKDWH 2025
+N + + +K K WH
Sbjct: 192 VSNIHKENS*EK-----KMWH 239
Score = 35.4 bits (80), Expect = 0.28
Identities = 16/49 (32%), Positives = 27/49 (54%)
Frame = +2
Query: 2015 QKMVTTYKDWHEMLPYALHGYRTTVRSSTGATPFSLVYGMEAVLPLEVE 2063
+ + ++ KDW L AL +T ++ G TPF +VY LP+E++
Sbjct: 227 KNVASSRKDWSSKLEDALWACKTAKKTPIGLTPFQMVYRKACHLPVELK 373
>BI424213
Length = 426
Score = 46.2 bits (108), Expect = 2e-04
Identities = 20/72 (27%), Positives = 39/72 (53%), Gaps = 1/72 (1%)
Frame = +1
Query: 1992 SSPYRPQMNGAVEATNKNIKRIVQKMVT-TYKDWHEMLPYALHGYRTTVRSSTGATPFSL 2050
S+ PQ +G + N+++ +++ ++ +K W E LP+ Y V +T +PF +
Sbjct: 55 STTCHPQTDGQTKVVNRSLSTLLRALLKGNHKSWDEYLPHVEFAYNRGVHRTTKQSPFEV 234
Query: 2051 VYGMEAVLPLEV 2062
VYG + PL++
Sbjct: 235 VYGFNPLTPLDL 270
>BG839293
Length = 781
Score = 39.3 bits (90), Expect = 0.019
Identities = 18/48 (37%), Positives = 30/48 (62%), Gaps = 2/48 (4%)
Frame = +1
Query: 1421 NFEFPVHDAENEEDDDI--PYEITRLLEQEGRAIQPHQEEIEIINFGI 1466
NFE E+E ++D+ P E+ R++ E + + PHQEE E+++ GI
Sbjct: 400 NFEQETSQTEDEGNEDVGLPPELERMVAHEDQEMGPHQEETELVDLGI 543
Database: GMGI
Posted date: Oct 22, 2004 4:58 PM
Number of letters in database: 37,918,896
Number of sequences in database: 63,676
Lambda K H
0.340 0.148 0.488
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 101,958,780
Number of Sequences: 63676
Number of extensions: 1613565
Number of successful extensions: 13216
Number of sequences better than 10.0: 60
Number of HSP's better than 10.0 without gapping: 12761
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 13184
length of query: 2185
length of database: 12,639,632
effective HSP length: 112
effective length of query: 2073
effective length of database: 5,507,920
effective search space: 11417918160
effective search space used: 11417918160
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 15 ( 7.4 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 39 (21.9 bits)
S2: 67 (30.4 bits)
Medicago: description of AC149039.3


